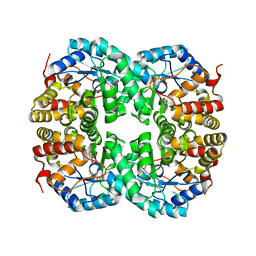
7C0D
 | |
7C0E
 
 | |
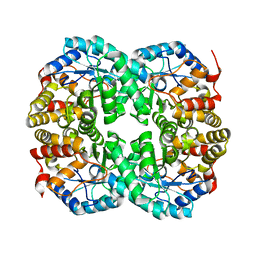
7C0C
 | |
7BYW
 
 | |
7BYU
 
 | | Crystal structure of Acidovorax avenae L-fucose mutarotase (apo form) | | 分子名称: | 1,2-ETHANEDIOL, 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, L-fucose mutarotase | | 著者 | Watanabe, Y, Fukui, Y, Watanabe, S. | | 登録日 | 2020-04-24 | | 公開日 | 2020-05-27 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.206 Å) | | 主引用文献 | Functional and structural characterization of a novel L-fucose mutarotase involved in non-phosphorylative pathway of L-fucose metabolism.
Biochem.Biophys.Res.Commun., 528, 2020
|
|
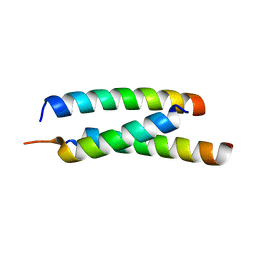
5JGE
 | |
6JNJ
 
 | |
6L06
 
 | |
6L07
 
 | |
6JNK
 
 | |
6J7C
 
 | | Crystal structure of proline racemase-like protein from Thermococcus litoralis in complex with proline | | 分子名称: | PROLINE, Proline racemase | | 著者 | Watanabe, Y, Watanabe, S, Itoh, Y, Watanabe, Y. | | 登録日 | 2019-01-17 | | 公開日 | 2019-02-27 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Crystal structure of substrate-bound bifunctional proline racemase/hydroxyproline epimerase from a hyperthermophilic archaeon.
Biochem. Biophys. Res. Commun., 511, 2019
|
|
6K9Y
 
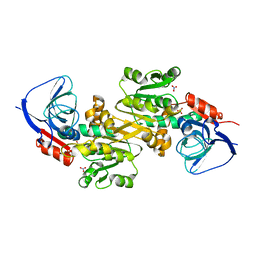 | | Crystal structure of human VAT-1 | | 分子名称: | NITRATE ION, Synaptic vesicle membrane protein VAT-1 homolog | | 著者 | Watanabe, Y, Endo, T. | | 登録日 | 2019-06-19 | | 公開日 | 2020-02-12 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Structural basis for interorganelle phospholipid transport mediated by VAT-1.
J.Biol.Chem., 295, 2020
|
|
2KZK
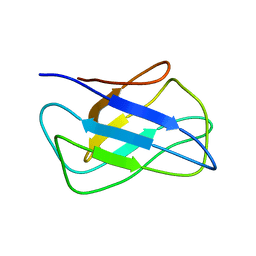 
 | | Solution structure of alpha-mannosidase binding domain of Atg34 | | 分子名称: | Uncharacterized protein YOL083W | | 著者 | Watanabe, Y, Noda, N, Kumeta, H, Suzuki, K, Ohsumi, Y, Inagaki, F. | | 登録日 | 2010-06-18 | | 公開日 | 2010-07-21 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Selective transport of alpha-mannosidase by autophagic pathways: structural basis for cargo recognition by Atg19 and Atg34.
J.Biol.Chem., 285, 2010
|
|
2KZB
 
