4KP1
 
 | | Crystal structure of IPM isomerase large subunit from methanococcus jannaschii (MJ0499) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2,4-dimethylpentane-2,4-diol, Isopropylmalate/citramalate isomerase large subunit, ... | | Authors: | Hwang, K.Y, Lee, E.H. | | Deposit date: | 2013-05-13 | | Release date: | 2014-04-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural characterization and comparison of the large subunits of IPM isomerase and homoaconitase from Methanococcus jannaschii
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3DNM
 
 | |
4LWM
 
 | |
3E6E
 
 | | Crystal structure of Alanine racemase from E.faecalis complex with cycloserine | | Descriptor: | Alanine racemase, D-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-N,O-CYCLOSERYLAMIDE | | Authors: | Hwang, K.Y, Priyadarshi, A, Lee, E.H, Sung, M.W. | | Deposit date: | 2008-08-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the alanine racemase from Enterococcus faecalis.
Biochim.Biophys.Acta, 1794, 2009
|
|
4LWK
 
 | |
3Q4R
 
 | |
3Q4N
 
 | |
3Q4Q
 
 | |
3Q4O
 
 | |
3H17
 
 | | Crystal structure of EstE5-PMSF (I) | | Descriptor: | Esterase/lipase, phenylmethanesulfonic acid | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2009-04-11 | | Release date: | 2009-04-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of an HSL-homolog EstE5 complex with PMSF reveals a unique configuration that inhibits the nucleophile Ser144 in catalytic triads.
Biochem.Biophys.Res.Commun., 389, 2009
|
|
3H18
 
 | | Crystal structure of EstE5-PMSF (II) | | Descriptor: | Esterase/lipase, phenylmethanesulfonic acid | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2009-04-11 | | Release date: | 2009-04-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The crystal structure of an HSL-homolog EstE5 complex with PMSF reveals a unique configuration that inhibits the nucleophile Ser144 in catalytic triads.
Biochem.Biophys.Res.Commun., 389, 2009
|
|
2MJP
 
 | | STRUCTURE-BASED IDENTIFICATION OF THE BIOCHEMICAL FUNCTION OF A HYPOTHETICAL PROTEIN FROM METHANOCOCCUS JANNASCHII:MJ0226 | | Descriptor: | PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, PYROPHOSPHATASE | | Authors: | Hwang, K.Y, Chung, J.H, Han, Y.S, Kim, S.H, Cho, Y, Berkeley Structural Genomics Center (BSGC) | | Deposit date: | 1999-01-27 | | Release date: | 2000-01-28 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based identification of a novel NTPase from Methanococcus jannaschii.
Nat.Struct.Biol., 6, 1999
|
|
3II1
 
 | |
2NX8
 
 | |
3G6N
 
 | | Crystal structure of an EfPDF complex with Met-Ala-Ser | | Descriptor: | FE (III) ION, Peptide deformylase, SODIUM ION, ... | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2009-02-07 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of an EfPDF complex with Met-Ala-Ser based on crystallographic packing.
Biochem.Biophys.Res.Commun., 381, 2009
|
|
5HXY
 
 | | Crystal structure of XerA recombinase | | Descriptor: | PHOSPHATE ION, Tyrosine recombinase XerA | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2016-01-31 | | Release date: | 2017-02-01 | | Last modified: | 2020-02-19 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of Thermoplasma acidophilum XerA recombinase shows large C-shape clamp conformation and cis-cleavage mode for nucleophilic tyrosine
FEBS Lett., 590, 2016
|
|
5HZT
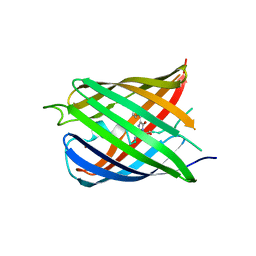 
 | | Crystal structure of Dronpa-Cu2+ | | Descriptor: | COPPER (II) ION, Fluorescent protein Dronpa | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2016-02-03 | | Release date: | 2017-03-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Crystal structures of Dronpa complexed with quenchable metal ions provide insight into metal biosensor development
FEBS Lett., 590, 2016
|
|
5HZU
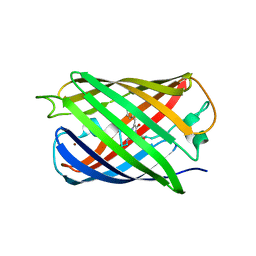 
 | | Crystal structure of Dronpa-Ni2+ | | Descriptor: | Fluorescent protein Dronpa, NICKEL (II) ION | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2016-02-03 | | Release date: | 2017-03-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Crystal structures of Dronpa complexed with quenchable metal ions provide insight into metal biosensor development
FEBS Lett., 590, 2016
|
|
5HZS
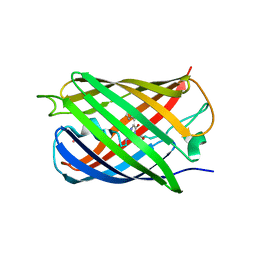 
 | | Crystal structure of Dronpa-Co2+ | | Descriptor: | COBALT (II) ION, Fluorescent protein Dronpa | | Authors: | Hwang, K.Y, Nam, K.H. | | Deposit date: | 2016-02-03 | | Release date: | 2017-03-15 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Crystal structures of Dronpa complexed with quenchable metal ions provide insight into metal biosensor development
FEBS Lett., 590, 2016
|
|
6A4V
 
 | | Open Reading frame 49 | | Descriptor: | 49 protein | | Authors: | Hwang, K.Y, Song, M.J, Kim, J.S, Cheong, W.C. | | Deposit date: | 2018-06-21 | | Release date: | 2019-06-26 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based mechanism of action of a viral poly(ADP-ribose) polymerase 1-interacting protein facilitating virus replication.
Iucrj, 5, 2018
|
|
6AGV
 
 | | Crystal structure of apo mouse MsrA | | Descriptor: | GLYCEROL, Mitochondrial peptide methionine sulfoxide reductase | | Authors: | Hwang, K.Y, Kim, J.S. | | Deposit date: | 2018-08-14 | | Release date: | 2019-08-28 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Crystal structure of apo mouse MsrA
To Be Published
|
|
3G9U
 
 | |
3G9Z
 
 | |
3H19
 
 | |
3GVY
 
 | |
