1NAN
 
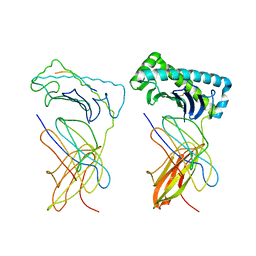 | | MCH CLASS I H-2KB MOLECULE COMPLEXED WITH PBM1 PEPTIDE | | Descriptor: | Beta-2-microglobulin, H-2 class I histocompatibility antigen, K-B alpha chain, ... | | Authors: | Reiser, J.-B, Darnault, C, Gregoire, C, Mosser, T, Mazza, G, Kearnay, A, van der Merwe, P.A, Fontecilla-Camps, J.C, Housset, D, Malissen, B. | | Deposit date: | 2002-11-28 | | Release date: | 2003-03-11 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | CDR3 loop flexibility contributes to the degeneracy of TCR recognition
Nat.Immunol., 4, 2003
|
|
2CDC
 
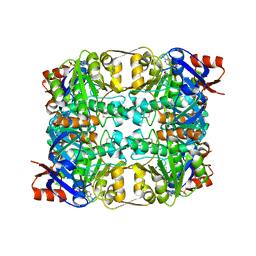 | | Sulfolobus solfataricus Glucose Dehydrogenase 1 in complex with NADP and Xylose | | Descriptor: | 1,2-ETHANEDIOL, GLUCOSE DEHYDROGENASE GLUCOSE 1-DEHYDROGENASE, DHG-1, ... | | Authors: | Milburn, C.C, Lamble, H.J, Theodossis, A, Hough, D.W, Danson, M.J, Taylor, G.L. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The structural basis of substrate promiscuity in glucose dehydrogenase from the hyperthermophilic archaeon Sulfolobus solfataricus.
J. Biol. Chem., 281, 2006
|
|
1L4I
 
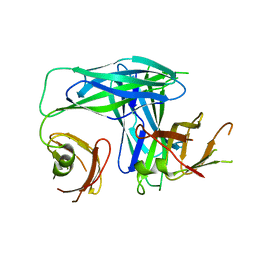 | | Crystal Structure of the Periplasmic Chaperone SfaE | | Descriptor: | SfaE PROTEIN | | Authors: | Knight, S.D, Choudhury, D, Hultgren, S, Pinkner, J, Stojanoff, V, Thompson, A. | | Deposit date: | 2002-03-05 | | Release date: | 2002-06-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the S pilus periplasmic chaperone SfaE at 2.2 A resolution.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1GUV
 
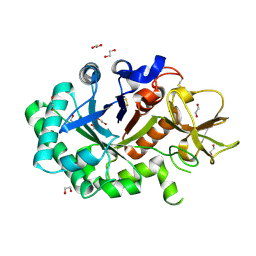 | | Structure of human chitotriosidase | | Descriptor: | 1,2-ETHANEDIOL, CHITOTRIOSIDASE | | Authors: | Von Moeller, H, Houston, D, Boot, R.G, Aerts, J.M.F.G, Van Aalten, D.M.F. | | Deposit date: | 2002-01-31 | | Release date: | 2003-03-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure of Human Chitotriosidase - Implications for Specific Inhibitor Design and Function of Mammalian Chitinase-Like Lectins
J.Biol.Chem., 277, 2002
|
|
3SG8
 
 | |
2BEN
 
 | |
1EG9
 
 | | NAPHTHALENE 1,2-DIOXYGENASE WITH INDOLE BOUND IN THE ACTIVE SITE. | | Descriptor: | FE (III) ION, FE2/S2 (INORGANIC) CLUSTER, INDOLE, ... | | Authors: | Carredano, E, Karlsson, A, Kauppi, B, Choudhury, D, Parales, R.E, Parales, J.V, Lee, K, Gibson, D.T, Eklund, H, Ramaswamy, S. | | Deposit date: | 2000-02-15 | | Release date: | 2000-05-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Substrate binding site of naphthalene 1,2-dioxygenase: functional implications of indole binding.
J.Mol.Biol., 296, 2000
|
|
2BEM
 
