4Q77
 
 | | Crystal structure of Rot, a global regulator of virulence genes in Staphylococcus aureus | | Descriptor: | GLYCEROL, HTH-type transcriptional regulator rot | | Authors: | Zhu, Y, Fan, X, Li, X, Teng, M. | | Deposit date: | 2014-04-24 | | Release date: | 2014-09-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structure of Rot, a global regulator of virulence genes in Staphylococcus aureus.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4PYR
 
 | | Structure of a putative branched-chain amino acid ABC transporter from Chromobacterium violaceum ATCC 12472 | | Descriptor: | GLUTATHIONE, Putative branched-chain amino acid ABC transporter | | Authors: | Filippova, E.V, Minasov, G, Shuvalova, L, Kiryukhina, O, Endres, M, Joachimiak, A, Anderson, W.F, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2014-03-27 | | Release date: | 2014-04-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure of a putative branched-chain amino acid ABC transporter from Chromobacterium violaceum ATCC 12472
To be Published
|
|
3OE4
 
 | | Rat catechol O-methyltransferase in complex with a catechol-type, purine-containing bisubstrate inhibitor - humanized form | | Descriptor: | Catechol O-methyltransferase, MAGNESIUM ION, N-[(E)-3-[(2R,3S,4R,5R)-3,4-dihydroxy-5-purin-9-yl-oxolan-2-yl]prop-2-enyl]-2,3-dihydroxy-5-nitro-benzamide | | Authors: | Ehler, A, Schlatter, D, Stihle, M, Benz, J, Rudolph, M.G. | | Deposit date: | 2010-08-12 | | Release date: | 2011-03-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Molecular Recognition at the Active Site of Catechol-O-methyltransferase (COMT): Adenine Replacements in Bisubstrate Inhibitors
Chemistry, 17, 2011
|
|
8QHG
 
 | | Human Carbonic Anhydrase IX mimic in complex with Lasamide (2,4-Dichloro 5-sulfamoyl benzoic acid) | | Descriptor: | 2,4-dichloro-5-sulfamoylbenzoic acid, Carbonic anhydrase 2, NICKEL (II) ION, ... | | Authors: | Baroni, C, Ferraroni, M, Carta, F. | | Deposit date: | 2023-09-08 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Human Carbonic Anhydrase IX mimic in complex with Lasamide (2,4-Dichloro 5-sulfamoyl benzoic acid)
To Be Published
|
|
4Q7M
 
 | | Structure of NBD288-Avi of TM287/288 | | Descriptor: | Uncharacterized ABC transporter ATP-binding protein TM_0288 | | Authors: | Bukowska, M.A, Hohl, M, Gruetter, M.G, Seeger, M.A. | | Deposit date: | 2014-04-25 | | Release date: | 2015-05-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A Transporter Motor Taken Apart: Flexibility in the Nucleotide Binding Domains of a Heterodimeric ABC Exporter.
Biochemistry, 54, 2015
|
|
4Q8S
 
 | | Crystal structure of mammalian Peptidoglycan recognition protein PGRP-S with paranitrophenyl palmitate and N-acetyl glucosamine at 2.09 A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-nitrophenyl hexadecanoate, GLYCEROL, ... | | Authors: | Yamini, S, Sharma, P, Sinha, M, Bhushan, A, Kaur, P, Sharma, S, Singh, T.P. | | Deposit date: | 2014-04-28 | | Release date: | 2014-05-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystal structure of mammalian Peptidoglycan recognition protein PGRP-S with paranitrophenyl palmitate and N-acetyl glucosamine at 2.09 A resolution
To be Published
|
|
4Q9E
 
 | | Structure of the ternary complex of peptidoglycan recognition protein, PGRP-S with N-acetyl glucosamine and paranitro benzaldehyde at 2.3 A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-nitrobenzaldehyde, GLYCEROL, ... | | Authors: | Yamini, S, Sharma, P, Yadav, S.P, Sinha, M, Kaur, P, Sharma, S, Singh, T.P. | | Deposit date: | 2014-05-01 | | Release date: | 2014-05-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Structure of the ternary complex of peptidoglycan recognition protein, PGRP-S with N-acetyl glucosamine and paranitro benzaldehyde at 2.3 A resolution
to be published
|
|
4QA9
 
 | | Ensemble refinement of an epoxide hydrolase from Streptomyces carzinostaticus subsp. neocarzinostaticus. | | Descriptor: | 1,2-ETHANEDIOL, Epoxide hydrolase, SULFATE ION | | Authors: | Wang, F, Tan, K, Bigelow, L, Clancy, S, Babnigg, G, Bingman, C.A, Yennamalli, R, Lohman, J, Ma, M, Shen, B, Joachimiak, A, Phillips Jr, G.N, Midwest Center for Structural Genomics (MCSG), Enzyme Discovery for Natural Product Biosynthesis (NatPro) | | Deposit date: | 2014-05-02 | | Release date: | 2014-05-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Ensemble refinement of an epoxide hydrolase from Streptomyces carzinostaticus subsp. neocarzinostaticus.
To be Published
|
|
4Q1I
 
