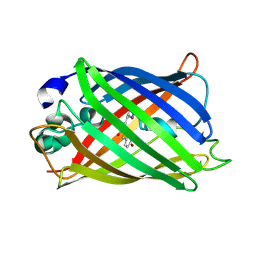
7A7Z
 | |
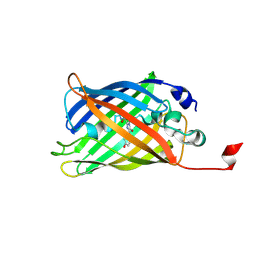
7A8L
 | |
2CTB
 
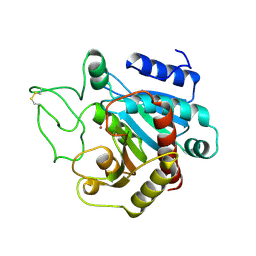 | | THE HIGH RESOLUTION CRYSTAL STRUCTURE OF THE COMPLEX BETWEEN CARBOXYPEPTIDASE A AND L-PHENYL LACTATE | | Descriptor: | CARBOXYPEPTIDASE A, ZINC ION | | Authors: | Teplyakov, A, Wilson, K.S, Orioli, P, Mangani, S. | | Deposit date: | 1993-04-02 | | Release date: | 1994-01-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | High-resolution structure of the complex between carboxypeptidase A and L-phenyl lactate.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
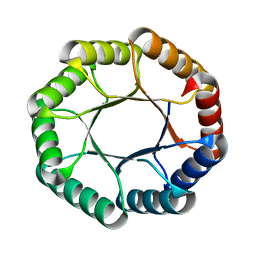
7UEK
 | |
1D7M
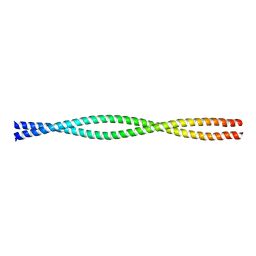 
 | | COILED-COIL DIMERIZATION DOMAIN FROM CORTEXILLIN I | | Descriptor: | CORTEXILLIN I | | Authors: | Burkhard, P, Kammerer, R.A, Steinmetz, M.O, Bourenkov, G.P, Aebi, U. | | Deposit date: | 1999-10-19 | | Release date: | 2000-03-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The coiled-coil trigger site of the rod domain of cortexillin I unveils a distinct network of interhelical and intrahelical salt bridges.
Structure Fold.Des., 8, 2000
|
|
7A8N
 
 | |
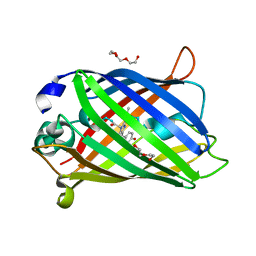
7A85
 | |
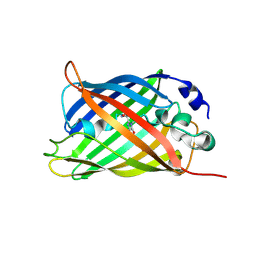
7A8E
 | |
7A8K
 
 | |
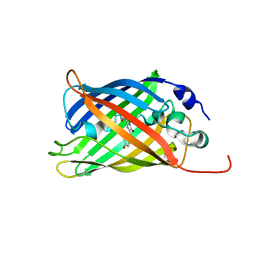
7A7O
 | |
2LRS
 
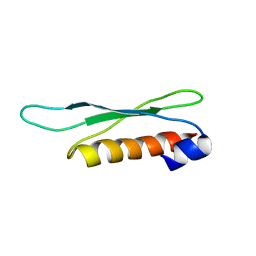 | | The second dsRBD domain from A. thaliana DICER-LIKE 1 | | Descriptor: | Endoribonuclease Dicer homolog 1 | | Authors: | Burdisso, P, Suarez, I, Bersch, B, Bologna, N, Palatnik, J, Boisbouvier, J, Rasia, R. | | Deposit date: | 2012-04-12 | | Release date: | 2013-01-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Second Double-Stranded RNA Binding Domain of Dicer-like Ribonuclease 1: Structural and Biochemical Characterization.
Biochemistry, 51, 2012
|
|
7A82
 
