9GKX
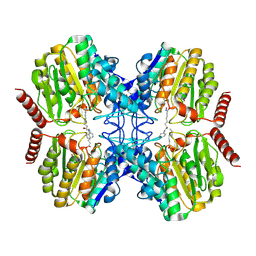 
 | | Crystal Structure of Rhizorhabdus wittichii Dimethoate hydrolase (DmhA) in complex with SAHA | | Descriptor: | Dimethoate hydrolase, OCTANEDIOIC ACID HYDROXYAMIDE PHENYLAMIDE, OCTANOIC ACID (CAPRYLIC ACID), ... | | Authors: | Graf, L.G, Lammers, M, Schulze, S, Palm, G.J. | | Deposit date: | 2024-08-26 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GKW
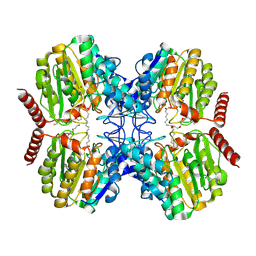 
 | | Crystal Structure of Dimethoate hydrolase (DmhA) of Rhizorhabdus wittichii in complex with octanoic acid | | Descriptor: | Dimethoate hydrolase, OCTANOIC ACID (CAPRYLIC ACID), PENTAETHYLENE GLYCOL, ... | | Authors: | Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-26 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GKZ
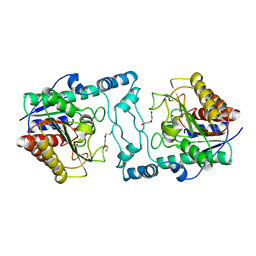 
 | | Crystal Structure of Acetylpolyamine amidohydrolase (ApaH) from Pseudomonas sp. M30-35 | | Descriptor: | ACETATE ION, Acetylpolyamine amidohydrolase, CHLORIDE ION, ... | | Authors: | Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-26 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GN7
 
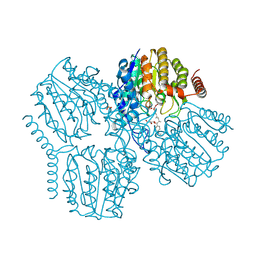 | | Crystal Structure of Deacetylase (HdaH) from Klebsiella pneumoniae subsp. ozaenae in Complex with the inhibitor TSA | | Descriptor: | Deacetylase, GLYCEROL, PHOSPHATE ION, ... | | Authors: | Qin, C, Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-30 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GN6
 
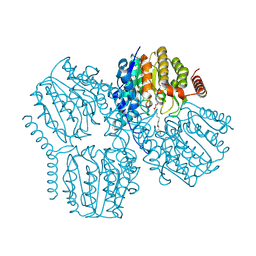 | | Crystal Structure of Deacetylase (HdaH) from Klebsiella pneumoniae subsp. ozaenae in complex with the inhibitor SAHA | | Descriptor: | Deacetylase, OCTANEDIOIC ACID HYDROXYAMIDE PHENYLAMIDE, POTASSIUM ION, ... | | Authors: | Qin, C, Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-30 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GL0
 
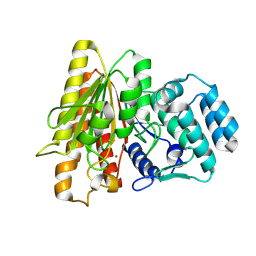 | | Crystal Structure of Acetylpolyamine aminohydrolase (ApaH) from Legionella pneumophila | | Descriptor: | Acetylpolyamine aminohydrolase, POTASSIUM ION, ZINC ION | | Authors: | Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-26 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GN1
 
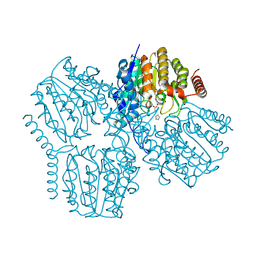 | | Crystal structure of inactive Deacetylase (HdaH) H144A from Klebsiella pneumoniae subsp. ozaenae | | Descriptor: | ACETATE ION, Deacetylase, IMIDAZOLE, ... | | Authors: | Qin, Q, Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-30 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GL1
 
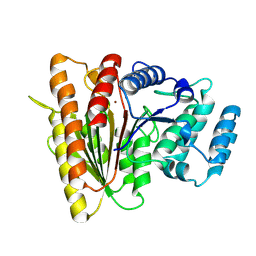 | | Crystal Structure of Acetylpolyamine aminohydrolase (ApaH) from Legionella cherrii | | Descriptor: | Acetylpolyamine aminohydrolase, POTASSIUM ION, ZINC ION | | Authors: | Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-26 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
9GLB
 