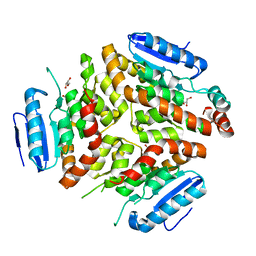 | |
4QGE
 
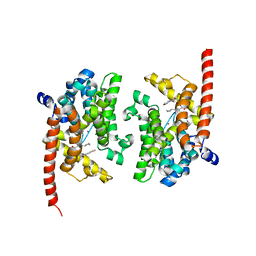 | | phosphodiesterase-9A in complex with inhibitor WYQ-C36D | | Descriptor: | MAGNESIUM ION, N~2~-(1-cyclopentyl-4-oxo-4,7-dihydro-1H-pyrazolo[3,4-d]pyrimidin-6-yl)-N-(4-methoxyphenyl)-D-alaninamide, Phosphodiesterase 9A, ... | | Authors: | Shao, Y.-X, Huang, M, Cui, W, Ke, H. | | Deposit date: | 2014-05-22 | | Release date: | 2014-12-10 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of a Phosphodiesterase 9A Inhibitor as a Potential Hypoglycemic Agent.
J.Med.Chem., 57, 2014
|
|
4QGX
 
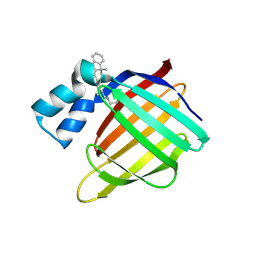 | | Crystal structure of the R132K:R111L:L121E mutant of Cellular Retinoic Acid Binding ProteinII complexed with a synthetic ligand (Merocyanine) at 1.47 angstrom resolution | | Descriptor: | (2E,4E,6E)-3-methyl-6-(1,3,3-trimethyl-1,3-dihydro-2H-indol-2-ylidene)hexa-2,4-dienal, Cellular retinoic acid-binding protein 2 | | Authors: | Nosrati, M, Yapici, I, Geiger, J.H. | | Deposit date: | 2014-05-26 | | Release date: | 2015-01-28 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.471 Å) | | Cite: | "Turn-on" protein fluorescence: in situ formation of cyanine dyes.
J.Am.Chem.Soc., 137, 2015
|
|
4QH5
 
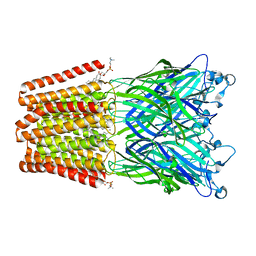 | | The GLIC pentameric Ligand-Gated Ion Channel (wild-type) crystallized in phosphate buffer | | Descriptor: | CHLORIDE ION, DIUNDECYL PHOSPHATIDYL CHOLINE, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Fourati, Z, Delarue, M, Sauguet, L. | | Deposit date: | 2014-05-26 | | Release date: | 2015-03-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural characterization of potential modulation sites in the extracellular domain of the prokaryotic pentameric proton-gated ion channel GLIC
Acta Crystallogr.,Sect.D, 2015
|
|
5MBZ
 
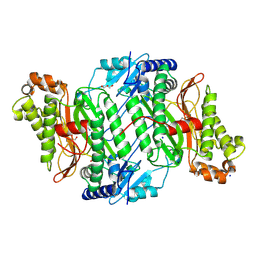 | | Crystal Structure of Ser202Phe mutant of Human Prolidase with Mn ions and GlyPro ligand | | Descriptor: | CHLORIDE ION, GLYCEROL, GLYCINE, ... | | Authors: | Wilk, P, Mueller, U, Dobbek, H, Weiss, M.S. | | Deposit date: | 2016-11-09 | | Release date: | 2017-12-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for prolidase deficiency disease mechanisms.
FEBS J., 285, 2018
|
|
8QK4
 
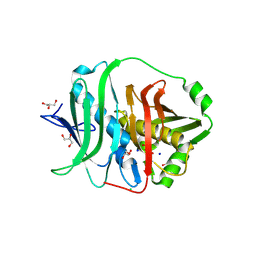 | |
8QPG
 
 | | Turret of Haloferax tailed virus 1 | | Descriptor: | MAGNESIUM ION, Prokaryotic polysaccharide deacetylase, ZINC ION, ... | | Authors: | Zhang, D, Daum, B, Isupov, M.N, McLaren, M. | | Deposit date: | 2023-10-02 | | Release date: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (2.36 Å) | | Cite: | CryoEM structure of Haloferax tailed virus 1
To Be Published
|
|
7T0J
 
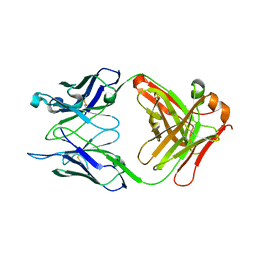 | |
8QPQ
 
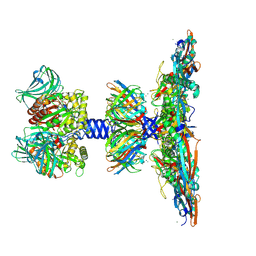 | | C1 turret to capsid interface of full Haloferax tailed virus 1 adjacent to the portal-capsid interface. | | Descriptor: | HK97 gp5-like major capsid protein, MAGNESIUM ION, Prokaryotic polysaccharide deacetylase, ... | | Authors: | Zhang, D, Daum, B, Isupov, M.N, McLaren, M. | | Deposit date: | 2023-10-02 | | Release date: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | CryoEM structure of Haloferax tailed virus 1
To Be Published
|
|
7T0H
 
