7L78
 
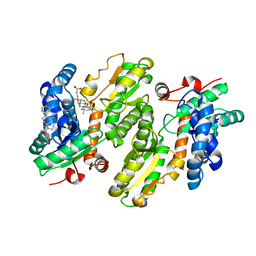 | | H235C variant of Yeast Ferrochelatase | | Descriptor: | CHOLIC ACID, Ferrochelatase, mitochondrial | | Authors: | Lanzilotta, W.N, Medlock, A.E. | | Deposit date: | 2020-12-27 | | Release date: | 2021-11-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Mitochondrial contact site and cristae organizing system (MICOS) machinery supports heme biosynthesis by enabling optimal performance of ferrochelatase.
Redox Biol, 46, 2021
|
|
4F4D
 
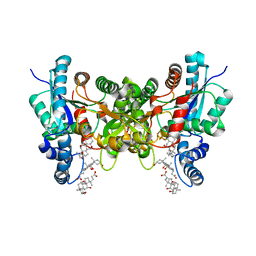 | | F337R variant of human ferrochelatase | | Descriptor: | CHLORIDE ION, CHOLIC ACID, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Lanzilotta, W.N, Medlock, A.E, Dailey, T.E, Dailey, H.A. | | Deposit date: | 2012-05-10 | | Release date: | 2012-09-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | F337R Variant of Human Ferrochelatase
TO BE PUBLISHED
|
|
6X6U
 
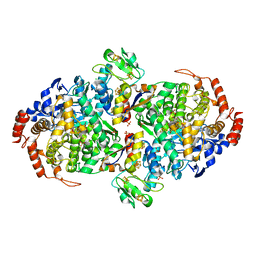 | | WOR5 from Pyrococcus furiosus, taurine-bound | | Descriptor: | 2-AMINOETHANESULFONIC ACID, CHLORIDE ION, Formaldehyde:ferredoxin oxidoreductase wor5, ... | | Authors: | Lanzilotta, W.N, Mathew, L.G. | | Deposit date: | 2020-05-29 | | Release date: | 2021-12-15 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.944 Å) | | Cite: | An unprecedented function for a tungsten-containing oxidoreductase.
J.Biol.Inorg.Chem., 27, 2022
|
|
3AQI
 
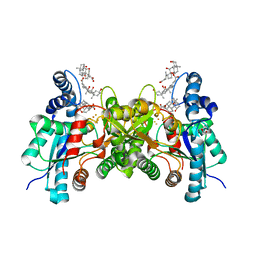 | | H240A variant of human ferrochelatase | | Descriptor: | CHOLIC ACID, FE2/S2 (INORGANIC) CLUSTER, Ferrochelatase, ... | | Authors: | Lanzilotta, W.N, Medlock, A.E, Dailey, T.A, Dailey, H.A. | | Deposit date: | 2010-11-03 | | Release date: | 2012-01-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | H240A variant of human ferrochelatase
To be published
|
|
1R9D
 
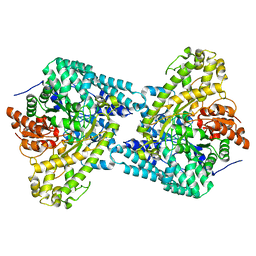 | | Glycerol bound form of the B12-independent glycerol dehydratase from Clostridium butyricum | | Descriptor: | GLYCEROL, glycerol dehydratase | | Authors: | Lanzilotta, W.N, O'Brien, J.R, Raynaud, C, Soucaille, P. | | Deposit date: | 2003-10-28 | | Release date: | 2004-06-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Insight into the mechanism of the B12-independent glycerol dehydratase from Clostridium butyricum: preliminary biochemical and structural characterization.
Biochemistry, 43, 2004
|
|
1R8W
 
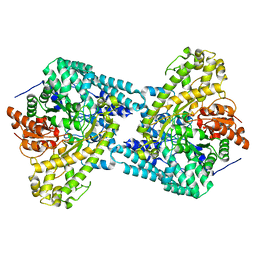 | |
4KMM
 
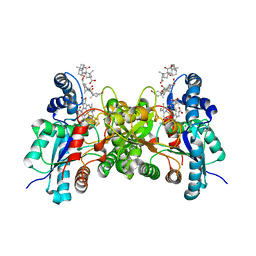 | |
4KLA
 
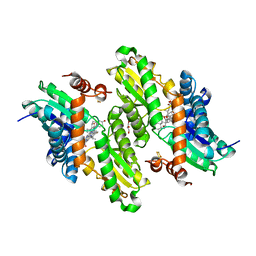 | |
4KLC
 
 | |
4KLR
 
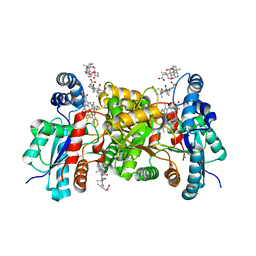 | |
4MK4
 
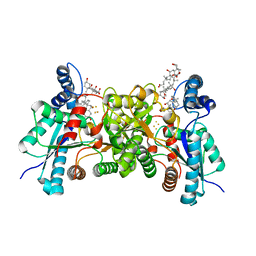 | | S197C variant of human ferrochelatase. | | Descriptor: | CHOLIC ACID, FE2/S2 (INORGANIC) CLUSTER, Ferrochelatase, ... | | Authors: | Lanzilotta, W.N, Medlock, A.E, Dailey, T.E, Dailey, H.A. | | Deposit date: | 2013-09-04 | | Release date: | 2013-09-25 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | S197C variant of human ferrochelatase.
To be Published
|
|
1FT9
 
