1RPU
 
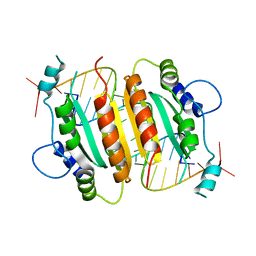 | | Crystal Structure of CIRV p19 bound to siRNA | | Descriptor: | 19 kDa protein, 5'-R(P*CP*GP*UP*AP*CP*GP*CP*GP*UP*CP*AP*CP*GP*CP*GP*UP*AP*CP*GP*UP*U)-3' | | Authors: | Vargason, J.M, Szittya, G, Burgyan, J, Hall, T.M.T. | | Deposit date: | 2003-12-03 | | Release date: | 2004-01-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Size selective recognition of siRNA by an RNA silencing suppressor
Cell(Cambridge,Mass.), 115, 2003
|
|
1IH3
 | |
1IH4
 | |
1IH6
 
 | |
1L6B
 | |
1IH2
 | |
1IH1
 | |
1F6C
 
 | |
1F6E
 
 | |
1F69
 | |
1F6I
 | |
1F6J
 
 | |
6U0T
 | | Protofilament Ribbon Flagellar Proteins Rib43a-S | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Ichikawa, M, Khalifa, A.A.Z, Vargas, J, Basu, K, Bui, K.H. | | Deposit date: | 2019-08-14 | | Release date: | 2019-09-25 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (4.16 Å) | | Cite: | Tubulin lattice in cilia is in a stressed form regulated by microtubule inner proteins.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6U0U
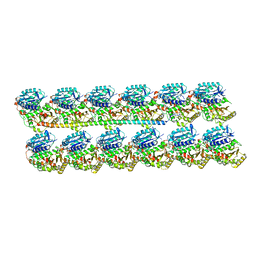 
 | | Protofilament Ribbon Flagellar Proteins Rib43a-L | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Ichikawa, M, Khalifa, A.A.Z, Vargas, J, Basu, K, Bui, K.H. | | Deposit date: | 2019-08-14 | | Release date: | 2019-09-25 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (4.16 Å) | | Cite: | Tubulin lattice in cilia is in a stressed form regulated by microtubule inner proteins.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6U0H
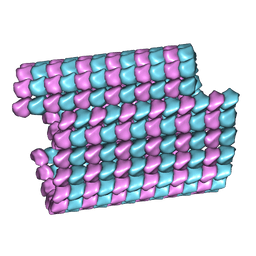 
 | | Tubulin lattice of the ciliary doublet microtubule from Tetrahymena thermophila | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Ichikawa, M, Khalifa, A.A.Z, Vargas, J, Basu, K, Bui, K.H. | | Deposit date: | 2019-08-14 | | Release date: | 2019-09-25 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Tubulin lattice in cilia is in a stressed form regulated by microtubule inner proteins.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
8QTZ
 
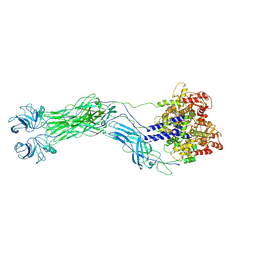 | | Cryo-EM reconstruction of VP5*/VP8* assembly from SA11 Rotavirus Tripsinized Triple Layered Particle | | Descriptor: | Outer capsid protein VP4 | | Authors: | Asensio-Cob, D, Perez-Mata, C, Gomez-Blanco, J, Vargas, J, Rodriguez, J.M, Luque, D. | | Deposit date: | 2023-10-13 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (4.27 Å) | | Cite: | Structural basis of rotavirus spike proteolytic activation
To Be Published
|
|
8SPK
 
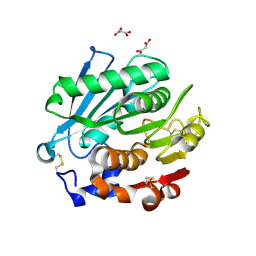 | | Crystal structure of Antarctic PET-degrading enzyme | | Descriptor: | Lipase 1, MALONATE ION | | Authors: | Furtado, A.A, Blazquez-Sanchez, P, Grinen, A, Vargas, J.A, Leonardo, D.A, Sculaccio, S.A, Pereira, H.M, Diez, B, Garratt, R.C, Ramirez-Sarmiento, C.A. | | Deposit date: | 2023-05-03 | | Release date: | 2023-08-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Engineering the catalytic activity of an Antarctic PET-degrading enzyme by loop exchange.
Protein Sci., 32, 2023
|
|
6W7W
 
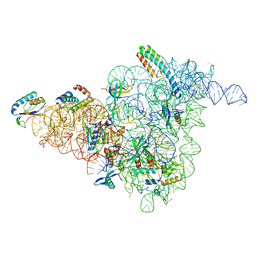 | | 30S-Inactive-low-Mg2+ Class B | | Descriptor: | 16S rRNA, 30S ribosomal protein S12, 30S ribosomal protein S15, ... | | Authors: | Jahagirdar, D, Jha, V, Basu, B, Gomez-Blanco, J, Vargas, J, Ortega, J. | | Deposit date: | 2020-03-19 | | Release date: | 2020-10-21 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.
Rna, 26, 2020
|
|
6W6K
 
