1QGB
 
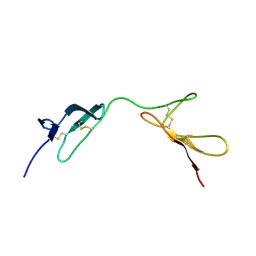 | | SOLUTION STRUCTURE OF THE N-TERMINAL F1 MODULE PAIR FROM HUMAN FIBRONECTIN | | Descriptor: | PROTEIN (FIBRONECTIN) | | Authors: | Potts, J.R, Bright, J.R, Bolton, D, Pickford, A.R, Campbell, I.D. | | Deposit date: | 1999-04-21 | | Release date: | 1999-12-08 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the N-terminal F1 module pair from human fibronectin.
Biochemistry, 38, 1999
|
|
7SIE
 
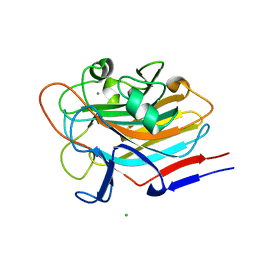 | | Structure of AAP A-domain (residues 351-605) from Staphylococcus epidermidis | | Descriptor: | Accumulation associated protein, CALCIUM ION, CHLORIDE ION | | Authors: | Atkin, K.E, Brentnall, A.S, Dodson, E.J, Whelan, F, Clark, L, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2021-10-13 | | Release date: | 2022-10-19 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Staphylococcal Periscope proteins Aap, SasG, and Pls project noncanonical legume-like lectin adhesin domains from the bacterial surface.
J.Biol.Chem., 299, 2023
|
|
6S5X
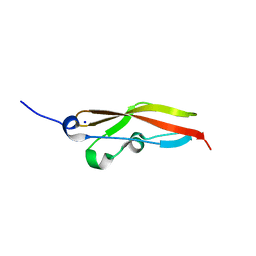 
 | | Structure of RibR, the most N-terminal Rib domain from Group B Streptococcus species Streptococcus agalactiae | | Descriptor: | Group B streptococcal R4 surface protein, SODIUM ION | | Authors: | Whelan, F, Turkenburg, J.P, Griffiths, S.C, Bateman, A, Potts, J.R. | | Deposit date: | 2019-07-02 | | Release date: | 2019-12-11 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Defining the remarkable structural malleability of a bacterial surface protein Rib domain implicated in infection.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6S5Y
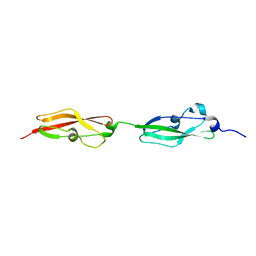 
 | |
8DEO
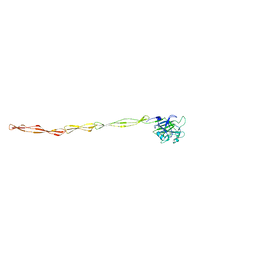 
 | | Structure of AAP A domain and B-repeats (residues 351-813) from Staphylococcus epidermidis | | Descriptor: | Accumulation associated protein, CALCIUM ION, CHLORIDE ION | | Authors: | Harris, G, Whelan, F, Clark, L, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2022-06-21 | | Release date: | 2023-05-03 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Staphylococcal Periscope proteins Aap, SasG, and Pls project noncanonical legume-like lectin adhesin domains from the bacterial surface.
J.Biol.Chem., 299, 2023
|
|
7AVH
 