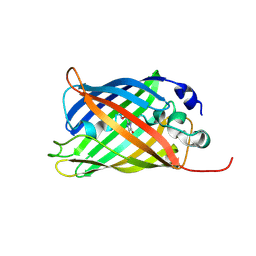 | |
7A87
 
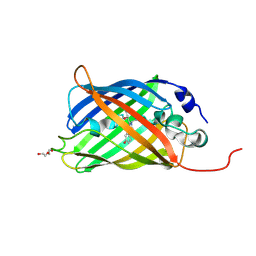 | |
7A8B
 | |
1MRN
 
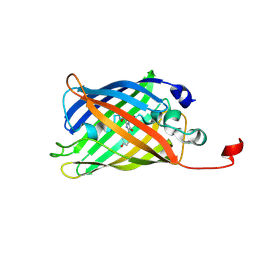
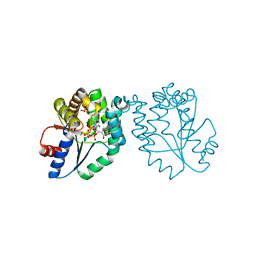 | | CRYSTAL STRUCTURE OF MYCOBACTERIUM TUBERCULOSIS THYMIDYLATE KINASE COMPLEXED WITH BISUBSTRATE INHIBITOR (TP5A) | | Descriptor: | MAGNESIUM ION, P1-(5'-ADENOSYL)P5-(5'-THYMIDYL)PENTAPHOSPHATE, SULFATE ION, ... | | Authors: | Haouz, A, Vanheusden, V, Munier-Lehmann, H, Froeyen, M, Herdewijn, P, Van Calenbergh, S, Delarue, M. | | Deposit date: | 2002-09-18 | | Release date: | 2003-01-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Enzymatic and structural analysis of inhibitors designed against Mycobacterium tuberculosis thymidylate kinase. New insights into the phosphoryl transfer mechanism.
J.Biol.Chem., 278, 2003
|
|
1MZV
 
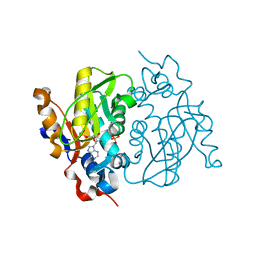 | | Crystal Structure of Adenine Phosphoribosyltransferase (APRT) From Leishmania tarentolae | | Descriptor: | ADENOSINE MONOPHOSPHATE, Adenine Phosphoribosyltransferase, PHOSPHATE ION | | Authors: | Thiemann, O.H, Silva, M, Oliva, G, Silva, C.H.T.P, Iulek, J. | | Deposit date: | 2002-10-10 | | Release date: | 2003-10-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of adenine phosphoribosyltransferase from Leishmania tarentolae: potential implications for APRT catalytic mechanism.
Biochim.Biophys.Acta, 1696, 2004
|
|
7A83
 | |
5OVW
 
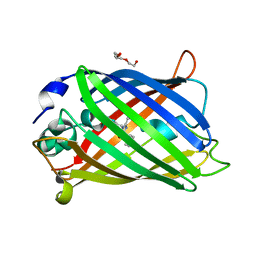
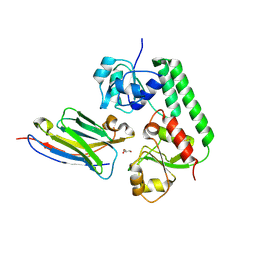 | | Nanobody-bound BtuF, the vitamin B12 binding protein in Escherichia coli | | Descriptor: | GLYCEROL, Nanobody, Vitamin B12-binding protein | | Authors: | Mireku, S.A, Sauer, M.M, Glockshuber, R, Locher, K.P. | | Deposit date: | 2017-08-30 | | Release date: | 2017-11-08 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.653 Å) | | Cite: | Structural basis of nanobody-mediated blocking of BtuF, the cognate substrate-binding protein of the Escherichia coli vitamin B12 transporter BtuCD.
Sci Rep, 7, 2017
|
|
2CYF
 
