6IR9
 
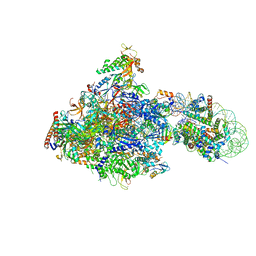 | | RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-1) of the nucleosome | | Descriptor: | DNA (198-MER), DNA-directed RNA polymerase subunit, DNA-directed RNA polymerase subunit beta, ... | | Authors: | Ehara, H, Kujirai, T, Fujino, Y, Shirouzu, M, Kurumizaka, H, Sekine, S. | | Deposit date: | 2018-11-12 | | Release date: | 2019-02-20 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural insight into nucleosome transcription by RNA polymerase II with elongation factors.
Science, 363, 2019
|
|
6R6N
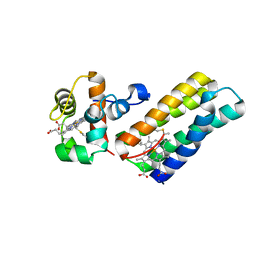 
 | |
3DD0
 
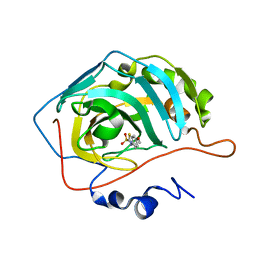 | | Use of Carbonic Anhydrase II, IX Active-Site Mimic, for the Purpose of Screening Inhibitors for Possible Anti-Cancer Properties | | Descriptor: | 6-ethoxy-1,3-benzothiazole-2-sulfonamide, Carbonic anhydrase 2, ZINC ION | | Authors: | Genis, C, Sippel, K.H, Case, N, Govindasamy, L, Agbandje-Mckenna, M, Mckenna, R. | | Deposit date: | 2008-06-04 | | Release date: | 2009-03-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Design of a carbonic anhydrase IX active-site mimic to screen inhibitors for possible anticancer properties
Biochemistry, 48, 2009
|
|
1HV5
 
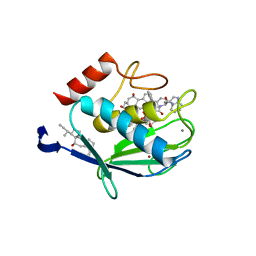 | | CRYSTAL STRUCTURE OF THE STROMELYSIN-3 (MMP-11) CATALYTIC DOMAIN COMPLEXED WITH A PHOSPHINIC INHIBITOR | | Descriptor: | 1-BENZYLOXYCARBONYLAMINO-2-PHENYL-ETHYL)-{2-[1-CARBAMOYL-2-(1H-INDOL-3-YL)-ETHYLCARBAMOYL]-5-PHENYL-PENTYL}-PHOSPHINIC ACID, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, CALCIUM ION, ... | | Authors: | Gall, A.L, Ruff, M, Kannan, R, Cuniasse, P, Yiotakis, A, Dive, V, Rio, M.C, Basset, P, Moras, D. | | Deposit date: | 2001-01-08 | | Release date: | 2001-03-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the stromelysin-3 (MMP-11) catalytic domain complexed with a phosphinic inhibitor mimicking the transition-state.
J.Mol.Biol., 307, 2001
|
|
4I29
 
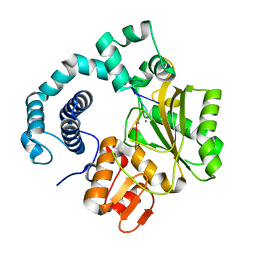 | | Binary complex of mouse TdT with ssDNA and Mn++ | | Descriptor: | 5'-D(*AP*AP*(8BA)P*AP*A)-3', DNA nucleotidylexotransferase, MANGANESE (II) ION, ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2012-11-21 | | Release date: | 2013-07-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of Intermediates along the Catalytic Cycle of Terminal Deoxynucleotidyltransferase: Dynamical Aspects of the Two-Metal Ion Mechanism.
J.Mol.Biol., 425, 2013
|
|
6E16
 
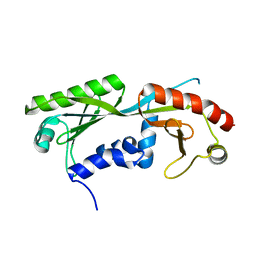 | | Ternary structure of c-Myc-TBP-TAF1 | | Descriptor: | Transcription initiation factor TFIID subunit 1,Myc proto-oncogene protein,TATA-box-binding protein | | Authors: | Wei, Y, Dong, A, Sunnerhagen, M, Penn, L, Tong, Y, Edwards, A.M, Arrowsmith, C.H, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-07-09 | | Release date: | 2019-10-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Multiple direct interactions of TBP with the MYC oncoprotein.
Nat.Struct.Mol.Biol., 26, 2019
|
|
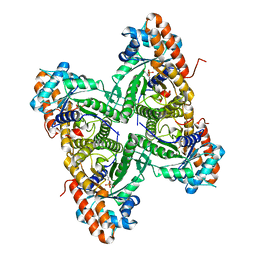
3DFS
 
