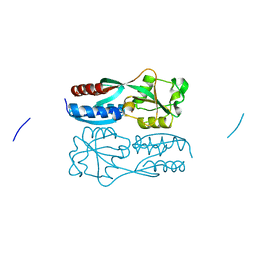
5B7D
 
 | |
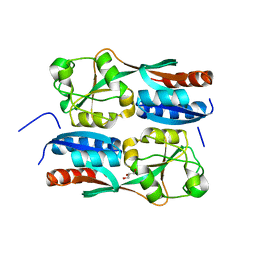
5B70
 
 | |
5YDV
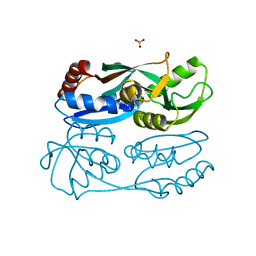 
 | | Regulatory domain of HypT from Salmonella typhimurium complexed with HOCl (HOCl-bound form) | | 分子名称: | Cell density-dependent motility repressor, SULFATE ION, hypochlorous acid | | 著者 | Jo, I, Hong, S, Ahn, J, Ha, N.C. | | 登録日 | 2017-09-14 | | 公開日 | 2018-11-28 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.752 Å) | | 主引用文献 | Structural basis for HOCl recognition and regulation mechanisms of HypT, a hypochlorite-specific transcriptional regulator.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
5YEZ
 
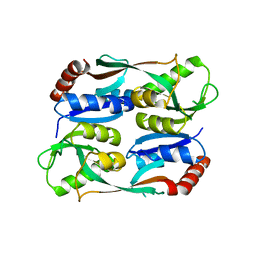 | | Regulatory domain of HypT M206Q mutant from Salmonella typhimurium | | 分子名称: | Cell density-dependent motility repressor | | 著者 | Jo, I, Hong, S, Ahn, J, Ha, N.C. | | 登録日 | 2017-09-20 | | 公開日 | 2018-10-03 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structural basis for HOCl recognition and regulation mechanisms of HypT, a hypochlorite-specific transcriptional regulator.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
5YDW
 
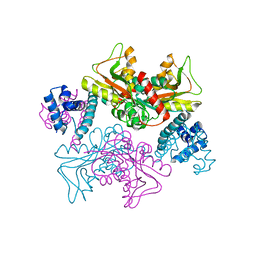 | | Full-length structure of HypT from Salmonella typhimuriuma (hypochlorite-specific LysR-type transcriptional regulator) | | 分子名称: | Cell density-dependent motility repressor | | 著者 | Jo, I, Hong, S, Ahn, J, Ha, N.C. | | 登録日 | 2017-09-15 | | 公開日 | 2018-11-28 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | Structural basis for HOCl recognition and regulation mechanisms of HypT, a hypochlorite-specific transcriptional regulator.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
5YER
 
 | | Regulatory domain of HypT from Salmonella typhimurium (Bromide ion-bound) | | 分子名称: | BROMIDE ION, Cell density-dependent motility repressor, SULFATE ION | | 著者 | Jo, I, Hong, S, Ahn, J, Ha, N.C. | | 登録日 | 2017-09-19 | | 公開日 | 2018-11-28 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.301 Å) | | 主引用文献 | Structural basis for HOCl recognition and regulation mechanisms of HypT, a hypochlorite-specific transcriptional regulator.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
5YDO
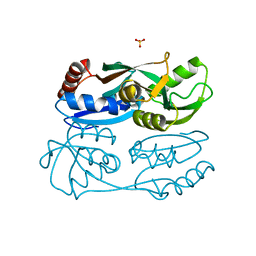 
 | | Regulatory domain of HypT from Salmonella typhimurium (apo-form) | | 分子名称: | Cell density-dependent motility repressor, SULFATE ION | | 著者 | Jo, I, Hong, S, Ahn, J, Ha, N.C. | | 登録日 | 2017-09-13 | | 公開日 | 2018-11-28 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural basis for HOCl recognition and regulation mechanisms of HypT, a hypochlorite-specific transcriptional regulator.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
5X0V
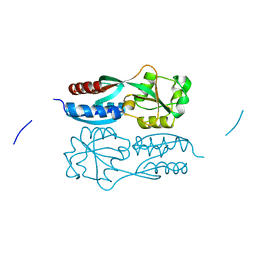 
 | |
5X0Q
 
