8YXK
 
 | |
8YXN
 
 | |
1ZZE
 
 | | X-ray Structure of NADPH-dependent Carbonyl Reductase from Sporobolomyces salmonicolor | | 分子名称: | Aldehyde reductase II, SULFATE ION | | 著者 | Kamitori, S, Iguchi, A, Ohtaki, A, Yamada, M, Kita, K. | | 登録日 | 2005-06-14 | | 公開日 | 2005-09-06 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | X-ray Structures of NADPH-dependent Carbonyl Reductase from Sporobolomyces salmonicolor Provide Insights into Stereoselective Reductions of Carbonyl Compounds
J.Mol.Biol., 352, 2005
|
|
7WRR
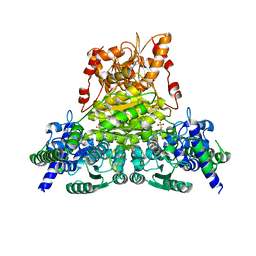 
 | |
7WRT
 
 | |
8GSY
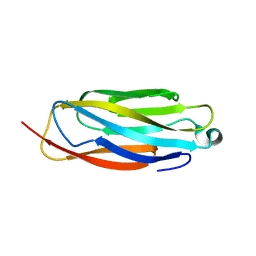 
 | |
8GSX
 
 | |
8WL7
 | |
8WL9
 | |
8WL5
 
 | |
8WLB
 | |
6L67
 
 | |
6L6B
 
 | | X-ray structure of human galectin-10 in complex with L-fucose | | 分子名称: | Galectin-10, beta-L-fucopyranose | | 著者 | Kamitori, S. | | 登録日 | 2019-10-28 | | 公開日 | 2020-03-04 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.802 Å) | | 主引用文献 | Structures of human galectin-10/monosaccharide complexes demonstrate potential of monosaccharides as effectors in forming Charcot-Leyden crystals.
Biochem.Biophys.Res.Commun., 2020
|
|
6L6C
 
 | |
6L64
 
 | | X-ray structure of human galectin-10 in complex with D-glucose | | 分子名称: | Galectin-10, beta-D-glucopyranose | | 著者 | Kamitori, S. | | 登録日 | 2019-10-28 | | 公開日 | 2020-03-04 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.08 Å) | | 主引用文献 | Structures of human galectin-10/monosaccharide complexes demonstrate potential of monosaccharides as effectors in forming Charcot-Leyden crystals.
Biochem.Biophys.Res.Commun., 2020
|
|
6L6A
 
 | | X-ray structure of human galectin-10 in complex with D-mannose | | 分子名称: | Galectin-10, beta-D-mannopyranose | | 著者 | Kamitori, S. | | 登録日 | 2019-10-28 | | 公開日 | 2020-03-04 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.81 Å) | | 主引用文献 | Structures of human galectin-10/monosaccharide complexes demonstrate potential of monosaccharides as effectors in forming Charcot-Leyden crystals.
Biochem.Biophys.Res.Commun., 2020
|
|
1BVZ
 
 | | ALPHA-AMYLASE II (TVAII) FROM THERMOACTINOMYCES VULGARIS R-47 | | 分子名称: | PROTEIN (ALPHA-AMYLASE II) | | 著者 | Kamitori, S, Kondo, S, Okuyama, K, Yokota, T, Shimura, Y, Tonozuka, T, Sakano, Y. | | 登録日 | 1998-09-22 | | 公開日 | 1999-03-02 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Crystal structure of Thermoactinomyces vulgaris R-47 alpha-amylase II (TVAII) hydrolyzing cyclodextrins and pullulan at 2.6 A resolution.
J.Mol.Biol., 287, 1999
|
|
1JI2
 