 | | Crystal structure of the Serratia marcescens chitin-binding protein CBP21 | | Descriptor: | 1,2-ETHANEDIOL, CBP21, SODIUM ION, ... | | Authors: | Vaaje-Kolstad, G, Houston, D.R, Eijsink, V.G.H, van Aalten, D.M.F. | | Deposit date: | 2004-11-26 | | Release date: | 2004-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal Structure and Binding Properties of the Serratia Marcescens Chitin-Binding Protein Cbp21
J.Biol.Chem., 280, 2005
|
|
4DCV
 
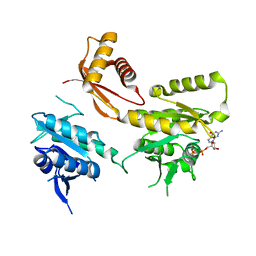 | | Crystal Structure of B. subtilis EngA in complex with GMPPCP | | Descriptor: | GTP-BINDING PROTEIN ENGA, PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER | | Authors: | Reiser, J.-B, Housset, D, Foucher, A.-E, Jault, J.-M. | | Deposit date: | 2012-01-18 | | Release date: | 2012-11-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Potassium Acts as a GTPase-Activating Element on Each Nucleotide-Binding Domain of the Essential Bacillus subtilis EngA.
Plos One, 7, 2012
|
|
2CD9
 
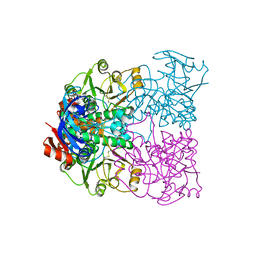 | | Sulfolobus solfataricus Glucose Dehydrogenase 1 - apo form | | Descriptor: | GLUCOSE DEHYDROGENASE, ZINC ION | | Authors: | Milburn, C.C, Lamble, H.J, Theodossis, A, Hough, D.W, Danson, M.J, Taylor, G.L. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-22 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Structural Basis of Substrate Promiscuity in Glucose Dehydrogenase from the Hyperthermophilic Archaeon Sulfolobus Solfataricus.
J.Biol.Chem., 281, 2006
|
|
2C7D
 
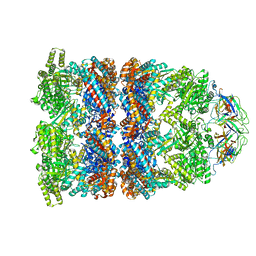 | | Fitted coordinates for GroEL-ADP7-GroES Cryo-EM complex (EMD-1181) | | Descriptor: | 10 KDA CHAPERONIN MOLECULE: GROES, PROTEIN CPN10, GROES PROTEIN, ... | | Authors: | Ranson, N.A, Clare, D.K, Farr, G.W, Houldershaw, D, Horwich, A.L, Saibil, H.R. | | Deposit date: | 2005-11-22 | | Release date: | 2006-01-25 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (8.7 Å) | | Cite: | Allosteric Signalling of ATP Hydrolysis in Groel-Groes Complexes.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2CDA
 
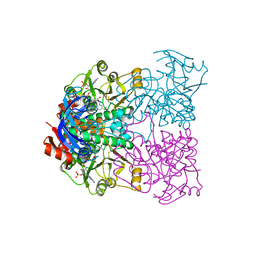 | | Sulfolobus solfataricus Glucose Dehydrogenase 1 in complex with NADP | | Descriptor: | GLUCOSE DEHYDROGENASE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ZINC ION | | Authors: | Milburn, C.C, Lamble, H.J, Theodossis, A, Hough, D.W, Danson, M.J, Taylor, G.L. | | Deposit date: | 2006-01-23 | | Release date: | 2006-03-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | The Structural Basis of Substrate Promiscuity in Glucose Dehydrogenase from the Hyperthermophilic Archaeon Sulfolobus Solfataricus.
J.Biol.Chem., 281, 2006
|
|
2C6X
 