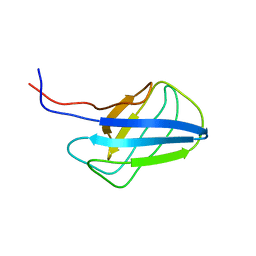 | | Solution structure of alpha-mannosidase binding domain of Atg19 | | 分子名称: | Autophagy-related protein 19 | | 著者 | Watanabe, Y, Noda, N, Kumeta, H, Suzuki, K, Ohsumi, Y, Inagaki, F. | | 登録日 | 2010-06-15 | | 公開日 | 2010-07-21 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Selective transport of alpha-mannosidase by autophagic pathways: structural basis for cargo recognition by Atg19 and Atg34.
J.Biol.Chem., 285, 2010
|
|
7YPD
 
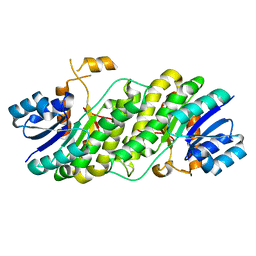 | |
5AUJ
 
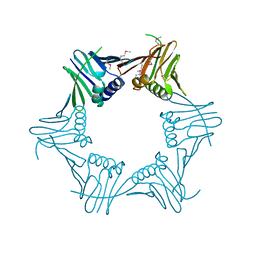 | |
4YTV
 
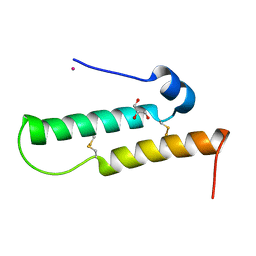 | | Crystal structure of Mdm35 | | 分子名称: | COBALT (II) ION, GLYCEROL, Mitochondrial distribution and morphology protein 35 | | 著者 | Watanabe, Y, Tamura, Y, Kawano, S, Endo, T. | | 登録日 | 2015-03-18 | | 公開日 | 2015-08-12 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.45 Å) | | 主引用文献 | Structural and mechanistic insights into phospholipid transfer by Ups1-Mdm35 in mitochondria.
Nat Commun, 6, 2015
|
|
4YTW
 
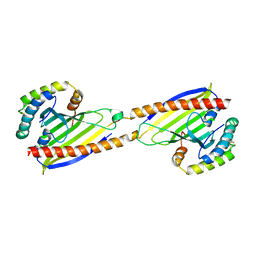 | | Crystal structure of Ups1-Mdm35 complex | | 分子名称: | Mitochondrial distribution and morphology protein 35, Protein UPS1, mitochondrial | | 著者 | Watanabe, Y, Tamura, Y, Kawano, S, Endo, T. | | 登録日 | 2015-03-18 | | 公開日 | 2015-08-12 | | 最終更新日 | 2020-02-05 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Structural and mechanistic insights into phospholipid transfer by Ups1-Mdm35 in mitochondria.
Nat Commun, 6, 2015
|
|
4YTX
 
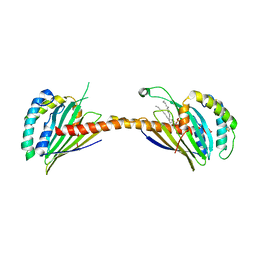 | | Crystal structure of Ups1-Mdm35 complex with PA | | 分子名称: | 1,2-DILAUROYL-SN-GLYCERO-3-PHOSPHATE, Mitochondrial distribution and morphology protein 35, Protein UPS1, ... | | 著者 | Watanabe, Y, Tamura, Y, Kawano, S, Endo, T. | | 登録日 | 2015-03-18 | | 公開日 | 2015-08-12 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Structural and mechanistic insights into phospholipid transfer by Ups1-Mdm35 in mitochondria.
Nat Commun, 6, 2015
|
|
5AZF
 
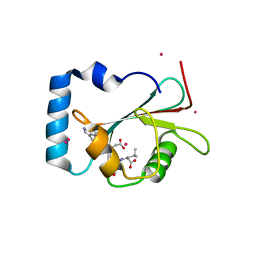 | | Crystal structure of LGG-1 complexed with a WEEL peptide | | 分子名称: | CADMIUM ION, Protein lgg-1, SULFATE ION, ... | | 著者 | Watanabe, Y, Noda, N.N. | | 登録日 | 2015-10-05 | | 公開日 | 2015-12-30 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Structural Basis of the Differential Function of the Two C. elegans Atg8 Homologs, LGG-1 and LGG-2, in Autophagy.
Mol.Cell, 60, 2015
|
|
5AZH
 
