2IUV
 
 | |
2Q7Q
 
 | |
3R6M
 
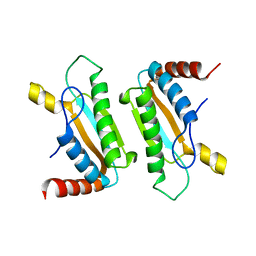
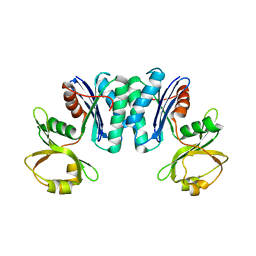 | | Crystal structure of Vibrio parahaemolyticus YeaZ | | Descriptor: | YeaZ, resuscitation promoting factor | | Authors: | Roujeinikova, A, Aydin, I. | | Deposit date: | 2011-03-21 | | Release date: | 2011-09-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Analysis of the Essential Resuscitation Promoting Factor YeaZ Suggests a Mechanism of Nucleotide Regulation through Dimer Reorganization.
Plos One, 6, 2011
|
|
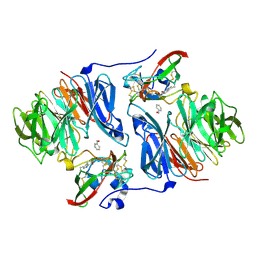
2OK6
 | |
3CYP
 
 | |
3DPP
 
 | |
3DPQ
 
 | |
3DPO
 
 | |
2FAC
 
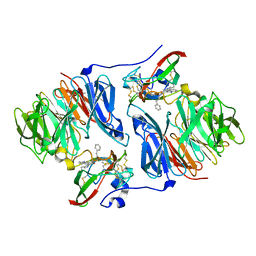
 | | Crystal structure of E. coli hexanoyl-ACP | | Descriptor: | Acyl carrier protein, S-(2-{[N-(2-HYDROXY-4-{[HYDROXY(OXIDO)PHOSPHINO]OXY}-3,3-DIMETHYLBUTANOYL)-BETA-ALANYL]AMINO}ETHYL) HEXANETHIOATE, ZINC ION | | Authors: | Roujeinikova, A. | | Deposit date: | 2005-12-07 | | Release date: | 2006-09-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural Studies of Fatty Acyl-(Acyl Carrier Protein) Thioesters Reveal a Hydrophobic Binding Cavity that Can Expand to Fit Longer Substrates.
J.Mol.Biol., 365, 2007
|
|
2FAE
 
 | | Crystal structure of E. coli decanoyl-ACP | | Descriptor: | Acyl carrier protein, O-PHOSPHOETHANOLAMINE, S-(2-{[N-(2-HYDROXY-4-{[HYDROXY(OXIDO)PHOSPHINO]OXY}-3,3-DIMETHYLBUTANOYL)-BETA-ALANYL]AMINO}ETHYL) DECANETHIOATE, ... | | Authors: | Roujeinikova, A. | | Deposit date: | 2005-12-07 | | Release date: | 2006-09-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural Studies of Fatty Acyl-(Acyl Carrier Protein) Thioesters Reveal a Hydrophobic Binding Cavity that Can Expand to Fit Longer Substrates.
J.Mol.Biol., 365, 2007
|
|
2FAD
 
 | | Crystal structure of E. coli heptanoyl-ACP | | Descriptor: | Acyl carrier protein, S-(2-{[N-(2-HYDROXY-4-{[HYDROXY(OXIDO)PHOSPHINO]OXY}-3,3-DIMETHYLBUTANOYL)-BETA-ALANYL]AMINO}ETHYL) HEPTANETHIOATE, SODIUM ION, ... | | Authors: | Roujeinikova, A. | | Deposit date: | 2005-12-07 | | Release date: | 2006-09-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural Studies of Fatty Acyl-(Acyl Carrier Protein) Thioesters Reveal a Hydrophobic Binding Cavity that Can Expand to Fit Longer Substrates.
J.Mol.Biol., 365, 2007
|
|
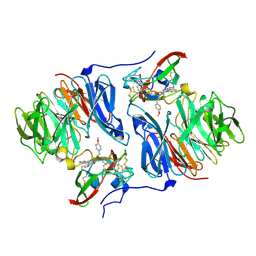
2HKR
 | |
2HJ4
 
