2CB4
 
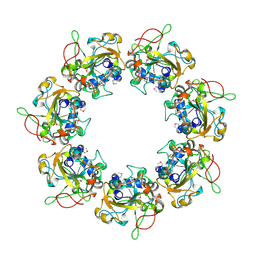 | | Crystal structure of the catalytic domain of the mosquitocidal toxin from Bacillus sphaericus, mutant E197Q | | Descriptor: | MOSQUITOCIDAL TOXIN | | Authors: | Reinert, D.J, Carpusca, I, Aktories, K, Schulz, G.E. | | Deposit date: | 2005-12-29 | | Release date: | 2006-02-22 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the Mosquitocidal Toxin from Bacillus Sphaericus.
J.Mol.Biol., 357, 2006
|
|
2CGL
 
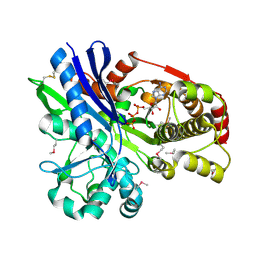 | |
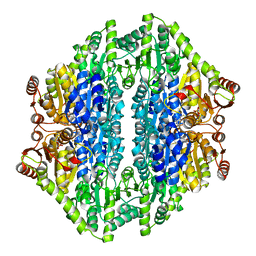
2AG0
 
 | |
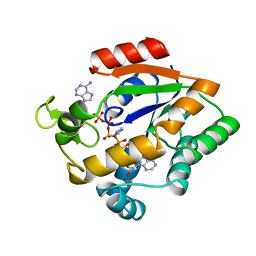
1AKY
 
 | |
1E4A
 
 | |
1E47
 
 | |
1E4C
 
 | |
1E49
 
 | |
1CGT
 
 | |
1E46
 
 | |
1E4B
 
 | |
1E48
 
 | |
2V2B
 
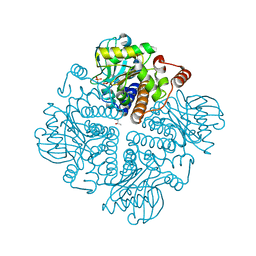 | |
1OG3
 
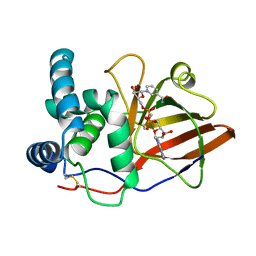 | | Crystal structure of the eukaryotic mono-ADP-ribosyltransferase ART2.2 mutant E189I in complex with NAD | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, T-CELL ECTO-ADP-RIBOSYLTRANSFERASE 2 | | Authors: | Ritter, H, Koch-Nolte, F, Marquez, V.E, Schulz, G.E. | | Deposit date: | 2003-04-24 | | Release date: | 2003-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Substrate Binding and Catalysis of Ecto-Adp-Ribosyltransferase 2.2 From Rat
Biochemistry, 42, 2003
|
|
1OG1
 
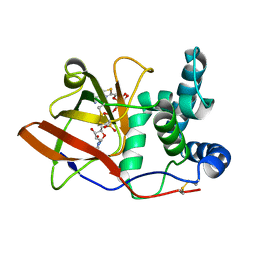 | | CRYSTAL STRUCTURE OF THE EUCARYOTIC MONO-ADP-RIBOSYLTRANSFERASE ART2.2 IN COMPLEX WITH TAD | | Descriptor: | BETA-METHYLENE-THIAZOLE-4-CARBOXYAMIDE-ADENINE DINUCLEOTIDE, T-CELL ECTO-ADP-RIBOSYLTRANSFERASE 2 | | Authors: | Ritter, H, Koch-Nolte, F, Marquez, V.E, Schulz, G.E. | | Deposit date: | 2003-04-23 | | Release date: | 2003-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Substrate Binding and Catalysis of Ecto-Adp-Ribosyltransferase 2.2 From Rat
Biochemistry, 42, 2003
|
|
1O9H
 
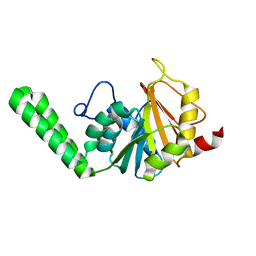 | |
1OG4
 
 | | Crystal Structure of the Eucaryotic Mono-ADP-Ribosyltransferase ART2.2 Mutant E189A in Complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, T-CELL ECTO-ADP-RIBOSYLTRANSFERASE 2 | | Authors: | Ritter, H, Koch-Nolte, F, Marquez, V.E, Schulz, G.E. | | Deposit date: | 2003-04-24 | | Release date: | 2003-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Substrate Binding and Catalysis of Ecto-Adp-Ribosyltransferase 2.2 From Rat
Biochemistry, 42, 2003
|
|
1RPX
 
 | |
1OKX
 
 | | Binding Structure of Elastase Inhibitor Scyptolin A | | Descriptor: | ELASTASE 1, SCYPTOLIN A | | Authors: | Matern, U, Schleberger, C, Jelakovic, S, Weckesser, J, Schulz, G.E. | | Deposit date: | 2003-07-31 | | Release date: | 2003-10-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Binding Structure of Elastase Inhibitor Scyptolin A
Chem.Biol., 10, 2003
|
|
1PBG
 
 | |
2PRN
 
 | | RHODOPSEUDOMONAS BLASTICA PORIN, TRIPLE MUTANT E1M, E99W, A116W | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, MAGNESIUM ION, PORIN | | Authors: | Maveyraud, L, Schmid, B, Schulz, G.E. | | Deposit date: | 1998-06-12 | | Release date: | 1999-01-13 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Porin mutants with new channel properties.
Protein Sci., 7, 1998
|
|
2SQC
 
 | |
2PAW
 
 | | THE CATALYTIC FRAGMENT OF POLY(ADP-RIBOSE) POLYMERASE | | Descriptor: | POLY(ADP-RIBOSE) POLYMERASE | | Authors: | Ruf, A, Schulz, G.E. | | Deposit date: | 1997-11-25 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Inhibitor and NAD+ binding to poly(ADP-ribose) polymerase as derived from crystal structures and homology modeling.
Biochemistry, 37, 1998
|
|
2PAX
 
 | |
1BH3
 
 | |