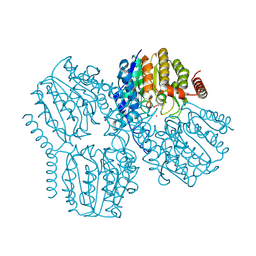 | | Crystal Structure of Deacetylase (HdaH) from Klebsiella pneumoniae subsp. ozaenae | | Descriptor: | ACETATE ION, Deacetylase, GLYCEROL, ... | | Authors: | Qin, C, Graf, L.G, Schulze, S, Palm, G.J, Lammers, M. | | Deposit date: | 2024-08-27 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Distribution and diversity of classical deacylases in bacteria
Nature Communications, 2024
|
|
8Y1J
 
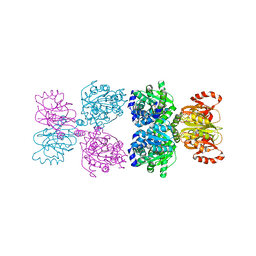 | |
8Y4Z
 
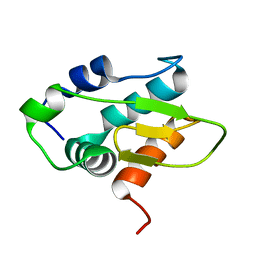 | | Monomeric HERC5 HECT c-lobe structure in solution | | Descriptor: | E3 ISG15--protein ligase HERC5 | | Authors: | Dag, C, Lambert, M, Kahraman, K, Lohn, F, Lee, W, Gocenler, O, Guntert, P, Dotsch, V. | | Deposit date: | 2024-01-31 | | Release date: | 2024-02-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Monomeric HERC5 HECT c-lobe structure in solution
To Be Published
|
|
3GWX
 
 | | MOLECULAR RECOGNITION OF FATTY ACIDS BY PEROXISOME PROLIFERATOR-ACTIVATED RECEPTORS | | Descriptor: | 5,8,11,14,17-EICOSAPENTAENOIC ACID, PROTEIN (PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR (PPAR-DELTA)) | | Authors: | Xu, H.E, Lambert, M.H, Montana, V.G, Parks, D.J, Blanchard, S.G, Brown, P.J, Sternbach, D.D, Lehmann, J.M, Wisely, G.B, Willson, T.M, Kliewer, S.A, Milburn, M.V. | | Deposit date: | 1999-04-26 | | Release date: | 2000-04-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Molecular recognition of fatty acids by peroxisome proliferator-activated receptors.
Mol.Cell, 3, 1999
|
|
3MW1
 
 | | p38 kinase Crystal structure in complex with small molecule inhibitor | | Descriptor: | 8-(2,6-dichlorophenyl)-4-(2,4-difluorophenyl)-2-piperidin-4-yl-1,7-naphthyridine 7-oxide, Mitogen-activated protein kinase 14 | | Authors: | Segarra, V, Caturla, F, Lumeras, W, Roca, R, Fisher, M, Lamers, M. | | Deposit date: | 2010-05-05 | | Release date: | 2011-04-27 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | 1,7-Naphthyridine 1-Oxides as Novel Potent and Selective Inhibitors of p38 Mitogen Activated Protein Kinase
J.Med.Chem., 54, 2011
|
|
3NGO
 
 | | Crystal structure of the human CNOT6L nuclease domain in complex with poly(A) DNA | | Descriptor: | 5'-D(*AP*AP*AP*A)-3', CCR4-NOT transcription complex subunit 6-like, MAGNESIUM ION | | Authors: | Wang, H, Morita, M, Yang, W, Bartlam, M, Yamamoto, T, Rao, Z. | | Deposit date: | 2010-06-12 | | Release date: | 2010-07-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the human CNOT6L nuclease domain reveals strict poly(A) substrate specificity.
Embo J., 2010
|
|
3NGQ
 
 | | Crystal structure of the human CNOT6L nuclease domain | | Descriptor: | 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, CCR4-NOT transcription complex subunit 6-like, MAGNESIUM ION | | Authors: | Wang, H, Morita, M, Yang, W, Bartlam, M, Yamamoto, T, Rao, Z. | | Deposit date: | 2010-06-13 | | Release date: | 2010-07-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the human CNOT6L nuclease domain reveals strict poly(A) substrate specificity.
Embo J., 2010
|
|
3NGN
 
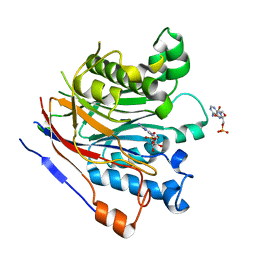 | | Crystal structure of the human CNOT6L nuclease domain in complex with AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, CCR4-NOT transcription complex subunit 6-like | | Authors: | Wang, H, Morita, M, Yang, W, Bartlam, M, Yamamoto, T, Rao, Z. | | Deposit date: | 2010-06-12 | | Release date: | 2010-07-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the human CNOT6L nuclease domain reveals strict poly(A) substrate specificity.
Embo J., 2010
|
|
1CGL
 
