8WCR
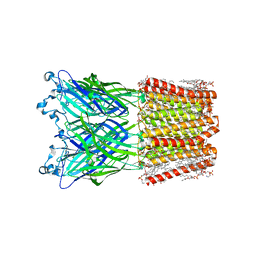 
 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | 著者 | Bharambe, N, Li, Z, Basak, S. | | 登録日 | 2023-09-13 | | 公開日 | 2024-04-17 | | 実験手法 | ELECTRON MICROSCOPY (2.74 Å) | | 主引用文献 | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
8WCQ
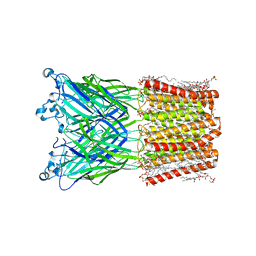 
 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | 著者 | Bharambe, N, Li, Z, Basak, S. | | 登録日 | 2023-09-13 | | 公開日 | 2024-04-17 | | 実験手法 | ELECTRON MICROSCOPY (3.35 Å) | | 主引用文献 | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
6QBO
 
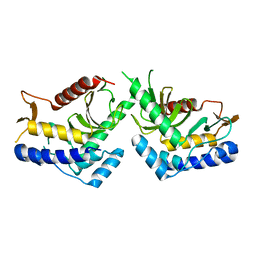 | | structure of the core domaine of Knr4, an intrinsically disordered protein from Saccharomyces cerevisiae - mutant S203A | | 分子名称: | Cell wall assembly regulator SMI1 | | 著者 | Ramos, N, Batista, M, Francois, J.M, Mourey, L, Maveyraud, L, Zerbib, D. | | 登録日 | 2018-12-21 | | 公開日 | 2020-01-29 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (2.75 Å) | | 主引用文献 | structure of the core domaine of Knr4, an intrinsically disordered protein from Saccharomyces cerevisiae - mutant S203A
To Be Published
|
|
6QBP
 
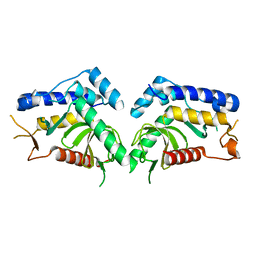 | | structure of the core domaine of Knr4, an intrinsically disordered protein from Saccharomyces cerevisiae - mutant S203D | | 分子名称: | Cell wall assembly regulator SMI1 | | 著者 | Ramos, N, Batista, M, Francois, J.M, Mourey, L, Maveyraud, L, Zerbib, D. | | 登録日 | 2018-12-21 | | 公開日 | 2020-01-29 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | structure of the core domaine of Knr4, an intrinsically disordered protein from Saccharomyces cerevisiae - mutant S203D
To Be Published
|
|
8I47
 
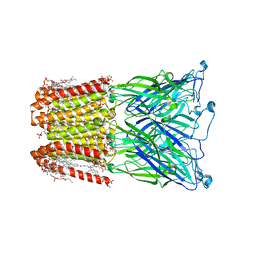 | |
8I42
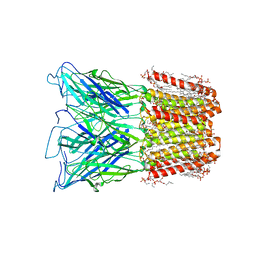 
 | |
8I41
 
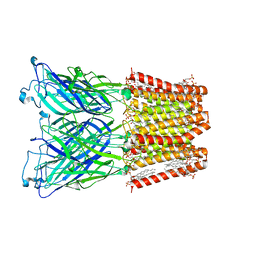 | |
8I48
 
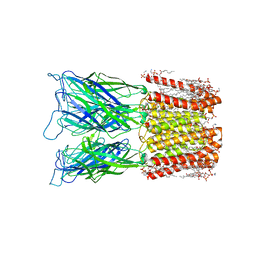 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | 著者 | Bharambe, N, Li, Z, Basak, S. | | 登録日 | 2023-01-18 | | 公開日 | 2024-04-17 | | 実験手法 | ELECTRON MICROSCOPY (2.74 Å) | | 主引用文献 | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
8JJ3
 
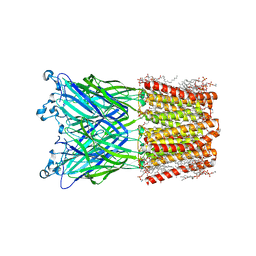 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5 | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | 著者 | Bharambe, N, Li, Z, Basak, S. | | 登録日 | 2023-05-29 | | 公開日 | 2024-04-17 | | 実験手法 | ELECTRON MICROSCOPY (2.6476 Å) | | 主引用文献 | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
4WP9
 
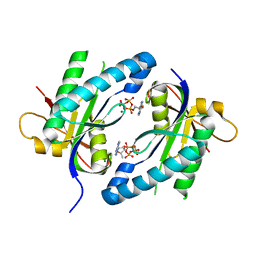 | | Crystal structure of Adenylyl cyclase MA1120 from Mycobacterium Avium bound to 2'5'-DD-3'-ATP, Calcium and Magnesium ion | | 分子名称: | 2',5'-dideoxyadenosine 3'-(tetrahydrogen triphosphate), CALCIUM ION, MAGNESIUM ION, ... | | 著者 | Bharambe, N.G, Barathy, D.V, Suguna, K. | | 登録日 | 2014-10-17 | | 公開日 | 2015-09-02 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.382 Å) | | 主引用文献 | Autoinhibitory mechanism and activity-related structural changes in a mycobacterial adenylyl cyclase
J.Struct.Biol., 190, 2015
|
|
4WPA
 
