7LZA
 
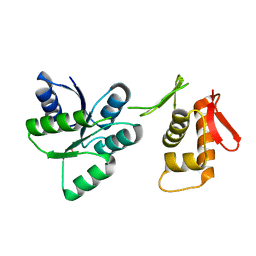 | | Activated form of VanR from S. coelicolor | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, MAGNESIUM ION, Putative two-component system response regulator | | Authors: | Maciunas, L.J, Loll, P.J. | | Deposit date: | 2021-03-09 | | Release date: | 2021-07-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Structures of full-length VanR from Streptomyces coelicolor in both the inactive and activated states.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7LZ9
 
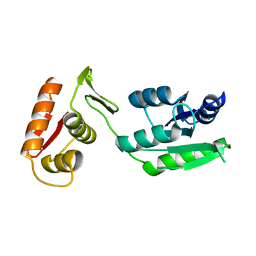 | | Inactive form of VanR from S. coelicolor | | Descriptor: | MAGNESIUM ION, Putative two-component system response regulator | | Authors: | Maciunas, L.J, Loll, P.J. | | Deposit date: | 2021-03-09 | | Release date: | 2021-07-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of full-length VanR from Streptomyces coelicolor in both the inactive and activated states.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
4BH1
 
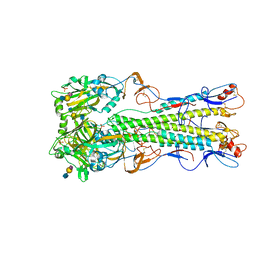 | | H5 (tyTy) Influenza Virus Haemagglutinin in Complex with Avian Receptor Analogue 3'-SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, HEMAGGLUTININ, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Xiong, X, Coombs, P.J, Martin, S.R, Liu, J, Xiao, H, McCauley, J.W, Locher, K, Walker, P.A, Collins, P.J, Kawaoka, Y, Skehel, J.J, Gamblin, S.J. | | Deposit date: | 2013-03-29 | | Release date: | 2013-04-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Receptor Binding by a Ferret-Transmissible H5 Avian Influenza Virus
Nature, 497, 2013
|
|
4BH3
 
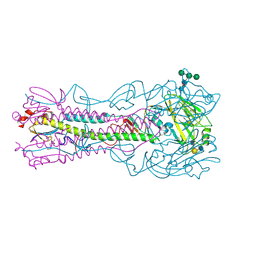 | | Haemagglutinin from a Transmissible Mutant H5 Influenza Virus in Complex with Human Receptor Analogue 6'-SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, HEMAGGLUTININ, ... | | Authors: | Xiong, X, Coombs, P.J, Martin, S.R, Liu, J, Xiao, H, McCauley, J.W, Locher, K, Walker, P.A, Collins, P.J, Kawaoka, Y, Skehel, J.J, Gamblin, S.J. | | Deposit date: | 2013-03-29 | | Release date: | 2013-04-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Receptor Binding by a Ferret-Transmissible H5 Avian Influenza Virus
Nature, 497, 2013
|
|
4BH4
 
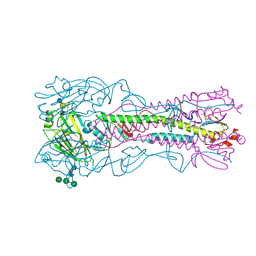 | | Haemagglutinin from a Transmissible Mutant H5 Influenza Virus in Complex with Avian Receptor Analogue 3'-SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, HEMAGGLUTININ, ... | | Authors: | Xiong, X, Coombs, P.J, Martin, S.R, Liu, J, Xiao, H, McCauley, J.W, Locher, K, Walker, P.A, Collins, P.J, Kawaoka, Y, Skehel, J.J, Gamblin, S.J. | | Deposit date: | 2013-03-29 | | Release date: | 2013-04-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Receptor Binding by a Ferret-Transmissible H5 Avian Influenza Virus
Nature, 497, 2013
|
|
4BH2
 
