1E5D
 
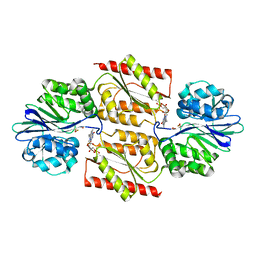 | | RUBREDOXIN OXYGEN:OXIDOREDUCTASE (ROO) FROM ANAEROBE DESULFOVIBRIO GIGAS | | Descriptor: | FLAVIN MONONUCLEOTIDE, MU-OXO-DIIRON, OXYGEN MOLECULE, ... | | Authors: | Frazao, C, Silva, G, Gomes, C.M, Matias, P, Coelho, R, Sieker, L, Macedo, S, Liu, M.Y, Oliveira, S, Teixeira, M, Xavier, A.V, Rodrigues-Pousada, C, Carrondo, M.A, Le Gall, J. | | Deposit date: | 2000-07-24 | | Release date: | 2000-11-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of a Dioxygen Reduction Enzyme from Desulfovibrio Gigas
Nat.Struct.Biol., 7, 2000
|
|
6XIR
 
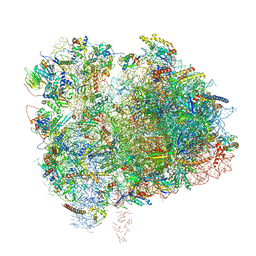 | | Cryo-EM Structure of K63 Ubiquitinated Yeast Translocating Ribosome under Oxidative Stress | | Descriptor: | 18S ribosomal RNA, 35S ribosomal RNA, 40S ribosomal protein S0-A, ... | | Authors: | Zhou, Y, Bartesaghi, A, Silva, G.M. | | Deposit date: | 2020-06-21 | | Release date: | 2020-08-26 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural impact of K63 ubiquitin on yeast translocating ribosomes under oxidative stress.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6XIQ
 
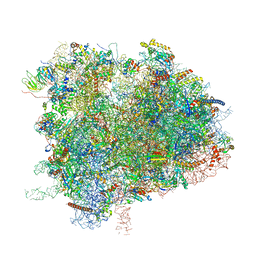 | | Cryo-EM Structure of K63R Ubiquitin Mutant Ribosome under Oxidative Stress | | Descriptor: | 18S ribosomal RNA, 35S ribosomal RNA, 40S ribosomal protein S0-A, ... | | Authors: | Zhou, Y, Bartesaghi, A, Silva, G.M. | | Deposit date: | 2020-06-21 | | Release date: | 2020-08-26 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Structural impact of K63 ubiquitin on yeast translocating ribosomes under oxidative stress.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
1B24
 
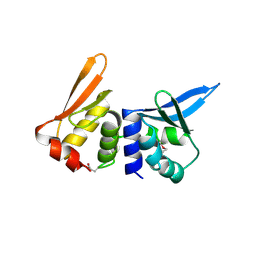 | | I-DMOI, INTRON-ENCODED ENDONUCLEASE | | Descriptor: | Homing endonuclease I-DmoI | | Authors: | Van Roey, P, Silva, G.H. | | Deposit date: | 1998-12-03 | | Release date: | 1999-03-24 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the thermostable archaeal intron-encoded endonuclease I-DmoI.
J.Mol.Biol., 286, 1999
|
|
2YPF
 
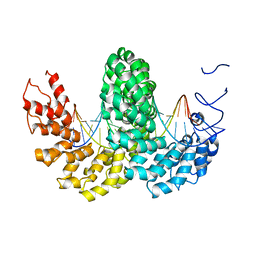 | | Structure of the AvrBs3-DNA complex provides new insights into the initial thymine-recognition mechanism | | Descriptor: | 5'-D(*TP*AP*GP*AP*GP*GP*GP*TP*TP*AP*GP*GP*TP*TP *TP*AP*TP*AP*TP*AP*AP)-3', 5'-D(*TP*TP*TP*AP*TP*AP*TP*AP*AP*AP*CP*CP*TP*AP *AP*CP*CP*CP*TP*CP*TP*AP)-3', AVRBS3 | | Authors: | Stella, S, Molina, R, Yefimenko, I, Prieto, J, Silva, G.H, Bertonati, C, Juillerat, A, Duchateau, P, Montoya, G. | | Deposit date: | 2012-10-30 | | Release date: | 2013-08-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structure of the Avrbs3-DNA Complex Provides New Insights Into the Initial Thymine-Recognition Mechanism
Acta Crystallogr.,Sect.D, 69, 2013
|
|
