8WCJ
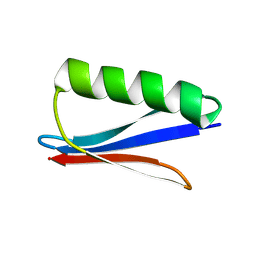 
 | | Crystal structure of GB3 penta mutation L5V/K10H/T16S/K19E/Y33I | | Descriptor: | Immunoglobulin G-binding protein G | | Authors: | Qin, M.M, Chen, X.X, Zhang, X.Y, Song, X.F, Yao, L.S. | | Deposit date: | 2023-09-12 | | Release date: | 2024-07-31 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Protein Allostery Study in Cells Using NMR Spectroscopy.
Anal.Chem., 96, 2024
|
|
5Z2U
 
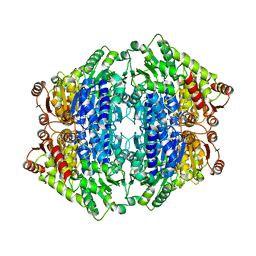 | | ThDP-Mn2+ complex of R395A variant of EcMenD soaked with 2-ketoglutarate for 5 min | | Descriptor: | (4S)-4-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3lambda~5~-thiazol-2-yl}-4-hydroxybutanoic acid, 1,2-ETHANEDIOL, 2-succinyl-5-enolpyruvyl-6-hydroxy-3-cyclohexene-1-carboxylate synthase, ... | | Authors: | Qin, M.M, Guo, Z.H. | | Deposit date: | 2018-01-04 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Two active site arginines are critical determinants of substrate binding and catalysis in MenD: a thiamine-dependent enzyme in menaquinone biosynthesis.
Biochem. J., 475, 2018
|
|
5Z2R
 
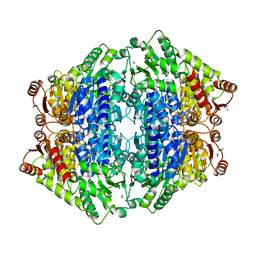 | | ThDP-Mn2+ complex of R395K variant of EcMenD soaked with 2-ketoglutarate for 5 min | | Descriptor: | (4S)-4-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3lambda~5~-thiazol-2-yl}-4-hydroxybutanoic acid, 2-succinyl-5-enolpyruvyl-6-hydroxy-3-cyclohexene-1-carboxylate synthase, FORMIC ACID, ... | | Authors: | Qin, M.M, Guo, Z.H. | | Deposit date: | 2018-01-03 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Two active site arginines are critical determinants of substrate binding and catalysis in MenD: a thiamine-dependent enzyme in menaquinone biosynthesis.
Biochem. J., 475, 2018
|
|
5Z2P
 
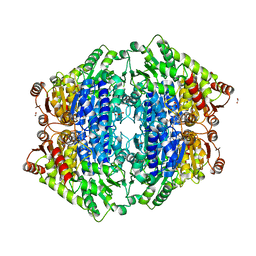 | | ThDP-Mn2+ complex of R413K variant of EcMenD soaked with 2-ketoglutarate for 5 min | | Descriptor: | (4S)-4-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3lambda~5~-thiazol-2-yl}-4-hydroxybutanoic acid, 2-succinyl-5-enolpyruvyl-6-hydroxy-3-cyclohexene-1-carboxylate synthase, FORMIC ACID, ... | | Authors: | Qin, M.M, Guo, Z.H. | | Deposit date: | 2018-01-03 | | Release date: | 2018-11-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Two active site arginines are critical determinants of substrate binding and catalysis in MenD: a thiamine-dependent enzyme in menaquinone biosynthesis.
Biochem. J., 475, 2018
|
|
8I3O
 
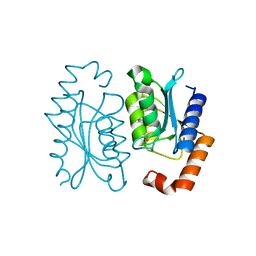 | |
8I3N
 
 | |
8I3P
 
 | | crystal structure of yeast cytosine deaminase mutant yCD-RQ-1/8SAH in complex with (R)-4-hydroxy-3,4-dihydropyrimidin-2(1H)-one | | Descriptor: | 1,2-ETHANEDIOL, 4-oxidanyl-3,4-dihydro-1H-pyrimidin-2-one, Cytosine deaminase, ... | | Authors: | Qin, M.M, Deng, H.Z, Yao, L.S. | | Deposit date: | 2023-01-17 | | Release date: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | crystal structure of yeast cytosine deaminase mutant yCD-RQ-1/8SAH in complex with (R)-4-hydroxy-3,4-dihydropyrimidin-2(1H)-one
To be published
|
|