 | | Streptococcal High Identity Repeats in Tandem (SHIRT) domains 3-4 from cell surface protein SGO_0707 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, LPXTG cell wall surface protein | | Authors: | Whelan, F, Jenkins, H.T, Potts, J.R. | | Deposit date: | 2020-11-05 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Periscope Proteins are variable-length regulators of bacterial cell surface interactions.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7AVK
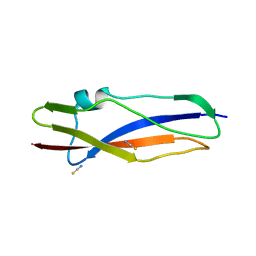 
 | | Streptococcal High Identity Repeats in Tandem (SHIRT) domain 10 from cell surface protein SGO_0707 | | Descriptor: | LPXTG cell wall surface protein, isothiocyanate | | Authors: | Degut, C, Gilburt, J, Whelan, F, Jenkins, H.T, Potts, J.R. | | Deposit date: | 2020-11-05 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (0.82 Å) | | Cite: | Periscope Proteins are variable-length regulators of bacterial cell surface interactions.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7AVJ
 | |
1HKY
 
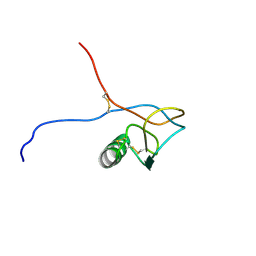 | | Solution structure of a PAN module from Eimeria tenella | | Descriptor: | MICRONEME PROTEIN 5 PRECURSOR | | Authors: | Brown, P.J, Mulvey, D, Potts, J.R, Tomley, F.M, Campbell, I.D. | | Deposit date: | 2002-10-03 | | Release date: | 2002-10-17 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a Pan Module from the Apicomplexan Parasite Eimeria Tenella
J.Struct.Funct.Genom., 4, 2003
|
|
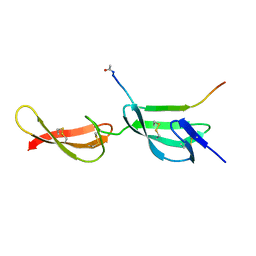
4PZ5
 
 | |
5DBL
 | |
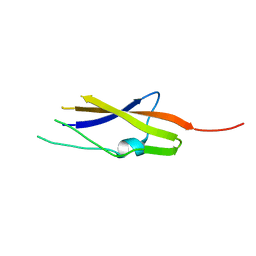
3TIQ
 | | Crystal structure of Staphylococcus aureus SasG G51-E-G52 module | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, MAGNESIUM ION, Surface protein G | | Authors: | Gruszka, D.T, Wojdyla, J.A, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2011-08-21 | | Release date: | 2012-04-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8739 Å) | | Cite: | Staphylococcal biofilm-forming protein has a contiguous rod-like structure.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
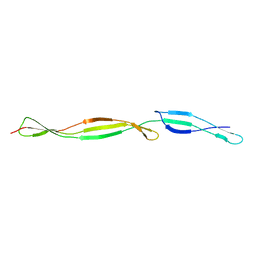
3TIP
 
 | |
7SJK
 
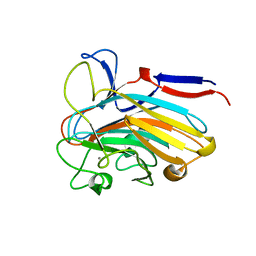 | | Structure of PLS A-domain (residues 391-656) from Staphylococcus aureus | | Descriptor: | CALCIUM ION, Pls Plasmin sensitive surface protein | | Authors: | Clark, L, Whelan, F, Atkin, K.E, Brentnall, A.S, Dodson, E.J, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2021-10-18 | | Release date: | 2022-10-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.208 Å) | | Cite: | Staphylococcal Periscope proteins Aap, SasG, and Pls project noncanonical legume-like lectin adhesin domains from the bacterial surface.
J.Biol.Chem., 299, 2023
|
|
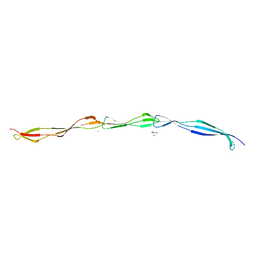
7SMH
 
