6K96
 
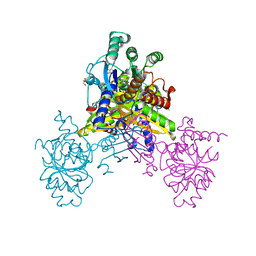 | | Crystal structure of Ari2 | | Descriptor: | Five-membered-cyclitol-phosphate synthase, GLYCEROL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Miyanaga, A, Tsunoda, T, Kudo, F, Eguchi, T. | | Deposit date: | 2019-06-14 | | Release date: | 2019-12-25 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Stereochemistry in the Reaction of themyo-Inositol Phosphate Synthase Ortholog Ari2 during Aristeromycin Biosynthesis.
Biochemistry, 58, 2019
|
|
6J39
 
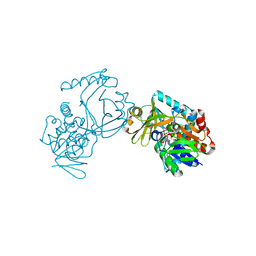 | | Crystal structure of CmiS2 with inhibitor | | Descriptor: | (3R)-3-[(carboxymethyl)sulfanyl]nonanoic acid, FAD-dependent glycine oxydase, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Kawasaki, D, Chisuga, T, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2019-01-04 | | Release date: | 2019-06-12 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structural Analysis of the Glycine Oxidase Homologue CmiS2 Reveals a Unique Substrate Recognition Mechanism for Formation of a beta-Amino Acid Starter Unit in Cremimycin Biosynthesis.
Biochemistry, 58, 2019
|
|
6K97
 
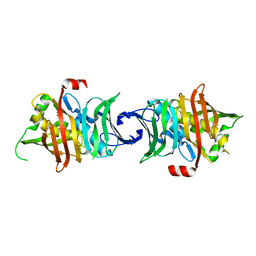 | | Crystal structure of fusion DH domain | | Descriptor: | Fusion DH, SULFATE ION | | Authors: | Kawasaki, D, Miyanaga, A, Chisuga, T, Kudo, F, Eguchi, T. | | Deposit date: | 2019-06-14 | | Release date: | 2019-11-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Functional and Structural Analyses of the Split-Dehydratase Domain in the Biosynthesis of Macrolactam Polyketide Cremimycin.
Biochemistry, 58, 2019
|
|
7F2R
 
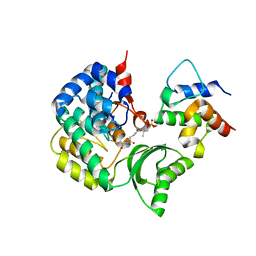 | | Crystal structure of VinK-VinL covalent complex formed with a pantetheineamide cross-linking probe | | Descriptor: | Acyl-carrier-protein, Malonyl-CoA-[acyl-carrier-protein] transacylase, N-[2-(acetylamino)ethyl]-N~3~-[(2R)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-beta-alaninamide | | Authors: | Miyanaga, A, Ouchi, R, Kudo, F, Eguchi, T. | | Deposit date: | 2021-06-14 | | Release date: | 2021-09-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Complex structure of the acyltransferase VinK and the carrier protein VinL with a pantetheine cross-linking probe.
Acta Crystallogr.,Sect.F, 77, 2021
|
|
7CL2
 
 | | The crystal structure of KanJ | | Descriptor: | GLYCEROL, Kanamycin B dioxygenase, NICKEL (II) ION, ... | | Authors: | Kitayama, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2020-07-20 | | Release date: | 2021-01-13 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis.
Chembiochem, 22, 2021
|
|
7CL5
 
 | | The crystal structure of KanJ in complex with kanamycin B and N-oxalylglycine | | Descriptor: | (1R,2S,3S,4R,6S)-4,6-DIAMINO-3-[(3-AMINO-3-DEOXY-ALPHA-D-GLUCOPYRANOSYL)OXY]-2-HYDROXYCYCLOHEXYL 2,6-DIAMINO-2,6-DIDEOXY-ALPHA-D-GLUCOPYRANOSIDE, Kanamycin B dioxygenase, N-OXALYLGLYCINE, ... | | Authors: | Kitayama, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2020-07-20 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis.
Chembiochem, 22, 2021
|
|
7CL4
 
 | | The crystal structure of KanJ in complex with N-oxalylglycine | | Descriptor: | Kanamycin B dioxygenase, N-OXALYLGLYCINE, NICKEL (II) ION | | Authors: | Kitayama, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2020-07-20 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis.
Chembiochem, 22, 2021
|
|
7CL6
 
