3GD0
 
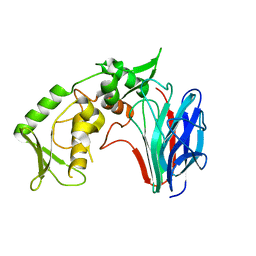 | | Crystal structure of laminaripentaose-producing beta-1,3-glucanase | | Descriptor: | Laminaripentaose-producing beta-1,3-guluase (LPHase) | | Authors: | Wu, H.M, Hsu, M.T, Liu, S.W, Lai, C.C, Li, Y.K, Wang, W.C. | | Deposit date: | 2009-02-23 | | Release date: | 2009-07-28 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Structure, mechanistic action, and essential residues of a GH-64 enzyme, laminaripentaose-producing beta-1,3-glucanase.
J.Biol.Chem., 284, 2009
|
|
4QI1
 
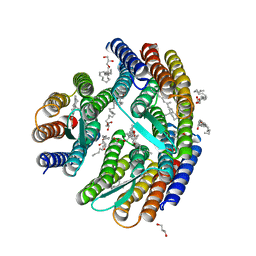 | | Crystal structure of H. walsbyi bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin-I, GLYCEROL, RETINAL, ... | | Authors: | Wang, A.H.J, Hsu, M.F, Yang, C.S, Fu, H.Y. | | Deposit date: | 2014-05-30 | | Release date: | 2015-07-29 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and Functional Studies of a Newly Grouped Haloquadratum walsbyi Bacteriorhodopsin Reveal the Acid-resistant Light-driven Proton Pumping Activity.
J. Biol. Chem., 290, 2015
|
|
4QID
 
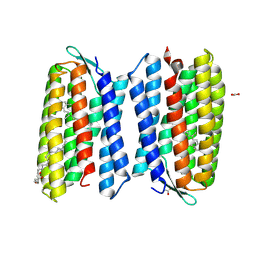 | | Crystal structure of Haloquadratum walsbyi bacteriorhodopsin | | Descriptor: | ACETATE ION, Bacteriorhodopsin-I, RETINAL, ... | | Authors: | Wang, A.H.J, Hsu, M.F, Yang, C.S, Fu, H.Y. | | Deposit date: | 2014-05-30 | | Release date: | 2015-07-29 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | An acid-tolerant light-driven proton pump
To be Published
|
|
1TI5
 
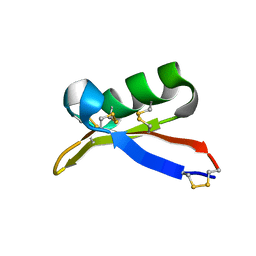 | | Solution structure of plant defensin | | Descriptor: | plant defensin | | Authors: | Liu, Y.J, Cheng, C.S, Liu, Y.N, Hsu, M.P, Chen, C.S, Lyu, P.C. | | Deposit date: | 2004-06-02 | | Release date: | 2005-07-26 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the plant defensin VrD1 from mung bean and its possible role in insecticidal activity against bruchids.
Proteins, 63, 2006
|
|
1Z10
 
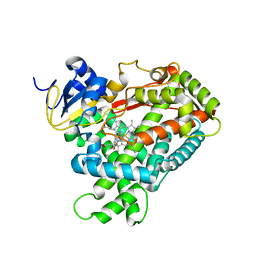 | | Crystal Structure of Human Microsomal P450 2A6 with Coumarin Bound | | Descriptor: | COUMARIN, GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Yano, J.K, Hsu, M.H, Griffin, K.J, Stout, C.D, Johnson, E.F. | | Deposit date: | 2005-03-02 | | Release date: | 2005-08-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of human microsomal cytochrome P450 2A6 complexed with coumarin and methoxsalen
Nat.Struct.Mol.Biol., 12, 2005
|
|
1Z11
 
