5DOT
 
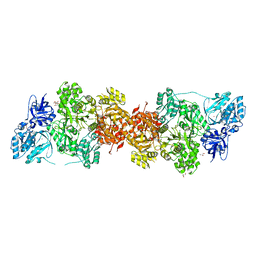 | | Crystal Structure of Human Carbamoyl phosphate synthetase I (CPS1), apo form | | Descriptor: | 1,2-ETHANEDIOL, Carbamoyl-phosphate synthase [ammonia], mitochondrial, ... | | Authors: | Polo, L.M, de Cima, S, Fita, I, Rubio, V. | | Deposit date: | 2015-09-11 | | Release date: | 2015-12-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis.
Sci Rep, 5, 2015
|
|
5DY1
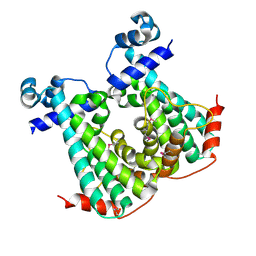 
 | |
2WXB
 
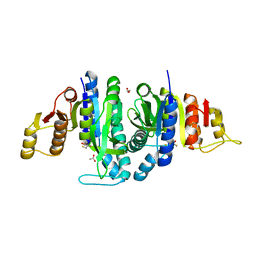 | | Acetylglutamate kinase from Escherichia coli free of substrates | | Descriptor: | (2R,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, 1,2-ETHANEDIOL, ACETATE ION, ... | | Authors: | Ramon-Maiques, S, Gil-Ortiz, F, Rubio, V. | | Deposit date: | 2009-11-06 | | Release date: | 2010-05-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Two Crystal Structures of Escherichia Coli N-Acetyl-L-Glutamate Kinase Demonstrate the Cycling between Open and Closed Conformations.
J.Mol.Biol., 399, 2010
|
|
2X2W
 
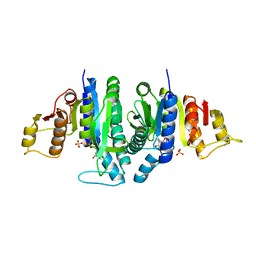 | |
2XKO
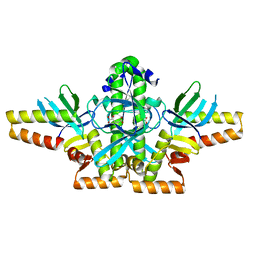 
 | |
2XLB
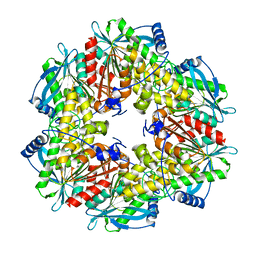 
 | | Acetyl xylan esterase from Bacillus pumilus without ligands | | Descriptor: | ACETYL XYLAN ESTERASE | | Authors: | Gil-Ortiz, F, Montoro-Garcia, S, Polo, L.M, Rubio, V, Sanchez-Ferrer, A. | | Deposit date: | 2010-07-20 | | Release date: | 2011-05-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Crystal Structure of the Cephalosporin Deacetylating Enzyme Acetyl Xylan Esterase Bound to Paraoxon Explains the Low Sensitivity of This Serine Hydrolase to Organophosphate Inactivation.
Biochem.J., 436, 2011
|
|
2XG8
 
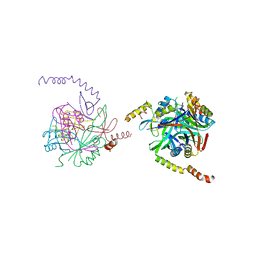 | |
2XKP
 
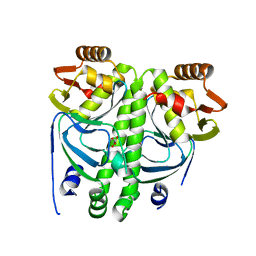 | |
2XLC
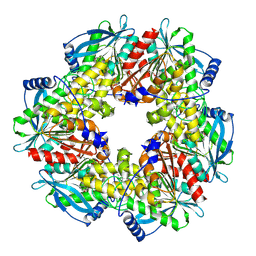 
 | | Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon | | Descriptor: | ACETYL XYLAN ESTERASE, DIETHYL PHOSPHONATE | | Authors: | Gil-Ortiz, F, Montoro-Garcia, S, Polo, L.M, Rubio, V, Sanchez-Ferrer, A. | | Deposit date: | 2010-07-20 | | Release date: | 2011-05-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Crystal Structure of the Cephalosporin Deacetylating Enzyme Acetyl Xylan Esterase Bound to Paraoxon Explains the Low Sensitivity of This Serine Hydrolase to Organophosphate Inactivation.
Biochem.J., 436, 2011
|
|
2XHK
 
