1VBI
 
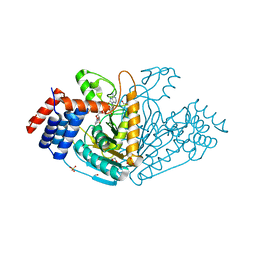 | |
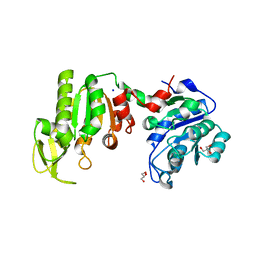
1V6S
 
 | |
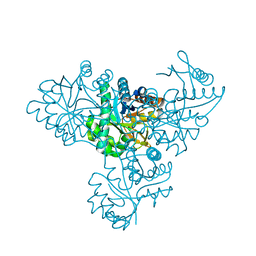
1V6T
 
 | |
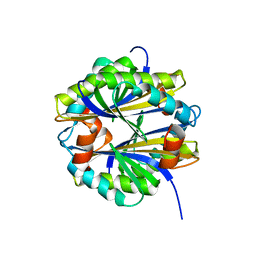
2EKY
 
 | |
2DXW
 
 | |
2EHC
 
 | |
2E17
 
 | |
2E5X
 
 | | Structure of nucleotide triphosphate pyrophosphatase from pyrococcus horikoshii OT3 | | Descriptor: | 1,2-ETHANEDIOL, Hypothetical protein PH1917, INOSINE 5'-TRIPHOSPHATE, ... | | Authors: | Mizutani, H, Lokanath, N.K, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-12-25 | | Release date: | 2007-06-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of nucleotide triphosphate pyrophosphatase from pyrococcus horikoshii OT3
To be Published
|
|
2EGB
 
 | | Crystal structure of Glu140 to Asn mutant of Diphthine synthase | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GLYCEROL, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Mizutani, H, Matsuura, Y, Krishna Swamy, B.S, Simanshu, D.K, Murthy, M.R.N, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-02-28 | | Release date: | 2007-08-28 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of diphthine synthase from Pyrococcus horikoshii OT3
To be Published
|
|
2E5W
 
 | |
2EHT
 
 | |
2EPI
 
 | |
2DXX
 
 | |
2E16
 
 | |
2EHS
 
 | |
2DXV
 
 | |
2E07
 
 | |
2E08
 
 | |
1EVI
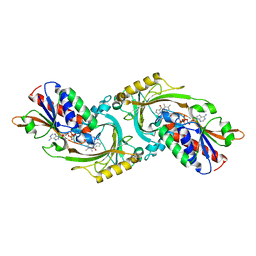 
 | | THREE-DIMENSIONAL STRUCTURE OF THE PURPLE INTERMEDIATE OF PORCINE KIDNEY D-AMINO ACID OXIDASE | | Descriptor: | 3,4-DIHYDRO-2H-PYRROLIUM-5-CARBOXYLATE, D-AMINO ACID OXIDASE, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Mizutani, H, Miyahara, I, Hirotsu, K, Nishina, Y, Shiga, K, Setoyama, C, Miura, R. | | Deposit date: | 2000-04-20 | | Release date: | 2000-10-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Three-dimensional structure of the purple intermediate of porcine kidney D-amino acid oxidase. Optimization of the oxidative half-reaction through alignment of the product with reduced flavin.
J.Biochem.(Tokyo), 128, 2000
|
|
2E7U
 
 | | Crystal structure of glutamate-1-semialdehyde 2,1-aminomutase from Thermus thermophilus HB8 | | Descriptor: | 4'-DEOXY-4'-AMINOPYRIDOXAL-5'-PHOSPHATE, GLYCEROL, Glutamate-1-semialdehyde 2,1-aminomutase, ... | | Authors: | Mizutani, H, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-14 | | Release date: | 2007-07-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of glutamate-1-semialdehyde 2,1-aminomutase from Thermus thermophilus HB8
To be Published
|
|
2E54
 
 | |
2EH6
 
 | |
2EO5
 
 | |
2EPJ
 
 | |
2EO4
 
 | |