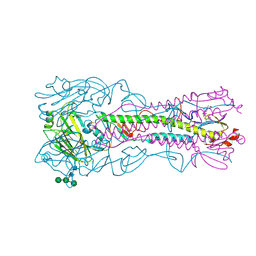 | | Crystal Structure of the Haemagglutinin from a Transmissible Mutant H5 Influenza Virus | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, HEMAGGLUTININ, ... | | Authors: | Xiong, X, Coombs, P.J, Martin, S.R, Liu, J, Xiao, H, McCauley, J.W, Locher, K, Walker, P.A, Collins, P.J, Kawaoka, Y, Skehel, J.J, Gamblin, S.J. | | Deposit date: | 2013-03-29 | | Release date: | 2013-04-24 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Receptor Binding by a Ferret-Transmissible H5 Avian Influenza Virus
Nature, 497, 2013
|
|
1YAZ
 
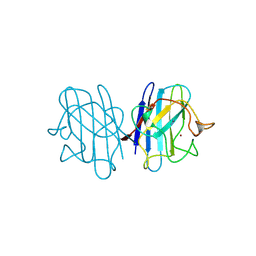 | | AZIDE-BOUND YEAST CU(II)/ZN SUPEROXIDE DISMUTASE ROOM TEMPERATURE (298K) STRUCTURE | | Descriptor: | AZIDE ION, COPPER (II) ION, PROTEIN (CU/ZN SUPEROXIDE DISMUTASE), ... | | Authors: | Hart, P.J, Balbirnie, M.M, Ogihara, N.L, Nersissian, A.M, Weiss, M.S, Valentine, J.S, Eisenberg, D. | | Deposit date: | 1998-12-23 | | Release date: | 2000-01-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A structure-based mechanism for copper-zinc superoxide dismutase.
Biochemistry, 38, 1999
|
|
1ZCT
 
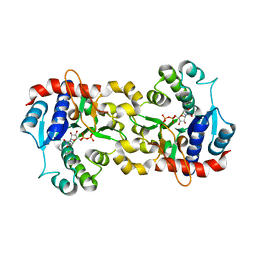 | | structure of glycogenin truncated at residue 270 in a complex with UDP | | Descriptor: | Glycogenin-1, MANGANESE (II) ION, URIDINE-5'-DIPHOSPHATE | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-13 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
1ZDF
 
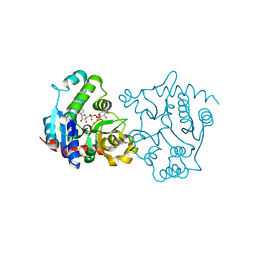 | | Ser162 mutant of glycogenin complexed with UDP-glucose and manganese | | Descriptor: | Glycogenin-1, MANGANESE (II) ION, SULFATE ION, ... | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-14 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
8SL0
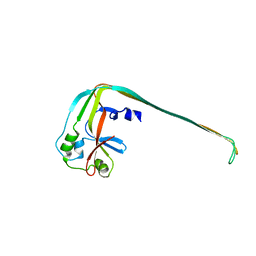 
 | |
1XC1
 
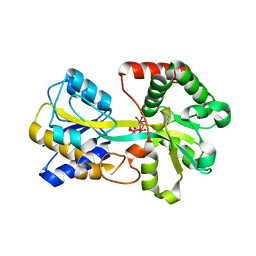 | | Oxo Zirconium(IV) Cluster in the Ferric Binding Protein (FBP) | | Descriptor: | OXO ZIRCONIUM(IV) CLUSTER, periplasmic iron-binding protein | | Authors: | Zhong, W, Alexeev, D, Harvey, I, Guo, M, Hunter, D.J.B, Zhu, H, Campopiano, D.J, Sadler, P.J. | | Deposit date: | 2004-08-31 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Assembly of an Oxo-Zirconium(IV) Cluster in a Protein Cleft
Angew.Chem.Int.Ed.Engl., 43, 2004
|
|
1ZCY
 