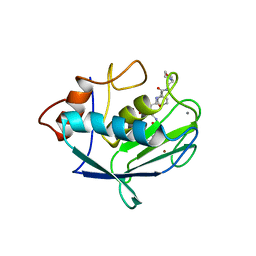 | | Structure of the catalytic domain of fibroblast collagenase complexed with an inhibitor | | Descriptor: | CALCIUM ION, FIBROBLAST COLLAGENASE, N-[(1S)-3-{[(benzyloxy)carbonyl]amino}-1-carboxypropyl]-L-leucyl-N-(2-morpholin-4-ylethyl)-L-phenylalaninamide, ... | | Authors: | Lovejoy, B, Cleasby, A, Hassell, A.M, Longley, K, Luther, M.A, Weigl, D, Mcgeehan, G, Mcelroy, A.B, Drewry, D, Lambert, M.H, Jordan, S.R. | | Deposit date: | 1993-11-17 | | Release date: | 1995-02-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the catalytic domain of fibroblast collagenase complexed with an inhibitor.
Science, 263, 1994
|
|
7PKK
 
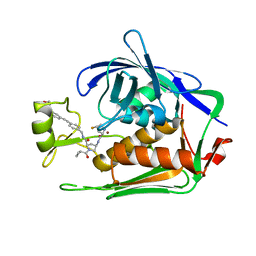 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2R,4R)-1-cyclopropylcarbonyl-4-[[(2S,4S)-4-fluoranylpyrrolidin-2-yl]carbonylamino]-N-[[4-[2-[4-(morpholin-4-ylmethyl)phenyl]ethynyl]phenyl]methyl]pyrrolidine-2-carboxamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-25 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
7PK8
 
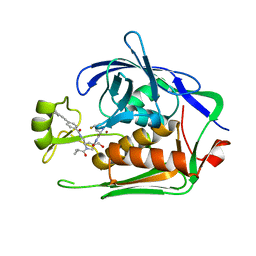 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2S,4S)-4-fluoranyl-N-[(3R,5R)-5-[[[4-[2-(4-methylphenyl)ethynyl]phenyl]carbonylamino]methyl]-1-(2-methylpropanoyl)pyrrolidin-3-yl]pyrrolidine-2-carboxamide, 1,2-ETHANEDIOL, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ... | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-25 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
7FJB
 
 | | KpAckA (PduW) with AMPPNP, sodium acetate complex structure | | Descriptor: | ACETATE ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Probable propionate kinase, ... | | Authors: | Wenyue, W, Zhang, Q, Bartlam, M. | | Deposit date: | 2021-08-03 | | Release date: | 2022-08-10 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | KpAckA (PduW) with AMPPNP, sodium acetate complex structure
To Be Published
|
|
7PHN
 
 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2~{R},4~{R})-4-[[(2~{S},4~{S})-4-fluoranylpyrrolidin-2-yl]carbonylamino]-1-(2-methylpropanoyl)-~{N}-[[4-(2-phenylethynyl)phenyl]methyl]pyrrolidine-2-carboxamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-17 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
4MFI
 
 | | Crystal structure of Mycobacterium tuberculosis UgpB | | Descriptor: | Sn-glycerol-3-phosphate ABC transporter substrate-binding protein UspB | | Authors: | Jiang, D, Bartlam, M, Rao, Z. | | Deposit date: | 2013-08-28 | | Release date: | 2014-07-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural analysis of Mycobacterium tuberculosis ATP-binding cassette transporter subunit UgpB reveals specificity for glycerophosphocholine
Febs J., 281, 2014
|
|
7PJG
 
 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2R,4R)-N-[[4-(4-cyclopropylbuta-1,3-diynyl)phenyl]methyl]-1-(2-methylpropanoyl)-4-[[(2S,4R)-4-oxidanylpyrrolidin-2-yl]carbonylamino]pyrrolidine-2-carboxamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-24 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
7PHJ
 
 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2~{R},4~{R})-4-[[(2~{S},4~{S})-4-fluoranylpyrrolidin-2-yl]carbonylamino]-1-(2-methylpropanoyl)-~{N}-[[4-(2-phenylethynyl)phenyl]methyl]pyrrolidine-2-carboxamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-17 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
7PJ2
 
 | | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria | | Descriptor: | (2R,4R)-4-[[(2S,4S)-4-fluoranylpyrrolidin-2-yl]carbonylamino]-1-(2-methylpropanoyl)-N-[[4-[2-[4-(morpholin-4-ylmethyl)phenyl]ethynyl]phenyl]methyl]pyrrolidine-2-carboxamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Ryan, M.D, Pallin, T.D, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2021-08-23 | | Release date: | 2022-09-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | LpxC Inhibitors With Fluoroproline As A Novel Zinc-Binding Group Can Serve As A Novel Class of Antibiotic With Activity Against Multidrug-Resistant Gram-Negative Bacteria
To Be Published
|
|
