4I3U
 | |
4I3W
 
 | |
4I3T
 
 | |
4I3X
 
 | |
4OK2
 
 | | Crystal Structure of Alg17c Mutant Y258A | | Descriptor: | Putative alginate lyase, ZINC ION | | Authors: | Nair, S.K, Park, D.S. | | Deposit date: | 2014-01-21 | | Release date: | 2014-01-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structure of a PL17 Family Alginate Lyase Demonstrates Functional Similarities among Exotype Depolymerases.
J.Biol.Chem., 289, 2014
|
|
4OJZ
 
 | |
4OK4
 
 | | Crystal Structure of Alg17c Mutant H202L | | Descriptor: | Putative alginate lyase, ZINC ION | | Authors: | Nair, S.K, Park, D.S. | | Deposit date: | 2014-01-21 | | Release date: | 2014-01-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of a PL17 Family Alginate Lyase Demonstrates Functional Similarities among Exotype Depolymerases.
J.Biol.Chem., 289, 2014
|
|
4H6W
 
 | |
4H6X
 
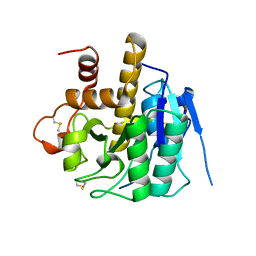
 | | Structure of Patellamide maturation protease PatG | | Descriptor: | Thiazoline oxidase/subtilisin-like protease | | Authors: | Nair, S.K, Agarwal, V. | | Deposit date: | 2012-09-19 | | Release date: | 2012-10-03 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of cyanobactin maturation enzymes define a family of transamidating proteases.
Chem.Biol., 19, 2012
|
|
4H6V
 
 | |
4NU6
 
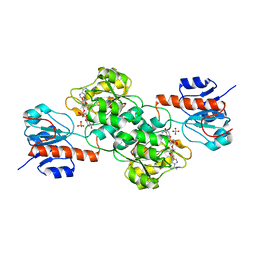 | | Crystal Structure of PTDH R301K | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Phosphonate dehydrogenase, SULFATE ION | | Authors: | Nair, S.K, Chekan, J.R. | | Deposit date: | 2013-12-03 | | Release date: | 2014-03-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Chemical rescue and inhibition studies to determine the role of arg301 in phosphite dehydrogenase.
Plos One, 9, 2014
|
|
4NU5
 
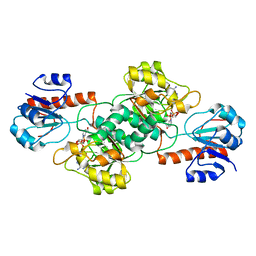 | | Crystal Structure of PTDH R301A | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Phosphonate dehydrogenase | | Authors: | Nair, S.K, Chekan, J.R. | | Deposit date: | 2013-12-03 | | Release date: | 2014-03-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Chemical rescue and inhibition studies to determine the role of arg301 in phosphite dehydrogenase.
Plos One, 9, 2014
|
|
3SJM
 
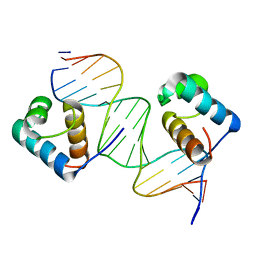 | | Crystal Structure Analysis of TRF2-Dbd-DNA complex | | Descriptor: | DNA (5'-D(*CP*TP*CP*TP*AP*GP*GP*GP*TP*TP*AP*GP*GP*GP*TP*TP*AP*G)-3'), DNA (5'-D(*TP*CP*TP*AP*AP*CP*CP*CP*TP*AP*AP*CP*CP*CP*TP*AP*GP*A)-3'), Telomeric repeat-binding factor 2 | | Authors: | Nair, S.K, Sliverman, S.K, Chen, J.H, Xiao, Y. | | Deposit date: | 2011-06-21 | | Release date: | 2012-10-31 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal Structure Analysis of TRF2-Dbd-DNA complex
To be Published
|
|
3R9F
 
 | |
3R9E
 
 | |
3R95
 
 | |
3R9G
 
 | |
3R96
 
 | |
3G7D
 
 | | Native PhpD with Cadmium Atoms | | Descriptor: | CADMIUM ION, PhpD | | Authors: | Nair, S.K. | | Deposit date: | 2009-02-09 | | Release date: | 2009-06-09 | | Last modified: | 2012-03-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | An unusual carbon-carbon bond cleavage reaction during phosphinothricin biosynthesis.
Nature, 459, 2009
|
|
2G02
 | | Nisin cyclase | | Descriptor: | Nisin biosynthesis protein nisC, ZINC ION | | Authors: | Nair, S.K. | | Deposit date: | 2006-02-10 | | Release date: | 2006-05-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and Mechanism of the Lantibiotic Cyclase Involved in Nisin Biosynthesis
Science, 311, 2006
|
|
8D0V
 
 | | Human LanCL1 C264A mutant bound to GSH | | Descriptor: | GLUTATHIONE, Glutathione S-transferase LANCL1, ZINC ION | | Authors: | Ongpipattanakul, C, Nair, S.K. | | Deposit date: | 2022-05-26 | | Release date: | 2023-01-25 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | The mechanism of thia-Michael addition catalyzed by LanC enzymes.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
6OM4
 
 | |
4WD9
 
 | |
4MGS
 
 | | BiXyn10A CBM1 APO | | Descriptor: | Putative glycosyl hydrolase family 10 | | Authors: | Chekan, J.R, Nair, S.K. | | Deposit date: | 2013-08-28 | | Release date: | 2014-08-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Xylan utilization in human gut commensal bacteria is orchestrated by unique modular organization of polysaccharide-degrading enzymes.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
5DZT
 
 | |
