7TJC
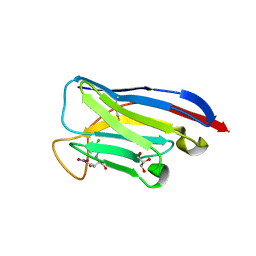 
 | | VHH Chl-B2 in complex with Chloramphenicol | | Descriptor: | CHLORAMPHENICOL, GLYCEROL, VHH-Chl-B2 | | Authors: | Nordeen, S.A, Schwartz, T.U. | | Deposit date: | 2022-01-16 | | Release date: | 2022-11-23 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structure and specificity of an anti-chloramphenicol single domain antibody for detection of amphenicol residues.
Protein Sci., 31, 2022
|
|
6X04
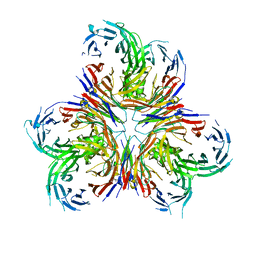 
 | |
6X02
 
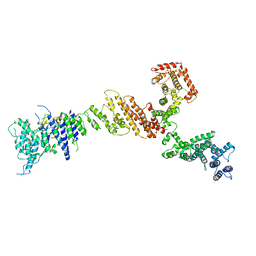 | |
6X03
 
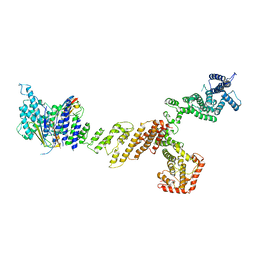 | |
6X08
 
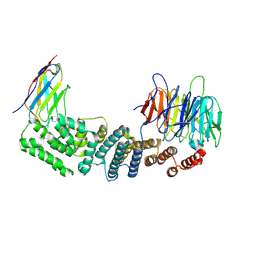 | | Nup85-Seh1 from S. cerevisiae bound by VHH-SAN2 | | Descriptor: | Nucleoporin NUP85, Nucleoporin SEH1, VHH-SAN2 | | Authors: | Nordeen, S.A, Schwartz, T.U. | | Deposit date: | 2020-05-15 | | Release date: | 2020-12-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (4.19 Å) | | Cite: | A nanobody suite for yeast scaffold nucleoporins provides details of the nuclear pore complex structure.
Nat Commun, 11, 2020
|
|
6X05
 
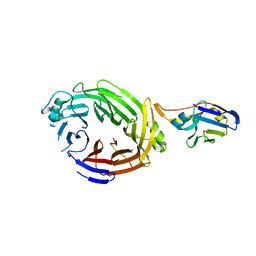 | |
6X07
 
 | |
6X06
 
 | |
