8IH0
 
 | | Crystal structure of GH11 from Thermoanaerobacterium saccharolyticum | | Descriptor: | ACETATE ION, Endo-1,4-beta-xylanase | | Authors: | Nam, K.H. | | Deposit date: | 2023-02-22 | | Release date: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Characterization and structural analysis of the endo-1,4-beta-xylanase GH11 from the hemicellulose-degrading Thermoanaerobacterium saccharolyticum useful for lignocellulose saccharification.
Sci Rep, 13, 2023
|
|
8IH1
 
 | |
8HVF
 
 | | Crystal structure of Thaumatin (100 ms) | | Descriptor: | 1,2-ETHANEDIOL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Nam, K.H. | | Deposit date: | 2022-12-26 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of Thaumatin (100 ms)
To Be Published
|
|
8HVE
 
 | | Crystal structure of Thaumatin (1 s) | | Descriptor: | 1,2-ETHANEDIOL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Nam, K.H. | | Deposit date: | 2022-12-26 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of Thaumatin (1 s)
To Be Published
|
|
8WFT
 
 | |
8WFU
 
 | |
8YUD
 
 | |
7WKR
 
 | |
7WUC
 
 | |
8GMW
 
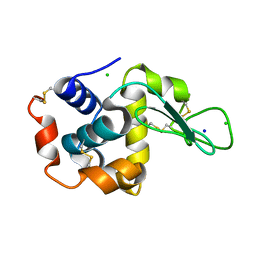 | | Crystal structure of lysozyme | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION | | Authors: | Nam, K.H. | | Deposit date: | 2022-08-22 | | Release date: | 2022-09-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structure of lysozyme
To Be Published
|
|
8GMV
 
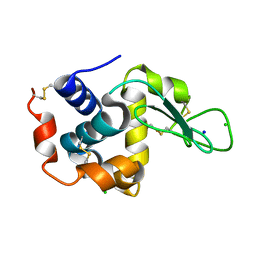 | | Crystal structure of lysozyme | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION | | Authors: | Nam, K.H. | | Deposit date: | 2022-08-22 | | Release date: | 2022-09-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of lysozyme
To Be Published
|
|
8WDG
 
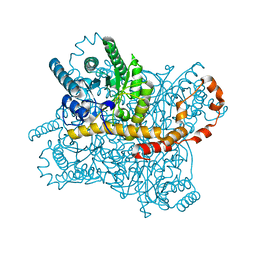 | |
8WGK
 
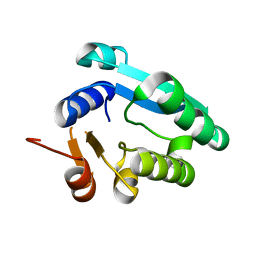 | |
8WFW
 
 | |
8WDI
 
 | |
8WXO
 
 | |
8WGL
 
 | | Crystal structure of Rhodothermus marinus substrate-binding protein (Hg soaking) | | Descriptor: | ABC-type uncharacterized transport system periplasmic component-like protein, MERCURY (II) ION | | Authors: | Nam, K.H. | | Deposit date: | 2023-09-22 | | Release date: | 2023-10-04 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and bioinformatics analysis of single-domain substrate-binding protein from Rhodothermus marinus.
Biochem Biophys Rep, 37, 2024
|
|
8X1D
 
 | |
8WFV
 
 | |
8WXM
 
 | |
8WXN
 
 | |
8WXP
 
 | |
8XPE
 
 | | Crystal structure of Tris-bound TsaBgl (DATA III) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SODIUM ION, beta-glucosidase | | Authors: | Nam, K.H. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-31 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural analysis of Tris binding in beta-glucosidases.
Biochem.Biophys.Res.Commun., 700, 2024
|
|
8XPC
 
 | | Crystal structure of Tris-bound TsaBgl (DATA I) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SODIUM ION, beta-glucosidase | | Authors: | Nam, K.H. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-31 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural analysis of Tris binding in beta-glucosidases.
Biochem.Biophys.Res.Commun., 700, 2024
|
|
8XPD
 
 | | Crystal structure of Tris-bound TsaBgl (DATA II) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SODIUM ION, beta-glucosidase | | Authors: | Nam, K.H. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-31 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural analysis of Tris binding in beta-glucosidases.
Biochem.Biophys.Res.Commun., 700, 2024
|
|