 | | OxyR2 E204G variant (Cl-bound) from Vibrio vulnificus | | 分子名称: | CHLORIDE ION, CITRIC ACID, LysR family transcriptional regulator | | 著者 | Jo, I, Ha, N.-C. | | 登録日 | 2017-01-23 | | 公開日 | 2017-03-15 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | The hydrogen peroxide hypersensitivity of OxyR2 in Vibrio vulnificus depends on conformational constraints
J. Biol. Chem., 292, 2017
|
|
4Y0M
 
 | |
4XWS
 
 | |
5B7H
 
 | |
5HFG
 
 | |
5HFI
 
 | |
4X6G
 
 | |
4PWO
 
 | | Crystal structure of DsbA from the Gram positive bacterium Corynebacterium diphtheriae | | 分子名称: | DsbA, GLYCEROL | | 著者 | Um, S.H, Kim, J.S, Jiao, L, Yoon, B.Y, Jo, I, Ha, N.C. | | 登録日 | 2014-03-21 | | 公開日 | 2015-03-25 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.52 Å) | | 主引用文献 | Crystal structure and biochemical characterization of DsbA from the Gram positive bacterium Corynebacterium diphtheriae
To be Published
|
|
4PWP
 
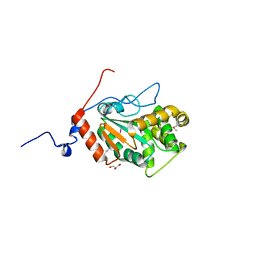 | | Crystal structure of DsbA from the Gram positive bacterium Corynebacterium diphtheriae | | 分子名称: | DsbA, GLYCEROL | | 著者 | Um, S.H, Kim, J.S, Jiao, L, Yoon, B.Y, Jo, I, Ha, N.C. | | 登録日 | 2014-03-21 | | 公開日 | 2015-03-25 | | 実験手法 | X-RAY DIFFRACTION (1.803 Å) | | 主引用文献 | Crystal structure and biochemical characterization of DsbA from the Gram positive bacterium Corynebacterium diphtheriae
To be Published
|
|
5Y9S
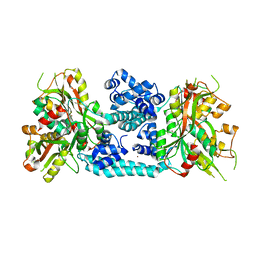 
 | | Crystal structure of VV2_1132, a LysR family transcriptional regulator | | 分子名称: | BROMIDE ION, VV2_1132 | | 著者 | Jang, Y, Hong, S, Jo, I, Ahn, J, Ha, N.C. | | 登録日 | 2017-08-28 | | 公開日 | 2018-03-28 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.199 Å) | | 主引用文献 | A Novel Tetrameric Assembly Configuration in VV2_1132, a LysR-Type Transcriptional Regulator inVibrio vulnificus
Mol. Cells, 41, 2018
|
|
5ZQS
 
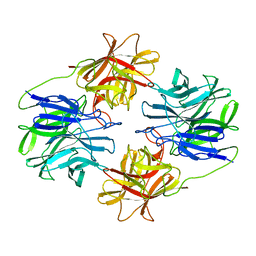 | |
5ZQJ
 
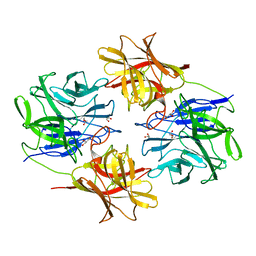 | |
5ZQX
 
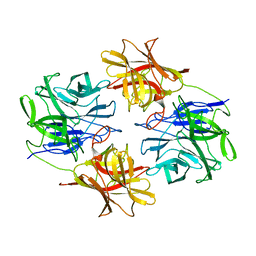 | |
7X5D
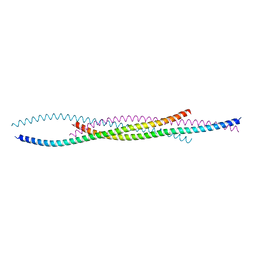 
 | |
5WU7
 
 | |
6JLB
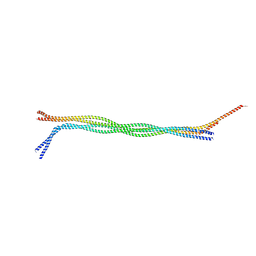 
 | |
5X15
 
 | |
