3ND3
 
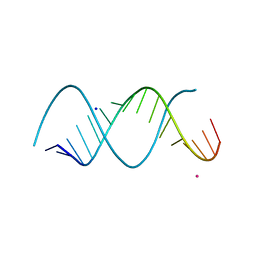 | | Uhelix 16-mer dsRNA | | Descriptor: | 5'-R(*AP*GP*AP*GP*AP*AP*GP*AP*UP*UP*UP*UP*UP*UP*UP*U)-3', POTASSIUM ION, SODIUM ION | | Authors: | Mooers, B.H, Singh, A. | | Deposit date: | 2010-06-07 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | The crystal structure of an oligo(U):pre-mRNA duplex from a trypanosome RNA editing substrate.
Rna, 17, 2011
|
|
5D99
 
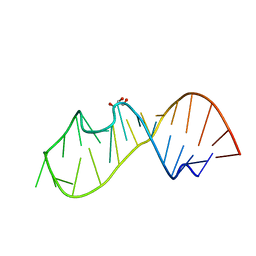 | | 3DW4 redetermined by direct methods starting from random phase angles | | Descriptor: | GLYCEROL, RNA (27-MER) hairpin from sarcin-ricin domain of E. coli 23S rRNA | | Authors: | Mooers, B.H.M. | | Deposit date: | 2015-08-18 | | Release date: | 2016-04-13 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (0.97 Å) | | Cite: | Direct-methods structure determination of a trypanosome RNA-editing substrate fragment with translational pseudosymmetry.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
5DA6
 
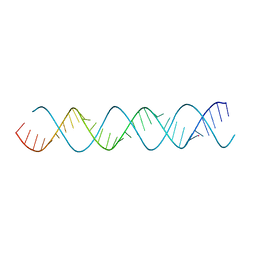 | |
4RBQ
 
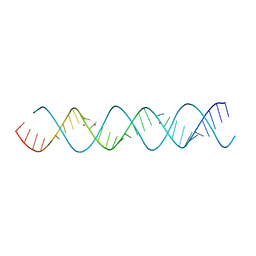 | | 32 base pair oligo(U) RNA | | Descriptor: | POTASSIUM ION, U-Helix RNA from Trypanosome editing | | Authors: | Mooers, B.H.M. | | Deposit date: | 2014-09-12 | | Release date: | 2015-11-11 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Structure of the Trypanosome RNA Editing U-Helix with 16 Contiguous Us
To be Published
|
|
3ND4
 
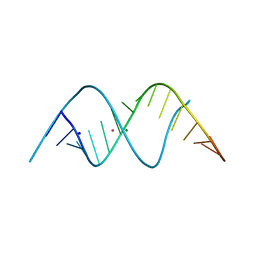 | | Watson-Crick 16-mer dsRNA | | Descriptor: | 5'-R(*AP*GP*AP*GP*AP*AP*GP*AP*UP*CP*UP*UP*CP*UP*CP*U)-3', MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Mooers, B.H, Singh, A. | | Deposit date: | 2010-06-07 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.524 Å) | | Cite: | The crystal structure of an oligo(U):pre-mRNA duplex from a trypanosome RNA editing substrate.
Rna, 17, 2011
|
|
4PCO
 
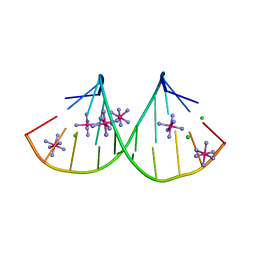 | |
1SX2
 
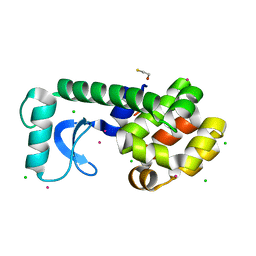 | |
1SX7
 
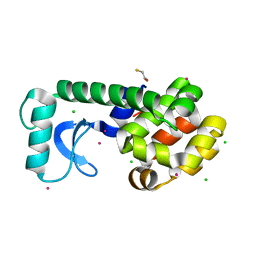 | |
1SWZ
 
 | |
3FI5
 
 | | Crystal Structure of T4 Lysozyme Mutant R96W | | Descriptor: | CHLORIDE ION, ISOPROPYL ALCOHOL, Lysozyme, ... | | Authors: | Mooers, B.H.M, Matthews, B.W. | | Deposit date: | 2008-12-11 | | Release date: | 2009-02-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Contributions of all 20 amino acids at site 96 to the stability and structure of T4 lysozyme.
Protein Sci., 18, 2009
|
|
3F9L
 
 | |
3F8V
 
 | |
3FA0
 
 | |
3FAD
 
 | |
312D
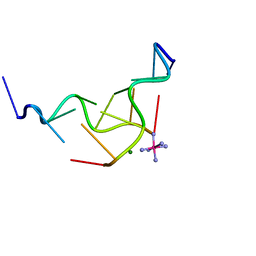 
 | | Z-DNA HEXAMER WITH 5' OVERHANGS THAT FORM A REVERSE WATSON-CRICK BASE PAIR | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*CP*CP*GP*CP*GP*CP*G)-3'), DNA (5'-D(*GP*CP*GP*CP*GP*CP*G)-3'), ... | | Authors: | Mooers, B.H.M, Eichman, B.F, Ho, P.S. | | Deposit date: | 1997-02-04 | | Release date: | 1997-08-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structures and relative stabilities of d(G x G) reverse Hoogsteen, d(G x T) reverse wobble, and d(G x C) reverse Watson-Crick base-pairs in DNA crystals.
J.Mol.Biol., 269, 1997
|
|
313D
 
 | | Z-DNA HEXAMER WITH 5' OVERHANGS THAT FORM A REVERSE HOOGSTEEN BASE PAIR | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*GP*(5CM)P*GP*CP*GP*CP*G)-3'), MAGNESIUM ION | | Authors: | Mooers, B.H.M, Eichman, B.F, Ho, P.S. | | Deposit date: | 1997-02-04 | | Release date: | 1997-08-05 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | The structures and relative stabilities of d(G x G) reverse Hoogsteen, d(G x T) reverse wobble, and d(G x C) reverse Watson-Crick base-pairs in DNA crystals.
J.Mol.Biol., 269, 1997
|
|
338D
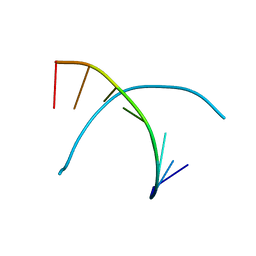 
 | |
339D
 
 | |
340D
 
 | |
343D
 
 | |
1ZEV
 
 | |
253D
 
 | |
257D
 
 | |
256D
 
 | |
2ANV
 
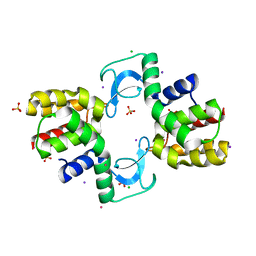 | | crystal structure of P22 lysozyme mutant L86M | | Descriptor: | CHLORIDE ION, IODIDE ION, Lysozyme, ... | | Authors: | Mooers, B.H, Matthews, B.W. | | Deposit date: | 2005-08-11 | | Release date: | 2006-02-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Extension to 2268 atoms of direct methods in the ab initio determination of the unknown structure of bacteriophage P22 lysozyme.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
