8AMM
 | | Crystal structure of AUGUGGCAU duplex with cesium ions | | Descriptor: | CESIUM ION, RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3') | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
8AMG
 
 | | Crystal structure of AUGUGGCAU duplex with barium ions (model 1) | | Descriptor: | BARIUM ION, RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3') | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
8AMI
 
 | | Crystal structure of AUGUGGCAU duplex with barium ions (model 2) | | Descriptor: | BARIUM ION, RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3'), SODIUM ION | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
8AML
 
 | | Crystal structure of AUGUGGCAU duplex with cadmium ions | | Descriptor: | CADMIUM ION, RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3') | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
8AMJ
 
 | | Crystal structure of unliganded AUGUGGCAU duplex | | Descriptor: | ACETATE ION, RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3'), SULFATE ION | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
8AMK
 
 | |
8AMN
 
 | | Crystal structure of AUGUGGCAU duplex with strontium ions | | Descriptor: | RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3'), STRONTIUM ION | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
9EN6
 
 | | Crystal structure of RNA G2C4 repeats - native model pH 6.5 | | Descriptor: | MAGNESIUM ION, RNA (5'-R(*GP*GP*CP*CP*CP*C)-3') | | Authors: | Mateja-Pluta, M, Kiliszek, A. | | Deposit date: | 2024-03-12 | | Release date: | 2024-05-01 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (0.918 Å) | | Cite: | Antisense RNA C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms a triplex-like structure and binds small synthetic ligand.
Nucleic Acids Res., 52, 2024
|
|
8QMI
 
 | | Crystal structure of RNA G2C4 repeats - native model | | Descriptor: | MAGNESIUM ION, RNA (5'-R(*GP*GP*CP*CP*CP*C)-3') | | Authors: | Blaszczyk, L, Kiliszek, A. | | Deposit date: | 2023-09-22 | | Release date: | 2024-05-01 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Antisense RNA C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms a triplex-like structure and binds small synthetic ligand.
Nucleic Acids Res., 52, 2024
|
|
4L3K
 
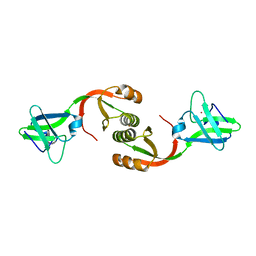 | | Crystal structure of Sporosarcina pasteurii UreE bound to Ni2+ and Zn2+ | | Descriptor: | NICKEL (II) ION, Urease accessory protein UreE, ZINC ION | | Authors: | Zambelli, B, Banaszak, K, Merloni, A, Kiliszek, A, Rypniewski, W.R, Ciurli, S. | | Deposit date: | 2013-06-06 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Selectivity of Ni(II) and Zn(II) binding to Sporosarcina pasteurii UreE, a metallochaperone in the urease assembly: a calorimetric and crystallographic study.
J.Biol.Inorg.Chem., 18, 2013
|
|
6IA2
 
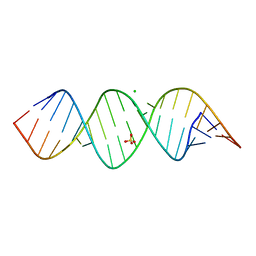 | | Crystal structure of a self-complementary RNA duplex recognized by Com | | Descriptor: | CHLORIDE ION, RNA (5'-R(*AP*GP*AP*GP*AP*AP*CP*CP*CP*GP*GP*AP*GP*UP*UP*CP*CP*CP*U)-3'), SULFATE ION | | Authors: | Nowacka, M, Fernandes, H, Kiliszek, A, Bernat, A, Lach, G, Bujnicki, J.M. | | Deposit date: | 2018-11-26 | | Release date: | 2019-03-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Specific interaction of zinc finger protein Com with RNA and the crystal structure of a self-complementary RNA duplex recognized by Com.
Plos One, 14, 2019
|
|