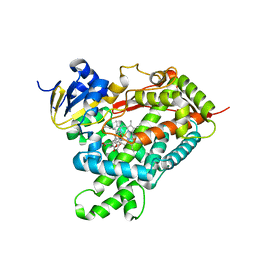 | | Crystal Structure of Human Microsomal P450 2A6 with Methoxsalen Bound | | Descriptor: | METHOXSALEN, PROTOPORPHYRIN IX CONTAINING FE, cytochrome P450, ... | | Authors: | Yano, J.K, Hsu, M.H, Griffin, K.J, Stout, C.D, Johnson, E.F. | | Deposit date: | 2005-03-02 | | Release date: | 2005-08-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structures of human microsomal cytochrome P450 2A6 complexed with coumarin and methoxsalen
Nat.Struct.Mol.Biol., 12, 2005
|
|
2GL1
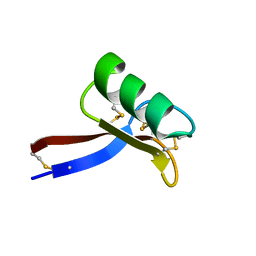 
 | | NMR solution structure of Vigna radiata Defensin 2 (VrD2) | | Descriptor: | PDF1 | | Authors: | Lin, K.F, Lee, T.R, Tsai, P.H, Hsu, M.P, Chen, C.S, Lyu, P.C. | | Deposit date: | 2006-04-04 | | Release date: | 2007-04-03 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure-based protein engineering for alpha-amylase inhibitory activity of plant defensin.
Proteins, 68, 2007
|
|
1ZJE
 
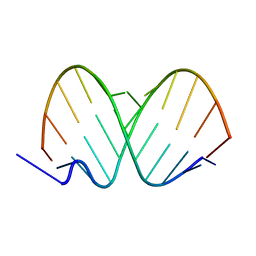 | | 12mer-spd | | Descriptor: | 5'-D(*AP*GP*GP*GP*GP*CP*GP*GP*GP*GP*CP*T)-3', 5'-D(*TP*AP*GP*CP*CP*CP*CP*GP*CP*CP*CP*C)-3' | | Authors: | Dohm, J.A, Hsu, M.H, Hwu, J.R, Huang, R.C, Moudrianakis, E.N, Lattman, E.E, Gittis, A.G. | | Deposit date: | 2005-04-28 | | Release date: | 2005-05-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Influence of Ions, Hydration, and the Transcriptional Inhibitor P4N on the Conformations of the Sp1 Binding Site.
J.Mol.Biol., 349, 2005
|
|
1ZJF
 
 | | 12mer-spd-P4N | | Descriptor: | 5'-D(*AP*GP*GP*GP*GP*CP*GP*GP*GP*GP*CP*T)-3', 5'-D(*TP*AP*GP*CP*CP*CP*CP*GP*CP*CP*CP*C)-3' | | Authors: | Dohm, J.A, Hsu, M.H, Hwu, J.R, Huang, R.C, Moudrianakis, E.N, Lattman, E.E, Gittis, A.G. | | Deposit date: | 2005-04-28 | | Release date: | 2005-05-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Influence of Ions, Hydration, and the Transcriptional Inhibitor P4N on the Conformations of the Sp1 Binding Site.
J.Mol.Biol., 349, 2005
|
|
1ZJG
 
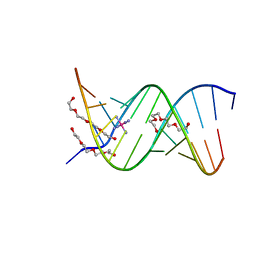 | | 13mer-co | | Descriptor: | 5'-D(*AP*TP*GP*GP*GP*GP*CP*GP*GP*GP*GP*CP*T)-3', 5'-D(*TP*AP*GP*CP*CP*CP*CP*GP*CP*CP*CP*CP*A)-3', COBALT HEXAMMINE(III), ... | | Authors: | Dohm, J.A, Hsu, M.H, Hwu, J.R, Huang, R.C, Moudrianakis, E.N, Lattman, E.E, Gittis, A.G. | | Deposit date: | 2005-04-28 | | Release date: | 2005-05-10 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Influence of Ions, Hydration, and the Transcriptional Inhibitor P4N on the Conformations of the Sp1 Binding Site.
J.Mol.Biol., 349, 2005
|
|