 | |
3P13
 
 | |
6ITF
 
 | |
1HZO
 | | STRUCTURE OF CLASS A CEPHALOSPORINASE FROM PROTEUS VULGARIS K1 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, BETA-LACTAMASE | | Authors: | Nukaga, M, Crichlow, G.V, Kuzin, A.P, Mayama, K, Knox, J.R. | | Deposit date: | 2001-01-25 | | Release date: | 2002-04-03 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of an extended-spectrum class A beta-lactamase from Proteus vulgaris K1.
J.Mol.Biol., 317, 2002
|
|
4I49
 
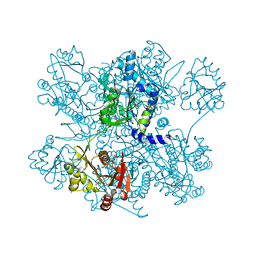 | | Structure of ngNAGS bound with bisubstrate analog CoA-NAG | | Descriptor: | (2S)-2-({(3S,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)tetrahydrofuran-2-yl]-3,5,9-trihydroxy-8,8-dimethyl-3,5-dioxido-10,14,20-trioxo-2,4,6-trioxa-18-thia-11,15-diaza-3lambda~5~,5lambda~5~-diphosphaicosan-20-yl}amino)pentanedioic acid (non-preferred name), Amino-acid acetyltransferase, SULFATE ION | | Authors: | Shi, D, Zhao, G, Allewell, N.M, Tuchman, M. | | Deposit date: | 2012-11-27 | | Release date: | 2013-01-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structure of the complex of Neisseria gonorrhoeae N-acetyl-l-glutamate synthase with a bound bisubstrate analog.
Biochem.Biophys.Res.Commun., 430, 2013
|
|
3DHZ
 
 | |
6E1R
 
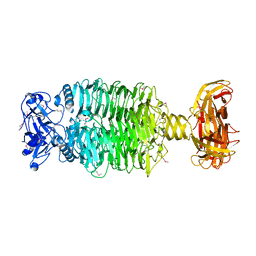 | | Crystal structure of the Acinetobacter phage vB_ApiP_P1 tailspike protein | | Descriptor: | CHLORIDE ION, SODIUM ION, Tailspike protein | | Authors: | Plattner, M, Shneider, M.M, Oliveira, H, Azeredo, J, Leiman, P.G. | | Deposit date: | 2018-07-10 | | Release date: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2.693 Å) | | Cite: | Crystal structure of the Acinetobacter phage vB_ApiP_P1 tailspike protein
To Be Published
|
|
3OP3
 
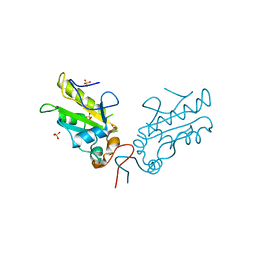 | | Crystal Structure of Cell Division Cycle 25C Protein Isoform A from Homo sapiens | | Descriptor: | M-phase inducer phosphatase 3, SULFATE ION | | Authors: | Kim, Y, Weger, A, Hatzos, C, Savitsky, P, Johansson, C, Ball, L, Barr, A, Vollmar, M, Muniz, J, Weigelt, J, Arrowsmith, C.H, Edwards, A, Bountra, C, Gileadi, O, von Delft, F, Knapp, S, Joachimiak, A, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-08-31 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Crystal Structure of Cell Division Cycle 25C Protein Isoform A from Homo sapiens
TO BE PUBLISHED
|
|
4MLS
 
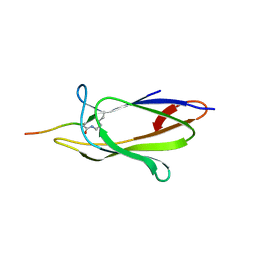 | | Crystal structure of the SpyTag and SpyCatcher-deltaN1 complex | | Descriptor: | Fibronectin binding protein, SpyTag | | Authors: | Li, L, Fierer, J.O, Rapoport, T.A, Howarth, M. | | Deposit date: | 2013-09-06 | | Release date: | 2013-11-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.984 Å) | | Cite: | Structural Analysis and Optimization of the Covalent Association between SpyCatcher and a Peptide Tag.
J.Mol.Biol., 426, 2014
|
|
3OP9
 