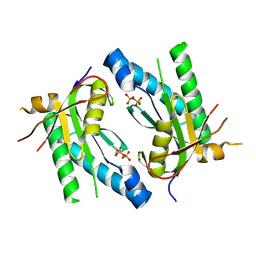 | |
5D0G
 
 | |
5D15
 
 | | Crystal structure of an adenylyl cyclase Ma1120 from Mycobacterium avium in complex with ATP and calcium ion | | 分子名称: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | 著者 | Bharambe, N.G, Barathy, D.V, Suguna, K. | | 登録日 | 2015-08-03 | | 公開日 | 2016-08-10 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Substrate specificity determinants of class III nucleotidyl cyclases
Febs J., 283, 2016
|
|
5D0E
 
 | | Crystal Structure of an adenylyl cyclase Ma1120-Cat in complex with GTP and calcium from Mycobacterium avium | | 分子名称: | CALCIUM ION, CHLORIDE ION, Cyclase, ... | | 著者 | Bharambe, N.G, Barathy, D.V, Suguna, K. | | 登録日 | 2015-08-03 | | 公開日 | 2016-08-10 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Substrate specificity determinants of class III nucleotidyl cyclases
Febs J., 283, 2016
|
|
5D0H
 
 | |
1Q4N
 
 | |
1SMD
 
 | | HUMAN SALIVARY AMYLASE | | 分子名称: | AMYLASE, CALCIUM ION, CHLORIDE ION | | 著者 | Ramasubbu, N. | | 登録日 | 1996-01-24 | | 公開日 | 1996-07-11 | | 最終更新日 | 2019-12-25 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Structure of human salivary alpha-amylase at 1.6 A resolution: implications for its role in the oral cavity.
Acta Crystallogr.,Sect.D, 52, 1996
|
|
1YHT
 
 | | Crystal structure analysis of Dispersin B | | 分子名称: | ACETIC ACID, DspB, GLYCEROL | | 著者 | Ramasubbu, N, Thomas, L.M, Ragunath, C, Kaplan, J.B. | | 登録日 | 2005-01-10 | | 公開日 | 2006-01-10 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural Analysis of Dispersin B, a Biofilm-releasing Glycoside Hydrolase from the Periodontopathogen Actinobacillus actinomycetemcomitans.
J.Mol.Biol., 349, 2005
|
|
1MFU
 
 | | Probing the role of a mobile loop in human salivary amylase: Structural studies on the loop-deleted mutant | | 分子名称: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, ... | | 著者 | Ramasubbu, N, Ragunath, C, Mishra, P.J. | | 登録日 | 2002-08-13 | | 公開日 | 2002-11-20 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Probing the role of a mobile loop in substrate binding and enzyme activity of
human salivary amylase.
J.Mol.Biol., 325, 2003
|
|
3BLP
 
 | | Role of aromatic residues in human salivary alpha-amylase | | 分子名称: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, Alpha-amylase 1, ... | | 著者 | Ramasubbu, N. | | 登録日 | 2007-12-11 | | 公開日 | 2008-11-18 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Structure-function relationships in human salivary alpha-amylase: role of aromatic residues in a secondary binding site
Biologia, 63, 2008
|
|
1NM9
 
 | | Crystal structure of recombinant human salivary amylase mutant W58A | | 分子名称: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, Alpha-amylase, ... | | 著者 | Ramasubbu, N, Ragunath, C, Mishra, P.J, Thomas, L.M. | | 登録日 | 2003-01-09 | | 公開日 | 2004-01-20 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Human salivary alpha-amylase Trp58 situated at subsite -2 is critical for enzyme activity.
Eur.J.Biochem., 271, 2004
|
|
3BLK
 
 | | Role of aromatic residues in starch binding | | 分子名称: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, Alpha-amylase 1, ... | | 著者 | Ramasubbu, N. | | 登録日 | 2007-12-11 | | 公開日 | 2008-11-25 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structure-function relationships in human salivary alpha-amylase: role of aromatic residues in a secondary binding site
Biologia, 63, 2008
|
|
1MFV
 
 | | Probing the role of a mobile loop in human slaivary amylase: Structural studies on the loop-deleted enzyme | | 分子名称: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, ... | | 著者 | Ramasubbu, N, Ragunath, C, Mishra, P.J. | | 登録日 | 2002-08-13 | | 公開日 | 2002-11-20 | | 最終更新日 | 2020-07-29 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Probing the role of a mobile loop in substrate binding and enzyme activity of
human salivary amylase.
J.Mol.Biol., 325, 2003
|
|
1C8Q
 
 | |
1AQP
 
 | | RIBONUCLEASE A COPPER COMPLEX | | 分子名称: | COPPER (II) ION, RIBONUCLEASE A | | 著者 | Ramasubbu, N. | | 登録日 | 1997-07-31 | | 公開日 | 1998-05-27 | | 最終更新日 | 2023-08-02 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Crystal structures of the copper and nickel complexes of RNase A: metal-induced interprotein interactions and identification of a novel copper binding motif.
Proc.Natl.Acad.Sci.USA, 94, 1997
|
|