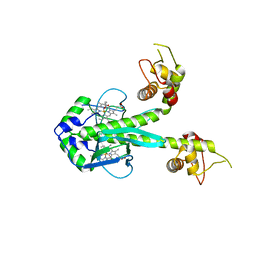 | | STRUCTURE OF THE REDUCED (FEII) CO-SENSING PROTEIN FROM R. RUBRUM | | Descriptor: | CARBON MONOXIDE OXIDATION SYSTEM TRANSCRIPTION REGULATOR, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Lanzilotta, W.N, Schuller, D.J, Poulos, T.L, Thorsteinsson, M.V, Kirby, B, Roberts, G.P. | | Deposit date: | 2000-09-12 | | Release date: | 2000-10-09 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the CO sensing transcription activator CooA.
Nat.Struct.Biol., 7, 2000
|
|
4MTJ
 
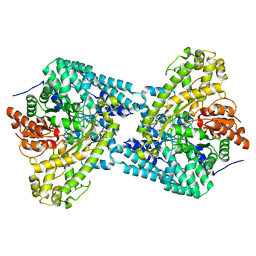 | | Structure of the b12-independent glycerol dehydratase with 1,2-propanediol bound | | Descriptor: | B12-independent glycerol dehydratase, S-1,2-PROPANEDIOL | | Authors: | LaMattina, J, Wright, A.V, Demick, J, Soucaille, P, Lanzilotta, W.N. | | Deposit date: | 2013-09-19 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | When Computational Chemistry and Modern Software Get It Right; New Insight Into the Mechanism of a Glycyl Radical Enzyme
To be Published
|
|
6X1O
 
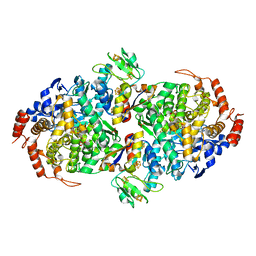 | | WOR5 from Pyrococcus furiosus, as crystallized | | Descriptor: | (1R)-1-hydroxybutane-1-sulfonic acid, CHLORIDE ION, Formaldehyde:ferredoxin oxidoreductase wor5, ... | | Authors: | Mathew, L.G, Lanzilotta, W.N. | | Deposit date: | 2020-05-19 | | Release date: | 2021-11-24 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.094 Å) | | Cite: | An unprecedented function for a tungsten-containing oxidoreductase.
J.Biol.Inorg.Chem., 27, 2022
|
|
3MPS
 
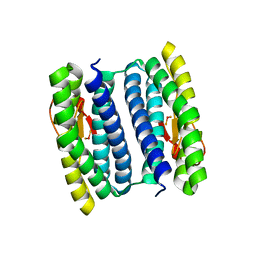 | | Peroxide Bound Oxidized Rubrerythrin from Pyrococcus furiosus | | Descriptor: | FE (III) ION, HYDROGEN PEROXIDE, MU-OXO-DIIRON, ... | | Authors: | Dillard, B.D, Adams, M.W.W, Lanzilotta, W.N. | | Deposit date: | 2010-04-27 | | Release date: | 2011-06-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A cryo-crystallographic time course for peroxide reduction by rubrerythrin from Pyrococcus furiosus.
J.Biol.Inorg.Chem., 16, 2011
|
|
5FFQ
 
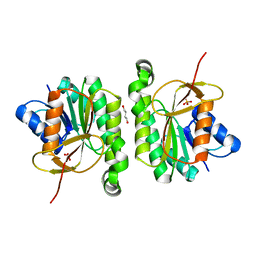 | | ChuY: An Anaerobillin Reductase from Escherichia coli O157:H7 | | Descriptor: | 1,4-BUTANEDIOL, PHOSPHATE ION, ShuY-like protein | | Authors: | LaMattina, J.W, Reedy, A.N, Uy, K.G, Lanzilotta, W.N. | | Deposit date: | 2015-12-18 | | Release date: | 2017-01-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Radical new paradigm for heme degradation in Escherichia coli O157:H7.
Proc. Natl. Acad. Sci. U.S.A., 113, 2016
|
|
4QGS
 