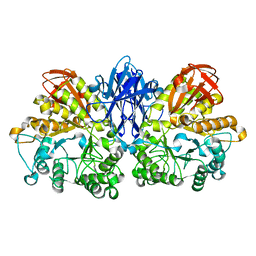 | | Improved X-ray Structure of Thermoactinomyces vulgaris R-47 alpha-Amylase 2 | | 分子名称: | ALPHA-AMYLASE II, CALCIUM ION | | 著者 | Kamitori, S, Abe, A, Ohtaki, A, Kaji, A, Tonozuka, T, Sakano, Y. | | 登録日 | 2001-06-28 | | 公開日 | 2002-06-05 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal structures and structural comparison of Thermoactinomyces vulgaris R-47 alpha-amylase 1 (TVAI) at 1.6 A resolution and alpha-amylase 2 (TVAII) at 2.3 A resolution.
J.Mol.Biol., 318, 2002
|
|
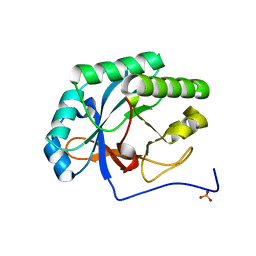
4KRU
 
 | |
4KRT
 
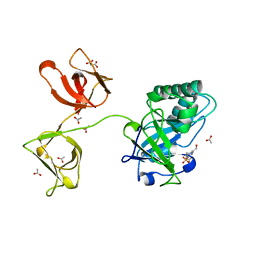 | |
2GS9
 
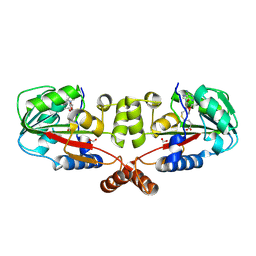 | | Crystal structure of TT1324 from Thermus thermophilis HB8 | | 分子名称: | FORMIC ACID, Hypothetical protein TT1324, S-ADENOSYL-L-HOMOCYSTEINE | | 著者 | Kamitori, S, Abe, A, Ebihara, A, Kanagawa, M, Nakagawa, N, Kuroishi, C, Agari, Y, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 2006-04-25 | | 公開日 | 2007-03-13 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Crystal structure of TT1324 from Thermus thermophilis HB8
To be Published
|
|
6IXZ
 | |
6IXY
 
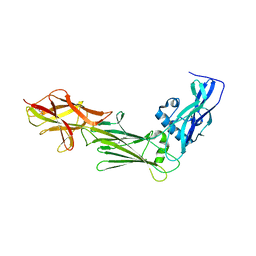 | |
1JI1
 
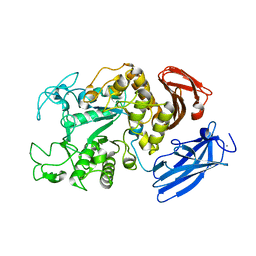 | | Crystal Structure Analysis of Thermoactinomyces vulgaris R-47 alpha-Amylase 1 | | 分子名称: | ALPHA-AMYLASE I, CALCIUM ION | | 著者 | Kamitori, S, Abe, A, Ohtaki, A, Kaji, A, Tonozuka, T, Sakano, Y. | | 登録日 | 2001-06-28 | | 公開日 | 2002-06-05 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Crystal structures and structural comparison of Thermoactinomyces vulgaris R-47 alpha-amylase 1 (TVAI) at 1.6 A resolution and alpha-amylase 2 (TVAII) at 2.3 A resolution.
J.Mol.Biol., 318, 2002
|
|
173D
 
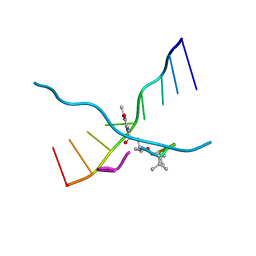 | | MULTIPLE BINDING MODES OF ANTICANCER DRUG ACTINOMYCIN D: X-RAY, MOLECULAR MODELING, AND SPECTROSCOPIC STUDIES OF D(GAAGCTTC)2-ACTINOMYCIN D COMPLEXES AND ITS HOST DNA | | 分子名称: | ACTINOMYCIN D, DNA (5'-D(*GP*AP*AP*GP*CP*TP*TP*C)-3') | | 著者 | Kamitori, S, Takusagawa, F. | | 登録日 | 1994-04-18 | | 公開日 | 1994-10-15 | | 最終更新日 | 2024-07-10 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Multiple Binding Modes of Anticancer Drug Actinomycin D: X-Ray, Molecular Modeling, and Spectroscopic Studies of D(Gaagcttc)2-Actinomycin D Complexes and its Host DNA
J.Am.Chem.Soc., 116, 1994
|
|