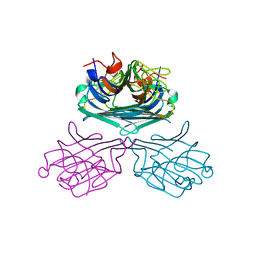 | | The Crystal Structure of Canavalia Maritima Lectin (ConM) in Complex with Trehalose and Maltose | | Descriptor: | CALCIUM ION, Lectin, MANGANESE (II) ION, ... | | Authors: | Delatorre, P, Rocha, B.A.M, Sousa, E.P, Gadelha, C.A.A, Azevedo Jr, W.F, Cavada, B.S. | | Deposit date: | 2005-07-06 | | Release date: | 2006-06-13 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a lectin from Canavalia maritima (ConM) in complex with trehalose and maltose reveals relevant mutation in ConA-like lectins
J.Struct.Biol., 154, 2006
|
|
7A8D
 | |
4CQ8
 
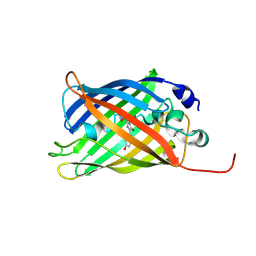
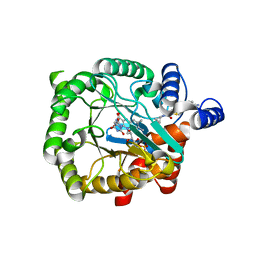 | | Plasmodium falciparum dihydroorotate dehydrogenase (DHODH) in complex with Genz-669178 | | Descriptor: | 5-(4-cyano-2-methyl-1H-benzimidazol-1-yl)-N-cyclopropylthiophene-2-carboxamide, DIHYDROOROTATE DEHYDROGENASE, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Rowland, P. | | Deposit date: | 2014-02-12 | | Release date: | 2014-04-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | In Vitro Resistance Selections for Plasmodium Falciparum Dihydroorotate Dehydrogenase Inhibitors Give Mutants with Multiple Point Mutations in the Drug-Binding Site and Altered Growth.
J.Biol.Chem., 289, 2014
|
|
7A8A
 
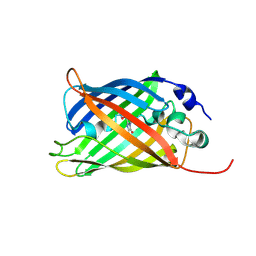 | |
7UOR
 
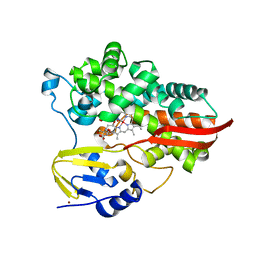 | | Crystal structure of cytochrome P450 enzyme CYP119 in complex with methyliridium(III) mesoporphyrin. | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Cytochrome, NICKEL (II) ION, ... | | Authors: | Pereira, J.H, Bloomer, B.J, Hartwig, J.F, Adams, P.D. | | Deposit date: | 2022-04-13 | | Release date: | 2023-02-22 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.16 Å) | | Cite: | Mechanistic and structural characterization of an iridium-containing cytochrome reveals kinetically relevant cofactor dynamics
Nat Catal, 6, 2023
|
|
7A8M
 
 | |
2GC3
 
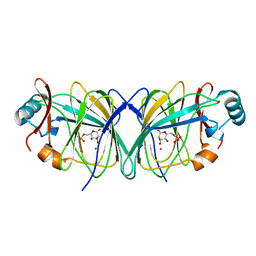 | |