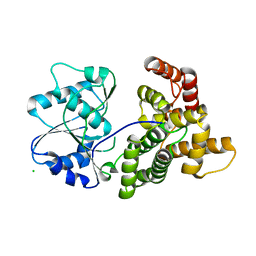 | |
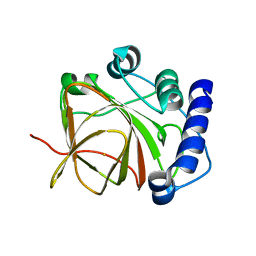
5CU1
 
 | |
6XFI
 
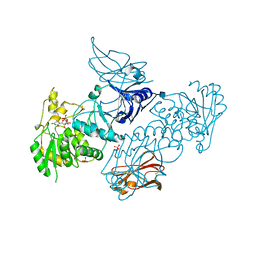 | | Crystal Structures of beta-1,4-N-Acetylglucosaminyltransferase 2 (POMGNT2): Structural Basis for Inherited Muscular Dystrophies | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PHOSPHATE ION, Protein O-linked-mannose beta-1,4-N-acetylglucosaminyltransferase 2, ... | | Authors: | Halmo, S.M, Yeh, J, Wells, L, Moremen, K.W, Lanzilotta, W.N. | | Deposit date: | 2020-06-15 | | Release date: | 2021-04-21 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of beta-1,4-N-acetylglucosaminyltransferase 2: structural basis for inherited muscular dystrophies.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
6XI2
 
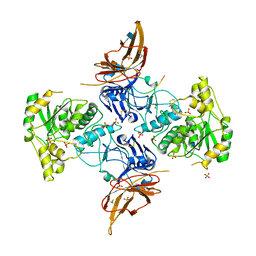 | | Apo form of POMGNT2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA-ALA, ALA-GLY-ALA-GLY-ALA-ALA-ALA-ALA-ALA-ALA, ... | | Authors: | Halmo, S.M, Yeh, J, Wells, L, Moremen, K.W, Lanzilotta, W.N. | | Deposit date: | 2020-06-19 | | Release date: | 2021-04-21 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Crystal structures of beta-1,4-N-acetylglucosaminyltransferase 2: structural basis for inherited muscular dystrophies.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
3QVD
 
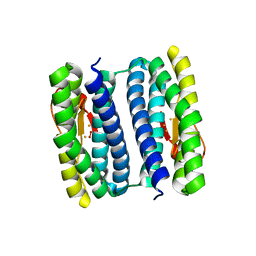 | | Exposure of rubrerythrin from Pyrococcus furiosus to peroxide, fifteen second time point. | | Descriptor: | FE (II) ION, FE (III) ION, HYDROGEN PEROXIDE, ... | | Authors: | Dillard, B.D, Demick, J.M, Adams, M.W.W, Lanzilotta, W.N. | | Deposit date: | 2011-02-25 | | Release date: | 2011-06-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A cryo-crystallographic time course for peroxide reduction by rubrerythrin from Pyrococcus furiosus.
J.Biol.Inorg.Chem., 16, 2011
|
|
3PWF
 
 | | High resolution structure of the fully reduced form of rubrerythrin from P. furiosus | | Descriptor: | FE (II) ION, Rubrerythrin | | Authors: | Dillard, B.D, Demick, J.M, Adams, M.W, Lanzilotta, W.N. | | Deposit date: | 2010-12-08 | | Release date: | 2011-06-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | A cryo-crystallographic time course for peroxide reduction by rubrerythrin from Pyrococcus furiosus.
J.Biol.Inorg.Chem., 16, 2011
|
|
3PZA
 
 | | Fully Reduced (All-ferrous) Pyrococcus rubrerythrin after a 10 second exposure to peroxide. | | Descriptor: | FE (II) ION, HYDROGEN PEROXIDE, Rubrerythrin | | Authors: | Dillard, B.D, Demick, J.M, Adams, M.W, Lanzilotta, W.N. | | Deposit date: | 2010-12-14 | | Release date: | 2011-06-22 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A cryo-crystallographic time course for peroxide reduction by rubrerythrin from Pyrococcus furiosus.
J.Biol.Inorg.Chem., 16, 2011
|
|
3TFI
 
 | | DMSP-dependent demethylase from P. ubique - with substrate DMSP | | Descriptor: | 3-(dimethyl-lambda~4~-sulfanyl)propanoic acid, GLYCEROL, GcvT-like Aminomethyltransferase protein, ... | | Authors: | Schuller, D.J, Reisch, C.R, Moran, M.A, Whitman, W.B, Lanzilotta, W.N. | | Deposit date: | 2011-08-15 | | Release date: | 2011-12-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structures of dimethylsulfoniopropionate-dependent demethylase from the marine organism Pelagabacter ubique.
Protein Sci., 21, 2012
|
|
3TFJ
 
 | | DMSP-dependent demethylase from P. ubique - with cofactor THF | | Descriptor: | (6S)-5,6,7,8-TETRAHYDROFOLATE, ACETATE ION, GLYCEROL, ... | | Authors: | Schuller, D.J, Reisch, C.R, Moran, M.A, Whitman, W.B, Lanzilotta, W.N. | | Deposit date: | 2011-08-15 | | Release date: | 2011-12-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structures of dimethylsulfoniopropionate-dependent demethylase from the marine organism Pelagabacter ubique.
Protein Sci., 21, 2012
|
|