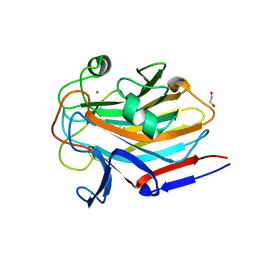 | | Structure of SASG A-domain (residues 163-419) from Staphylococcus aureus | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Surface protein G | | Authors: | Atkin, K.E, Whelan, F, Brentnall, A.S, Dodson, E.J, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2021-10-25 | | Release date: | 2022-11-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Staphylococcal Periscope proteins Aap, SasG, and Pls project noncanonical legume-like lectin adhesin domains from the bacterial surface.
J.Biol.Chem., 299, 2023
|
|
5FRH
 
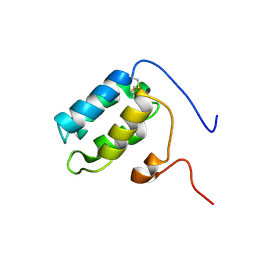 | | Solution structure of oxidised RsrA | | Descriptor: | ANTI-SIGMA FACTOR RSRA | | Authors: | Zdanowski, K, Pecqueur, L, Werner, J, Potts, J.R, Kleanthous, C. | | Deposit date: | 2015-12-17 | | Release date: | 2016-08-03 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | The Anti-Sigma Factor Rsra Responds to Oxidative Stress by Reburying its Hydrophobic Core.
Nat.Commun., 7, 2016
|
|
7SP2
 
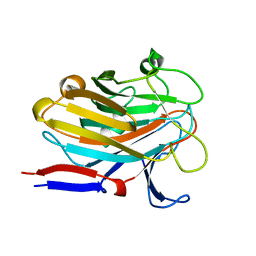 | | Structure of PLS A-domain (residues 391-656; 513-518 deletion mutant) from Staphylococcus aureus | | Descriptor: | CALCIUM ION, Plasmin Sensitive Protein Pls | | Authors: | Clark, L, Whelan, F, Atkin, K.E, Brentnall, A.S, Dodson, E.J, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2021-11-02 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structure of PLS A-domain (residues 391-65) from Staphylococcus aureus
Not Published
|
|
4WVE
 
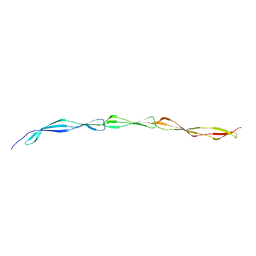 | |
5FRF
 
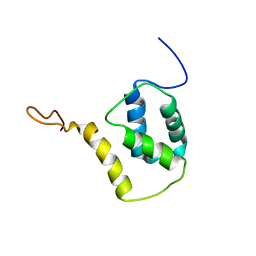 | | Solution structure of reduced and zinc-bound RsrA | | Descriptor: | ANTI-SIGMA FACTOR RSRA, ZINC ION | | Authors: | Zdanowski, K, Pecqueur, L, Werner, J, Potts, J.R, Kleanthous, C. | | Deposit date: | 2015-12-17 | | Release date: | 2016-08-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Anti-Sigma Factor Rsra Responds to Oxidative Stress by Reburying its Hydrophobic Core.
Nat.Commun., 7, 2016
|
|
3ZRZ
 
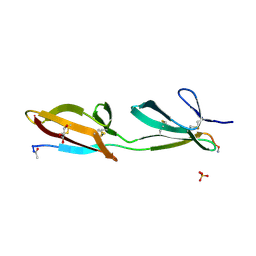 | | Crystal structure of the second and third fibronectin F1 modules in complex with a fragment of Streptococcus pyogenes SfbI-5 | | Descriptor: | FIBRONECTIN, FIBRONECTIN-BINDING PROTEIN, GLYCEROL, ... | | Authors: | Norris, N.C, Bingham, R.J, Potts, J.R. | | Deposit date: | 2011-06-21 | | Release date: | 2011-09-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and Functional Analysis of the Tandem Beta-Zipper Interaction of a Streptococcal Protein with Human Fibronectin.
J.Biol.Chem., 286, 2011
|
|
4B5Z
 
