1HZF
 
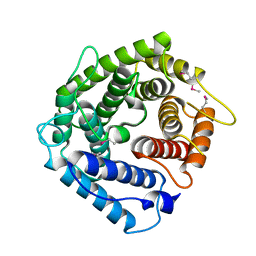 | | C4ADG FRAGMENT OF HUMAN COMPLEMENT FACTOR C4A | | Descriptor: | COMPLEMENT FACTOR C4A | | Authors: | van den Elsen, J.M.H, Martin, A, Wong, V, Clemenza, L, Rose, D.R, Isenman, D.E. | | Deposit date: | 2001-01-24 | | Release date: | 2002-10-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | X-ray crystal structure of the C4d fragment of human complement component C4.
J.Mol.Biol., 322, 2002
|
|
2LZP
 
 | |
2LZQ
 | |
2MKB
 
 | |
4O8Y
 
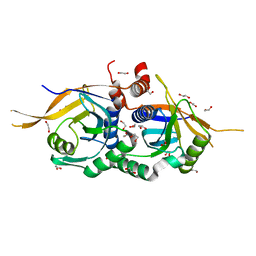 | | Zinc-free Rpn11 in complex with Rpn8 | | Descriptor: | 1,2-ETHANEDIOL, 26S proteasome regulatory subunit RPN11, 26S proteasome regulatory subunit RPN8 | | Authors: | Worden, E.J, Padovani, C, Martin, A. | | Deposit date: | 2013-12-30 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of the Rpn11-Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4O8X
 
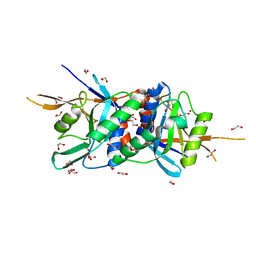 | | Zinc-bound Rpn11 in complex with Rpn8 | | Descriptor: | 1,2-ETHANEDIOL, 26S proteasome regulatory subunit RPN11, 26S proteasome regulatory subunit RPN8, ... | | Authors: | Worden, E.J, Padovani, C, Martin, A. | | Deposit date: | 2013-12-30 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.991 Å) | | Cite: | Structure of the Rpn11-Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation.
Nat.Struct.Mol.Biol., 21, 2014
|
|
1AZ4
 
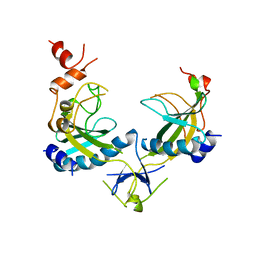 | | ECORV ENDONUCLEASE, UNLIGANDED, FORM B, T93A MUTANT | | Descriptor: | ECORV ENDONUCLEASE | | Authors: | Perona, J, Martin, A. | | Deposit date: | 1997-11-24 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Conformational transitions and structural deformability of EcoRV endonuclease revealed by crystallographic analysis.
J.Mol.Biol., 273, 1997
|
|
1AZ3
 
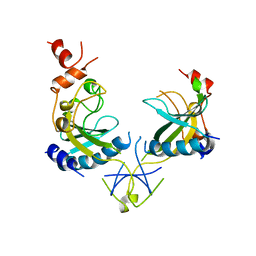 | | ECORV ENDONUCLEASE, UNLIGANDED, FORM B | | Descriptor: | ECORV ENDONUCLEASE | | Authors: | Perona, J, Martin, A. | | Deposit date: | 1997-11-24 | | Release date: | 1998-05-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Conformational transitions and structural deformability of EcoRV endonuclease revealed by crystallographic analysis.
J.Mol.Biol., 273, 1997
|
|
5U4P
 | | Protein-protein complex between 26S proteasome regulatory subunit RPN8, RPN11, and Ubiquitin S31 | | Descriptor: | 26S proteasome regulatory subunit RPN11, 26S proteasome regulatory subunit RPN8, Ubiquitin-40S ribosomal protein S31, ... | | Authors: | Worden, E.J, Dong, K.C, Martin, A. | | Deposit date: | 2016-12-05 | | Release date: | 2017-09-06 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | An AAA Motor-Driven Mechanical Switch in Rpn11 Controls Deubiquitination at the 26S Proteasome.
Mol. Cell, 67, 2017
|
|