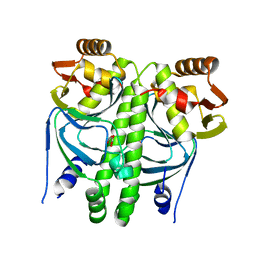 | |
2XGX
 
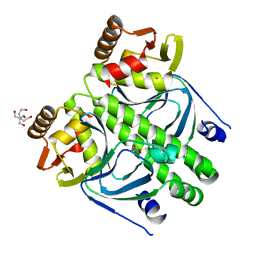 | | Crystal structure of transcription factor NtcA from Synechococcus elongatus (mercury derivative) | | Descriptor: | 2-OXOGLUTARIC ACID, 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLOBAL NITROGEN REGULATOR, ... | | Authors: | Llacer, J.L, Castells, M.A, Rubio, V. | | Deposit date: | 2010-06-08 | | Release date: | 2010-08-18 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural Basis for the Regulation of Ntca-Dependent Transcription by Proteins Pipx and Pii.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2YFK
 
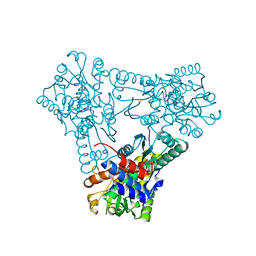 | |
4CR8
 
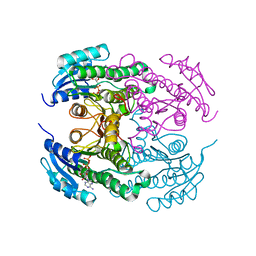 | | Crystal structure of the N-acetyl-D-mannosamine dehydrogenase with NAD | | Descriptor: | N-ACYLMANNOSAMINE 1-DEHYDROGENASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Gil-Ortiz, F, Sola-Carvajal, A, Garcia-Carmona, F, Sanchez-Ferrer, A, Rubio, V. | | Deposit date: | 2014-02-25 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structures and Functional Studies Clarify Substrate Selectivity and Catalytic Residues for the Unique Orphan Enzyme N-Acetyl-D-Mannosamine Dehydrogenase.
Biochem.J., 462, 2014
|
|
4CR7
 
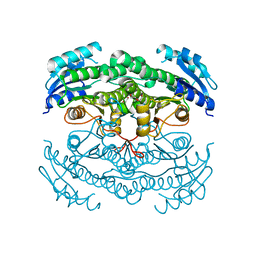 | | Crystal structure of the N-acetyl-D-mannosamine dehydrogenase with n-acetylmannosamine | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-mannopyranose, N-ACYLMANNOSAMINE 1-DEHYDROGENASE, TETRAETHYLENE GLYCOL, ... | | Authors: | Gil-Ortiz, F, Sola-Carvajal, A, Garcia-Carmona, F, Sanchez-Ferrer, A, Rubio, V. | | Deposit date: | 2014-02-25 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal Structures and Functional Studies Clarify Substrate Selectivity and Catalytic Residues for the Unique Orphan Enzyme N-Acetyl-D-Mannosamine Dehydrogenase.
Biochem.J., 462, 2014
|
|
4CR6
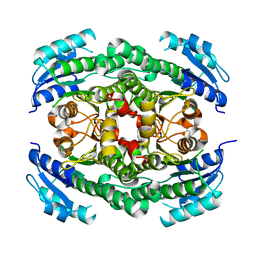 
 | | Crystal structure of the N-acetyl-D-mannosamine dehydrogenase without substrates | | Descriptor: | N-ACYLMANNOSAMINE 1-DEHYDROGENASE, alpha-D-mannopyranose | | Authors: | Gil-Ortiz, F, Sola-Carvajal, A, Garcia-Carmona, F, Sanchez-Ferrer, A, Rubio, V. | | Deposit date: | 2014-02-25 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structures and Functional Studies Clarify Substrate Selectivity and Catalytic Residues for the Unique Orphan Enzyme N-Acetyl-D-Mannosamine Dehydrogenase.
Biochem.J., 462, 2014
|
|
5NLC
 
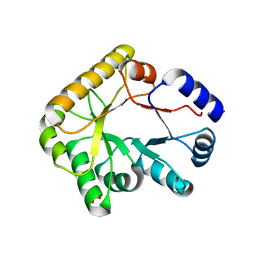 | | Structure of PipY, the COG0325 family member of Synechococcus elongatus PCC7942,without PLP | | Descriptor: | PHOSPHATE ION, PipY | | Authors: | Tremino, L, Forcada-Nadal, A, Contreras, A, Rubio, V. | | Deposit date: | 2017-04-04 | | Release date: | 2017-09-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Studies on cyanobacterial protein PipY shed light on structure, potential functions, and vitamin B6 -dependent epilepsy.
FEBS Lett., 591, 2017
|
|
5NM8
 
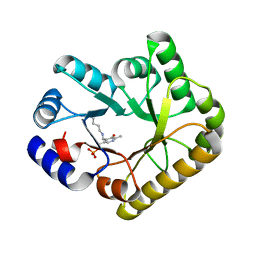 | | Structure of PipY, the COG0325 family member of Synechococcus elongatus PCC7942, with PLP bound | | Descriptor: | CALCIUM ION, PYRIDOXAL-5'-PHOSPHATE, PipY | | Authors: | Tremino, L, Forcada-Nadal, A, Contreras, A, Rubio, V. | | Deposit date: | 2017-04-05 | | Release date: | 2017-09-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Studies on cyanobacterial protein PipY shed light on structure, potential functions, and vitamin B6 -dependent epilepsy.
FEBS Lett., 591, 2017
|
|