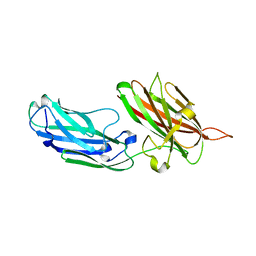 | | Structure of the apo-rFnBPA(189-505) from Staphylococcus aureus | | Descriptor: | FIBRONECTIN-BINDING PROTEIN A | | Authors: | Stemberk, V, Moroz, O, Atkin, K.E, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2012-08-08 | | Release date: | 2013-10-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Evidence for Steric Regulation of Fibrinogen Binding to Staphylococcus Aureus Fibronectin-Binding Protein a (Fnbpa).
J.Biol.Chem., 289, 2014
|
|
4B60
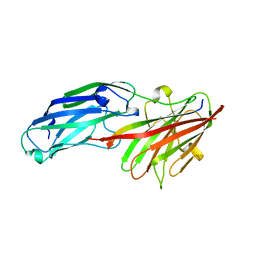 
 | | Structure of rFnBPA(189-505) in complex with fibrinogen gamma chain C- terminal peptide | | Descriptor: | CALCIUM ION, FIBRINOGEN GAMMA CHAIN, FIBRONECTIN-BINDING PROTEIN A | | Authors: | Stemberk, V, Moroz, O, Atkin, K.E, Turkenburg, J.P, Potts, J.R. | | Deposit date: | 2012-08-08 | | Release date: | 2013-10-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Evidence for Steric Regulation of Fibrinogen Binding to Staphylococcus Aureus Fibronectin-Binding Protein a (Fnbpa).
J.Biol.Chem., 289, 2014
|
|
2JKE
 
 | | Structure of a family 97 alpha-glucosidase from Bacteroides thetaiotaomicron in complex with deoxynojirimycin | | Descriptor: | 1,2-ETHANEDIOL, 1-DEOXYNOJIRIMYCIN, ALPHA-GLUCOSIDASE (ALPHA-GLUCOSIDASE SUSB), ... | | Authors: | Gloster, T.M, Turkenburg, J.P, Potts, J.R, Henrissat, B, Davies, G.J. | | Deposit date: | 2008-08-28 | | Release date: | 2008-09-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Divergence of Catalytic Mechanism within a Glycosidase Family Provides Insight Into Evolution of Carbohydrate Metabolism by Human Gut Flora.
Chem.Biol., 15, 2008
|
|
2JKP
 
 | | Structure of a family 97 alpha-glucosidase from Bacteroides thetaiotaomicron in complex with castanospermine | | Descriptor: | 1,2-ETHANEDIOL, ALPHA-GLUCOSIDASE (ALPHA-GLUCOSIDASE SUSB), CALCIUM ION, ... | | Authors: | Gloster, T.M, Turkenburg, J.P, Potts, J.R, Henrissat, B, Davies, G.J. | | Deposit date: | 2008-08-29 | | Release date: | 2008-09-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Divergence of Catalytic Mechanism within a Glycosidase Family Provides Insight Into Evolution of Carbohydrate Metabolism by Human Gut Flora.
Chem.Biol., 15, 2008
|
|
2JKA
 
 | | Native structure of a family 97 alpha-glucosidase from Bacteroides thetaiotaomicron | | Descriptor: | 1,2-ETHANEDIOL, ALPHA-GLUCOSIDASE (ALPHA-GLUCOSIDASE SUSB), CALCIUM ION | | Authors: | Gloster, T.M, Turkenburg, J.P, Potts, J.R, Henrissat, B, Davies, G.J. | | Deposit date: | 2008-08-23 | | Release date: | 2008-09-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Divergence of Catalytic Mechanism within a Glycosidase Family Provides Insight Into Evolution of Carbohydrate Metabolism by Human Gut Flora.
Chem.Biol., 15, 2008
|
|