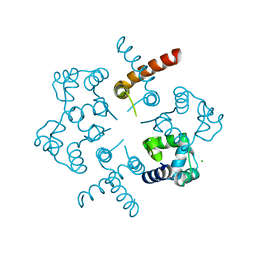 | |
4MLZ
 
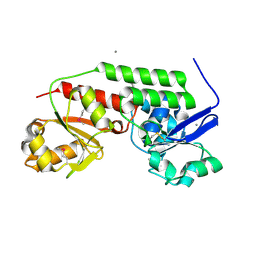 | | Crystal structure of periplasmic binding protein from Jonesia denitrificans | | Descriptor: | CALCIUM ION, POTASSIUM ION, Periplasmic binding protein | | Authors: | Chang, C, Chhor, G, Endres, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2013-09-06 | | Release date: | 2013-09-18 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Crystal structure of periplasmic binding protein from Jonesia denitrificans
To be Published
|
|
6QZ0
 | | The cryo-EM structure of the head of the genome empited bacteriophage phi29 | | Descriptor: | Capsid fiber protein, Major capsid protein | | Authors: | Xu, J, Wang, D, Gui, M, Xiang, Y. | | Deposit date: | 2019-03-10 | | Release date: | 2019-06-12 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural assembly of the tailed bacteriophage φ29.
Nat Commun, 10, 2019
|
|
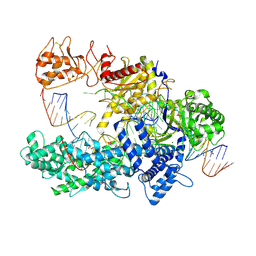
6I1K
 
 | |
6I19
 
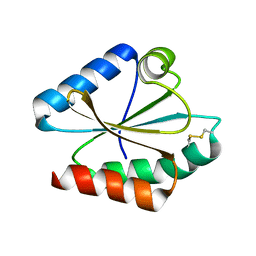 | | Crystal structure of Chlamydomonas reinhardtii thioredoxin h1 | | Descriptor: | Thioredoxin H-type | | Authors: | Lemaire, S.D, Tedesco, D, Crozet, P, Michelet, L, Fermani, S, Zaffagnini, M, Henri, J. | | Deposit date: | 2018-10-27 | | Release date: | 2018-12-05 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.378 Å) | | Cite: | Crystal Structure of Chloroplastic Thioredoxin f2 fromChlamydomonas reinhardtiiReveals Distinct Surface Properties.
Antioxidants (Basel), 7, 2018
|
|
6RB9
 | | The pore structure of Clostridium perfringens epsilon toxin | | Descriptor: | Epsilon-toxin type B | | Authors: | Savva, C.G, Clark, A.R, Naylor, C.E, Popoff, M.R, Moss, D.S, Basak, A.K, Titball, R.W, Bokori-Brown, M. | | Deposit date: | 2019-04-09 | | Release date: | 2019-06-19 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The pore structure of Clostridium perfringens epsilon toxin.
Nat Commun, 10, 2019
|
|
6I2S
 
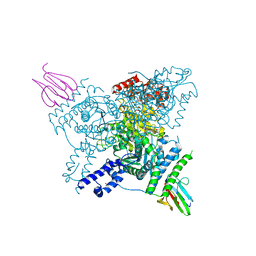 | | Crystal structure of the SucA domain of Mycobacterium smegmatis KGD (R802A) in complex with GarA, following 2-oxoglutarate soak | | Descriptor: | (4S)-4-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3lambda~5~-thiazol-2-yl}-4-hydroxybutanoic acid, CALCIUM ION, Glycogen accumulation regulator GarA, ... | | Authors: | Wagner, T, Bellinzoni, M, Alzari, P.M. | | Deposit date: | 2018-11-01 | | Release date: | 2019-05-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural insights into the functional versatility of an FHA domain protein in mycobacterial signaling.
Sci.Signal., 12, 2019
|
|
3P4E
 
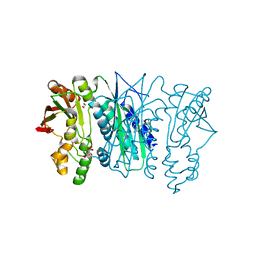 | | Phosphoribosylformylglycinamidine cyclo-ligase from Vibrio cholerae | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE MONOPHOSPHATE, CITRIC ACID, ... | | Authors: | Osipiuk, J, Zhou, M, Papazisi, L, Anderson, W.F, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2010-10-06 | | Release date: | 2010-10-20 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Phosphoribosylformylglycinamidine cyclo-ligase from Vibrio cholerae.
To be Published
|
|
4I2D
 
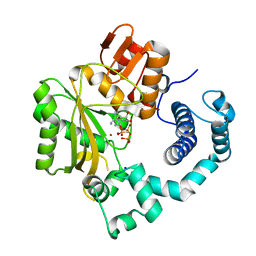 | | Binary complex of mouse TdT with AMPcPP | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, DNA nucleotidylexotransferase, MAGNESIUM ION, ... | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2012-11-21 | | Release date: | 2013-07-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of Intermediates along the Catalytic Cycle of Terminal Deoxynucleotidyltransferase: Dynamical Aspects of the Two-Metal Ion Mechanism.
J.Mol.Biol., 425, 2013
|
|
6RBT
 
 | |