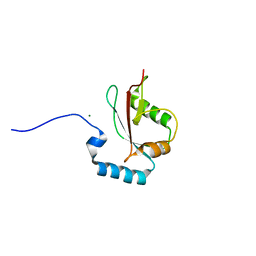 | | Crystal structure of LGG-2 fused with an EEEWEEL peptide | | 分子名称: | EEEWEEL peptide,Protein lgg-2, MAGNESIUM ION | | 著者 | Watanabe, Y, Fujioka, Y, Noda, N.N. | | 登録日 | 2015-10-05 | | 公開日 | 2015-12-30 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structural Basis of the Differential Function of the Two C. elegans Atg8 Homologs, LGG-1 and LGG-2, in Autophagy.
Mol.Cell, 60, 2015
|
|
5AZG
 
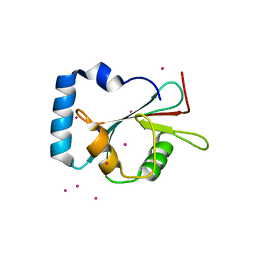 | | Crystal structure of LGG-1 complexed with a UNC-51 peptide | | 分子名称: | CADMIUM ION, Protein lgg-1, Serine/threonine-protein kinase unc-51 | | 著者 | Watanabe, Y, Fujioka, Y, Noda, N.N. | | 登録日 | 2015-10-05 | | 公開日 | 2015-12-30 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.81 Å) | | 主引用文献 | Structural Basis of the Differential Function of the Two C. elegans Atg8 Homologs, LGG-1 and LGG-2, in Autophagy.
Mol.Cell, 60, 2015
|
|
3VU4
 
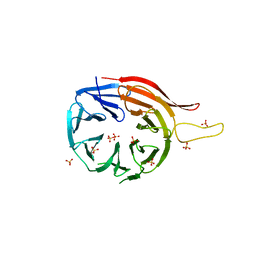 | |
3WQH
 
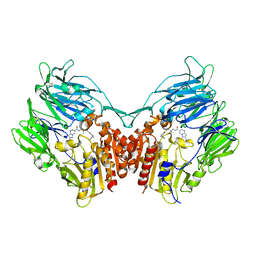 | | Crystal Structure of human DPP-IV in complex with Anagliptin | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Dipeptidyl peptidase 4, N-[2-({2-[(2S)-2-cyanopyrrolidin-1-yl]-2-oxoethyl}amino)-2-methylpropyl]-2-methylpyrazolo[1,5-a]pyrimidine-6-carboxamide | | 著者 | Watanabe, Y.S, Okada, S, Motoyama, T, Takahashi, R, Adachi, H, Oka, M. | | 登録日 | 2014-01-27 | | 公開日 | 2015-07-15 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (2.85 Å) | | 主引用文献 | Anagliptin, a potent dipeptidyl peptidase IV inhibitor: its single-crystal structure and enzyme interactions.
J Enzyme Inhib Med Chem, 30, 2015
|
|
8GST
 
 | | Crystal structure of L-2,4-diketo-3-deoxyrhamnonate hydrolase from Sphingomonas sp. (pyruvate bound-form) | | 分子名称: | L-2,4-diketo-3-deoxyrhamnonate hydrolase, MAGNESIUM ION, PYRUVIC ACID | | 著者 | Fukuhara, S, Watanabe, Y, Watanabe, S, Nishiwaki, H. | | 登録日 | 2022-09-07 | | 公開日 | 2023-02-08 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.71 Å) | | 主引用文献 | Crystal Structure of l-2,4-Diketo-3-deoxyrhamnonate Hydrolase Involved in the Nonphosphorylated l-Rhamnose Pathway from Bacteria.
Biochemistry, 62, 2023
|
|