 | | The crystal structure of KanJ in complex with neamine and N-oxalylglycine | | Descriptor: | (1R,2R,3S,4R,6S)-4,6-diamino-2,3-dihydroxycyclohexyl 2,6-diamino-2,6-dideoxy-alpha-D-glucopyranoside, Kanamycin B dioxygenase, N-OXALYLGLYCINE, ... | | Authors: | Kitayama, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2020-07-20 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis.
Chembiochem, 22, 2021
|
|
7CL3
 
 | | The crystal structure of KanJ in complex with kanamycin B | | Descriptor: | (1R,2S,3S,4R,6S)-4,6-DIAMINO-3-[(3-AMINO-3-DEOXY-ALPHA-D-GLUCOPYRANOSYL)OXY]-2-HYDROXYCYCLOHEXYL 2,6-DIAMINO-2,6-DIDEOXY-ALPHA-D-GLUCOPYRANOSIDE, Kanamycin B dioxygenase, NICKEL (II) ION, ... | | Authors: | Kitayama, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2020-07-20 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Stepwise Post-glycosylation Modification of Sugar Moieties in Kanamycin Biosynthesis.
Chembiochem, 22, 2021
|
|
4ZM4
 
 | | Complex structure of PctV K276R mutant with PMP and 3-dehydroshkimate | | Descriptor: | (3E,4R,5R)-4,5-dihydroxy-3-{[(Z)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4(1H)-ylidene}methyl]imino}cyclohex-1-ene-1-carboxylic acid, Aminotransferase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Hirayama, A, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2015-05-02 | | Release date: | 2015-10-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Mechanism-Based Trapping of the Quinonoid Intermediate by Using the K276R Mutant of PLP-Dependent 3-Aminobenzoate Synthase PctV in the Biosynthesis of Pactamycin.
Chembiochem, 16, 2015
|
|
3WV5
 
 | | Complex structure of VinN with 3-methylaspartate | | Descriptor: | (2S,3S)-3-methyl-aspartic acid, Non-ribosomal peptide synthetase | | Authors: | Miyanaga, A, Cieslak, J, Shinohara, Y, Kudo, F, Eguchi, T. | | Deposit date: | 2014-05-15 | | Release date: | 2014-10-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure of the adenylation enzyme VinN reveals a unique beta-amino acid recognition mechanism
J.Biol.Chem., 289, 2014
|
|
3WMR
 
 | | Crystal structure of VinJ | | Descriptor: | 1-ETHOXY-2-(2-ETHOXYETHOXY)ETHANE, GLYCEROL, Proline iminopeptidase | | Authors: | Shinohara, Y, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2013-11-22 | | Release date: | 2014-02-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The crystal structure of the amidohydrolase VinJ shows a unique hydrophobic tunnel for its interaction with polyketide substrates
Febs Lett., 588, 2014
|
|
3WVN
 
 | | Complex structure of VinN with L-aspartate | | Descriptor: | ASPARTIC ACID, Non-ribosomal peptide synthetase | | Authors: | Miyanaga, A, Cieslak, J, Shinohara, Y, Kudo, F, Eguchi, T. | | Deposit date: | 2014-05-30 | | Release date: | 2014-10-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure of the adenylation enzyme VinN reveals a unique beta-amino acid recognition mechanism
J.Biol.Chem., 289, 2014
|
|
4ZM3
 
 | | Crystal structure of PLP-Dependent 3-Aminobenzoate Synthase PctV wild-type | | Descriptor: | Aminotransferase, DI(HYDROXYETHYL)ETHER, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Hirayama, A, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2015-05-02 | | Release date: | 2015-10-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Mechanism-Based Trapping of the Quinonoid Intermediate by Using the K276R Mutant of PLP-Dependent 3-Aminobenzoate Synthase PctV in the Biosynthesis of Pactamycin.
Chembiochem, 16, 2015
|
|
3WV4
 
 | | Crystal structure of VinN | | Descriptor: | Non-ribosomal peptide synthetase | | Authors: | Miyanaga, A, Cieslak, J, Shinohara, Y, Kudo, F, Eguchi, T. | | Deposit date: | 2014-05-15 | | Release date: | 2014-10-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The crystal structure of the adenylation enzyme VinN reveals a unique beta-amino acid recognition mechanism
J.Biol.Chem., 289, 2014
|
|