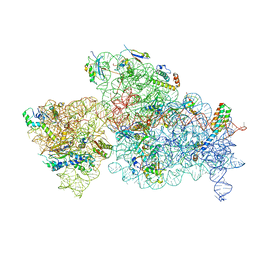 | | 30S-Activated-high-Mg2+ | | Descriptor: | 16S rRNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | Authors: | Jahagirdar, D, Jha, V, Basu, B, Gomez-Blanco, J, Vargas, J, Ortega, J. | | Deposit date: | 2020-03-17 | | Release date: | 2020-10-21 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.
Rna, 26, 2020
|
|
6W77
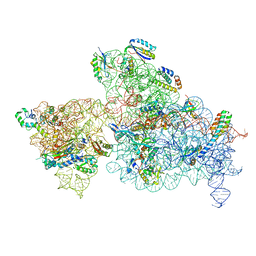 
 | | 30S-Inactivated-high-Mg2+ Class A | | Descriptor: | 30S ribosomal protein S10, 30S ribosomal protein S11, 30S ribosomal protein S12, ... | | Authors: | Jahagirdar, D, Jha, V, Basu, B, Gomez-Blanco, J, Vargas, J, Ortega, J. | | Deposit date: | 2020-03-18 | | Release date: | 2020-10-21 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.
Rna, 26, 2020
|
|
6W7M
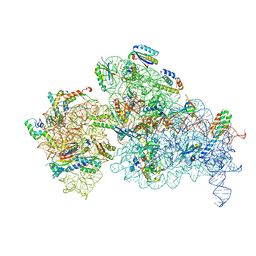 
 | | 30S-Inactive-high-Mg2+ + carbon layer | | Descriptor: | 16S rRNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | Authors: | Jahagirdar, D, Jha, V, Basu, B, Gomez-Blanco, J, Vargas, J, Ortega, J. | | Deposit date: | 2020-03-19 | | Release date: | 2020-10-21 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.
Rna, 26, 2020
|
|
6W7N
 
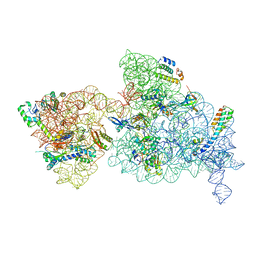 | | 30S-Inactive-low-Mg2+ Class A | | Descriptor: | 16S rRNA, 30S ribosomal protein S10, 30S ribosomal protein S12, ... | | Authors: | Jahagirdar, D, Jha, V, Basu, B, Gomez-Blanco, J, Vargas, J, Ortega, J. | | Deposit date: | 2020-03-19 | | Release date: | 2020-10-21 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.
Rna, 26, 2020
|
|
8OLE
 
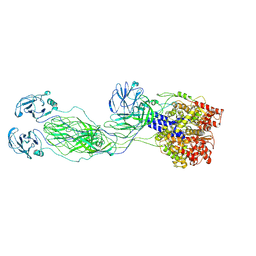 | | Cryo-EM reconstruction of VP4 assembly from SA11 Rotavirus Non-Tripsinized Triple Layered Particle | | Descriptor: | Outer capsid protein VP4 | | Authors: | Asensio-Cob, D, Perez-Mata, C, Gomez-Blanco, J, Vargas, J, Rodriguez, J.M, Luque, D. | | Deposit date: | 2023-03-30 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structural basis of rotavirus spike proteolytic activation
To Be Published
|
|
8OLC
 
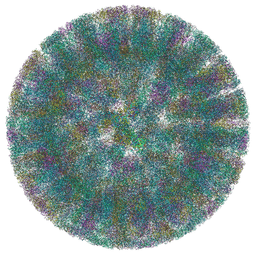 | | SA11 Rotavirus Trypsinized Triple Layered Particle | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Inner capsid protein VP2, ... | | Authors: | Asensio-Cob, D, Perez-Mata, C, Gomez-Blanco, J, Vargas, J, Rodriguez, J.M, Luque, D. | | Deposit date: | 2023-03-30 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (3.48 Å) | | Cite: | Structural basis of rotavirus spike proteolytic activation
To Be Published
|
|
8OLB
 
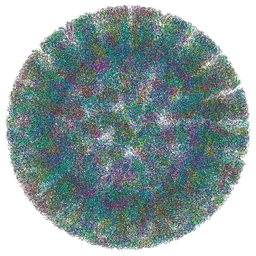 | | SA11 Rotavirus Non-tripsinized Triple Layered Particle | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Inner capsid protein VP2, ... | | Authors: | Asensio-Cob, D, Perez-Mata, C, Gomez-Blanco, J, Vargas, J, Rodriguez, J.M, Luque, D. | | Deposit date: | 2023-03-30 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural basis of rotavirus spike proteolytic activation
To Be Published
|
|