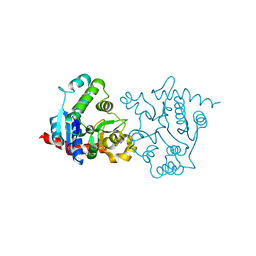 | | apo form of a mutant of glycogenin in which Asp159 is replaced by Ser | | Descriptor: | Glycogenin-1, SULFATE ION | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-13 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
1ZCU
 
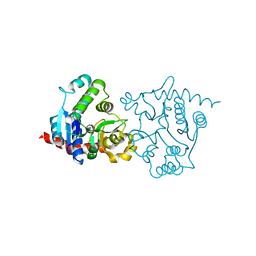 | | apo form of the 162S mutant of glycogenin | | Descriptor: | Glycogenin-1 | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-13 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
1Y7J
 
 | | NMR structure family of Human Agouti Signalling Protein (80-132: Q115Y, S124Y) | | Descriptor: | Agouti Signaling Protein | | Authors: | McNulty, J.C, Jackson, P.J, Thompson, D.A, Chai, B, Gantz, I, Barsh, G.S, Dawson, P.E, Millhauser, G.L. | | Deposit date: | 2004-12-08 | | Release date: | 2005-02-15 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Structures of the agouti signaling protein.
J.Mol.Biol., 346, 2005
|
|
5IR0
 
 | | Crystal structure of protein of unknown function ORF19 from Vibrio cholerae O1 PICI-like element, C57S I109M mutant | | Descriptor: | CITRIC ACID, Uncharacterized protein ORF19 | | Authors: | Stogios, P.J, Wawrzak, Z, Skarina, T, Di Leo, R, Yim, V, Savchenko, A, Anderson, W.F, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2016-03-11 | | Release date: | 2016-03-30 | | Method: | X-RAY DIFFRACTION (3.297 Å) | | Cite: | Crystal structure of protein of unknown function ORF19 from Vibrio cholerae O1 PICI-like element, C57S I109M mutant
To Be Published
|
|
1Y7K
 
 | | NMR structure family of Human Agouti Signalling Protein (80-132: Q115Y, S124Y) | | Descriptor: | Agouti Signaling Protein | | Authors: | McNulty, J.C, Jackson, P.J, Thompson, D.A, Chai, B, Gantz, I, Barsh, G.S, Dawson, P.E, Millhauser, G.L. | | Deposit date: | 2004-12-08 | | Release date: | 2005-02-15 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Structures of the agouti signaling protein.
J.Mol.Biol., 346, 2005
|
|
1ZDG
 
 | | Ser159 mutant of glycogenin complexed with UDP-glucose and manganese | | Descriptor: | Glycogenin-1, MANGANESE (II) ION, SULFATE ION, ... | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-14 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
1ZCV
 
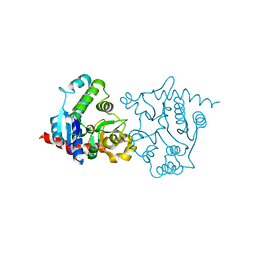 | | apo form of a mutant of glycogenin in which Asp159 is replaced by Asn | | Descriptor: | Glycogenin-1, SULFATE ION | | Authors: | Hurley, T.D, Stout, S.L, Miner, E, Zhou, J, Roach, P.J. | | Deposit date: | 2005-04-13 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Requirements for catalysis in mammalian glycogenin.
J.Biol.Chem., 280, 2005
|
|
1LEH
 
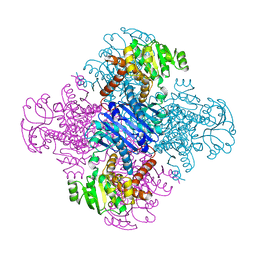 | | LEUCINE DEHYDROGENASE FROM BACILLUS SPHAERICUS | | Descriptor: | LEUCINE DEHYDROGENASE | | Authors: | Baker, P.J, Turnbull, A.P, Sedelnikova, S.E, Stillman, T.J, Rice, D.W. | | Deposit date: | 1995-06-09 | | Release date: | 1996-12-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A role for quaternary structure in the substrate specificity of leucine dehydrogenase.
Structure, 3, 1995
|
|
4CQQ
 