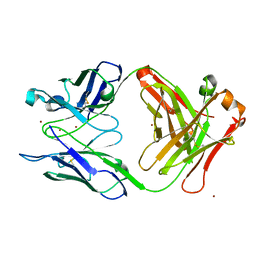 | |
8QH8
 
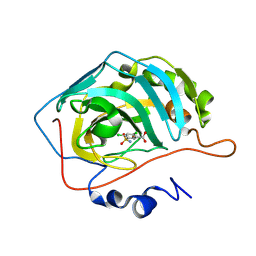 | | Human Carbonic Anhydrase II in complex with Lasamide (2,4-Dichloro 5-sulfamoyl benzoic acid) | | Descriptor: | 2,4-dichloro-5-sulfamoylbenzoic acid, Carbonic anhydrase 2, ZINC ION | | Authors: | Baroni, C, Ferraroni, M, Carta, F. | | Deposit date: | 2023-09-06 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Human Carbonic Anhydrase II in complex with Lasamide (2,4-Dichloro 5-sulfamoyl benzoic acid)
To Be Published
|
|
7T0I
 
 | |
7T0G
 
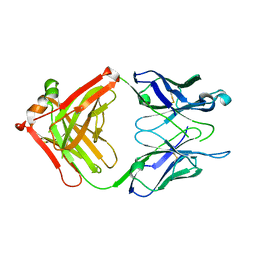
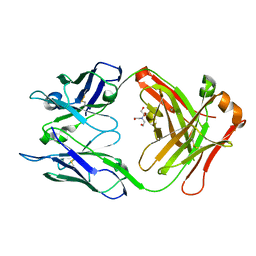 | | Crystal structure of S25-39 Fab Unliganded 1 | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, S25-39 Fab heavy chain, S25-39 Fab light chain | | Authors: | Legg, M.S.G, Blackler, R.J, Evans, S.V. | | Deposit date: | 2021-11-29 | | Release date: | 2022-04-20 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Antigen binding by conformational selection in near-germline antibodies.
J.Biol.Chem., 298, 2022
|
|
8QKZ
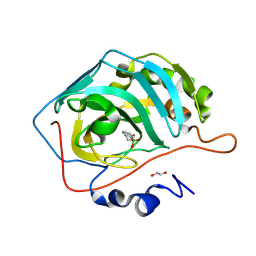 
 | | Human Carbonic Anhydrase II in complex with 3,4-dihydro-1H-benzo[c][1,2]oxaborinin-1-ol at pH 9.0 | | Descriptor: | 1,1-bis(oxidanyl)-3,4-dihydro-2,1$l^{4}-benzoxaborinine, 1,2-ETHANEDIOL, Carbonic anhydrase 2, ... | | Authors: | Angeli, A, Ferraroni, M. | | Deposit date: | 2023-09-18 | | Release date: | 2024-10-02 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Human Carbonic Anhydrase II in complex with 3,4-dihydro-1H-benzo[c][1,2]oxaborinin-1-ol at pH 9.0
To Be Published
|
|
8QKW
 
 | | Crystal structure of Levansucrase from Pseudomonas syringae in complex with a tetravalent iminosugar | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-(hydroxymethyl)-1-[6-[4-[[3-[[3-[6-[(2~{S},3~{R},4~{S})-2-(hydroxymethyl)-3,4-bis(oxidanyl)pyrrolidin-1-yl]hexyl]-1,2,3-triazol-4-yl]methoxy]-2,2-bis[[1-[6-[2-(hydroxymethyl)-3,4-bis(oxidanyl)pyrrolidin-1-yl]hexyl]-1,2,3-triazol-4-yl]methoxymethyl]propoxy]methyl]-1,2,3-triazol-1-yl]hexyl]pyrrolidine-3,4-diol, DIMETHYL SULFOXIDE, ... | | Authors: | Ferraroni, M, Canovai, A. | | Deposit date: | 2023-09-18 | | Release date: | 2024-10-02 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of Levansucrase from Pseudomonas syringae in complex with a tetravalent iminosugar
To Be Published
|
|
8QUF
 
 | |
4Q6B
 
 | | Crystal Structure of ABC Transporter Substrate-Binding Protein fromDesulfitobacterium hafniense complex with Leu | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Extracellular ligand-binding receptor, ... | | Authors: | Kim, Y, Chhor, G, Endres, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2014-04-22 | | Release date: | 2014-07-02 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.667 Å) | | Cite: | Crystal Structure of ABC Transporter Substrate-Binding Protein fromDesulfitobacterium hafniense complex with Leu
To be Published, 2014
|
|