 | |
2HJB
 
 | |
2HKM
 
 | |
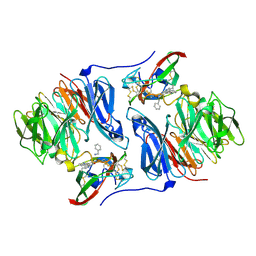
2HXC
 | |
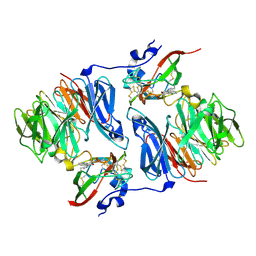
2I0R
 | |
3CYQ
 
 | |
6OFQ
 
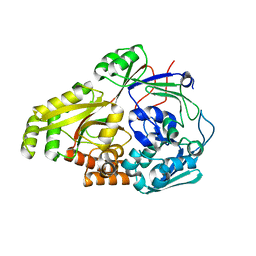 | | ABC transporter-associated periplasmic binding protein DppA from Helicobacter pylori in complex with peptide STSA | | Descriptor: | Heme-binding protein A / AI-2 binding protein A, SER-THR-SER-ALA | | Authors: | Rahman, M.M, Machuca, M.A, Khan, M.F, Barlow, C.K, Schittenhelm, R.B, Roujeinikova, A. | | Deposit date: | 2019-03-31 | | Release date: | 2019-08-21 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Molecular Basis of Unexpected Specificity of ABC Transporter-Associated Substrate-Binding Protein DppA from Helicobacter pylori.
J.Bacteriol., 201, 2019
|
|
6Q0G
 
 | |
6PY3
 
 | |
6PU3
 
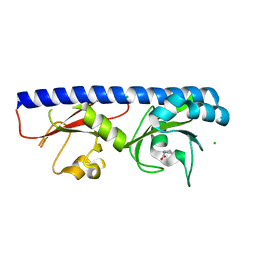
 | | ABC transporter-associated periplasmic binding protein DppA from Helicobacter pylori | | Descriptor: | Heme-binding protein A, SER-THR-SER-ALA | | Authors: | Rahman, M.M, Machuca, M.A, Khan, M.F, Barlow, C.K, Schittenhelm, R.B, Roujeinikova, A. | | Deposit date: | 2019-07-16 | | Release date: | 2019-08-21 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Molecular Basis of Unexpected Specificity of ABC Transporter-Associated Substrate-Binding Protein DppA from Helicobacter pylori.
J.Bacteriol., 201, 2019
|
|
6PXY
 
 | |
5WBF
 
 | | Double CACHE (dCACHE) sensing domain of TlpC chemoreceptor from Helicobacter pylori | | Descriptor: | GLYCEROL, LACTIC ACID, Methyl-accepting chemotaxis transducer (TlpC) | | Authors: | Machuca, M.A, Johnson, K.S, Liu, Y.C, Steer, D.L, Ottemann, K.M, Roujeinikova, A. | | Deposit date: | 2017-06-28 | | Release date: | 2017-11-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Helicobacter pylori chemoreceptor TlpC mediates chemotaxis to lactate.
Sci Rep, 7, 2017
|
|
6W3O
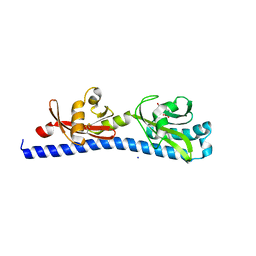 
 | | Crystal structure of ligand-binding domain of Campylobacter jejuni chemoreceptor Tlp3 in complex with 4-methylisoleucine | | Descriptor: | 4-methylisoleucine, CHLORIDE ION, Methyl-accepting chemotaxis protein, ... | | Authors: | Khan, M.F, Machuca, M.A, Rahman, M.M, Roujeinikova, A. | | Deposit date: | 2020-03-09 | | Release date: | 2020-05-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Structure-Activity Relationship Study Reveals the Molecular Basis for Specific Sensing of Hydrophobic Amino Acids by theCampylobacter jejuniChemoreceptor Tlp3.
Biomolecules, 10, 2020
|
|