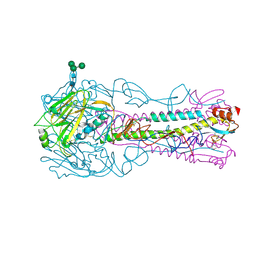 | | H5 (VN1194) Ser227Asn/Gln196Arg Mutant Haemagglutinin in Complex with Avian Receptor Analogue 3'SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, ... | | Authors: | Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Liu, J, Collins, P.J, Vachieri, S.G, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2014-02-21 | | Release date: | 2014-05-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
4CQY
 
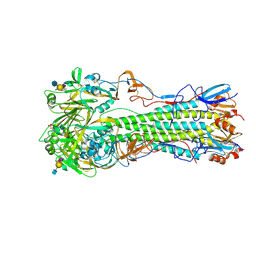 | | H5 (tyTy) Del133/Ile155Thr Mutant Haemagglutinin in Complex with Avian Receptor Analogue LSTa | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, HAEMAGGLUTININ HA1, HAEMAGGLUTININ HA2, ... | | Authors: | Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Liu, J, Collins, P.J, Vachieri, S.G, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2014-02-21 | | Release date: | 2014-05-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
4CQV
 
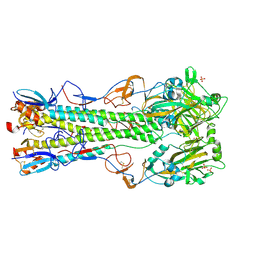 | | Crystal structure of H5 (tyTy) Del133/Ile155Thr Mutant Haemagglutinin | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, HAEMAGGLUTININ HA1, HAEMAGGLUTININ HA2, ... | | Authors: | Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Liu, J, Collins, P.J, Vachieri, S.G, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2014-02-21 | | Release date: | 2014-05-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
1ZUW
 
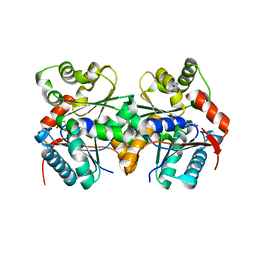 | | Crystal structure of B.subtilis glutamate racemase (RacE) with D-Glu | | Descriptor: | D-GLUTAMIC ACID, glutamate racemase 1 | | Authors: | Ruzheinikov, S.N, Taal, M.A, Sedelnikova, S.E, Baker, P.J, Rice, D.W. | | Deposit date: | 2005-06-01 | | Release date: | 2005-12-06 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Substrate-Induced Conformational Changes in Bacillus subtilis Glutamate Racemase and Their Implications for Drug Discovery
Structure, 13, 2005
|
|
4CQZ
 
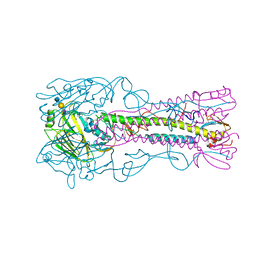 | | Crystal Structure of H5 (VN1194) Gln196Arg Mutant Haemagglutinin | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Liu, J, Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Collins, P.J, Vachieri, S.G, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2014-02-21 | | Release date: | 2014-05-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
4CR0
 
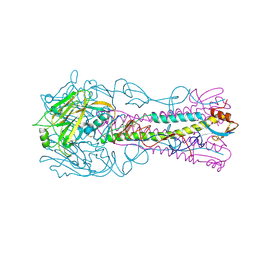 | | Crystal Structure of H5 (VN1194) Asn186Lys/Gly143Arg Mutant Haemagglutinin | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin HA1, Hemagglutinin HA2 | | Authors: | Collins, P.J, Vachieri, S.G, Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Liu, J, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2014-02-21 | | Release date: | 2014-05-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