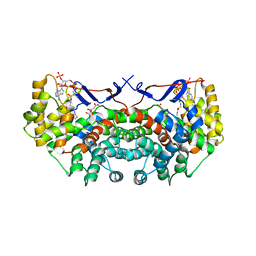 | |
3SG9
 
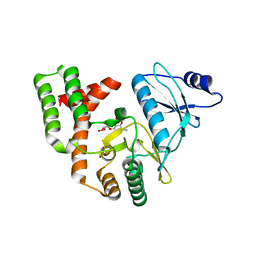 | |
1KJ3
 
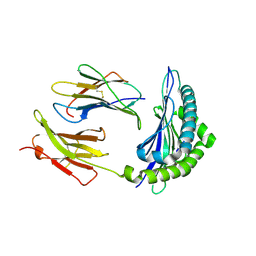 | | Mhc Class I H-2Kb molecule complexed with pKB1 peptide | | Descriptor: | BETA-2 MICROGLOBULIN, H-2KB MHC CLASS I MOLECULE ALPHA CHAIN, NATURALLY PROCESSED OCTAPEPTIDE PKB1 | | Authors: | Reiser, J.-B, Gregoire, C, Darnault, C, Mosser, T, Guimezanes, A, Schmitt-Verhulst, A.-M, Fontecilla-Camps, J.C, Mazza, G, Malissen, B, Housset, D. | | Deposit date: | 2001-12-04 | | Release date: | 2002-03-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A T cell receptor CDR3beta loop undergoes conformational changes of unprecedented magnitude upon binding to a peptide/MHC class I complex.
Immunity, 16, 2002
|
|
2ELG
 
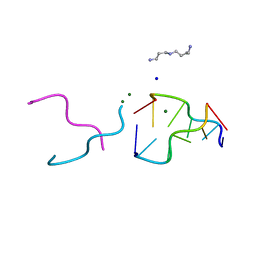 | | The rare crystallographic structure of d(CGCGCG)2: The natural spermidine molecule bound to the minor groove of left-handed Z-DNA d(CGCGCG)2 at 10 degree celsius | | Descriptor: | DNA (5'-D(*DCP*DGP*DCP*DGP*DCP*DG)-3'), MAGNESIUM ION, SODIUM ION, ... | | Authors: | Ohishi, H, Tozuka, Y, Zhou, D.Y, Ishida, T, Nakatani, K. | | Deposit date: | 2007-03-27 | | Release date: | 2008-04-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | The rare crystallographic structure of d(CGCGCG)(2): The natural spermidine molecule bound to the minor groove of left-handed Z-DNA d(CGCGCG)(2) at 10 degrees C
Biochem.Biophys.Res.Commun., 358, 2007
|
|
3TNX
 
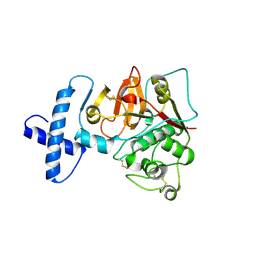 | | Structure of the precursor of a thermostable variant of papain at 2.6 Angstroem resolution | | Descriptor: | CHLORIDE ION, Papain | | Authors: | Roy, S, Choudhury, D, Dattagupta, J.K, Biswas, S. | | Deposit date: | 2011-09-02 | | Release date: | 2012-09-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | The structure of a thermostable mutant of pro-papain reveals its activation mechanism
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3RQC
 
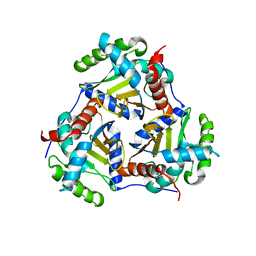 | | Crystal structure of the catalytic core of the 2-oxoacid dehydrogenase multienzyme complex from Thermoplasma acidophilum | | Descriptor: | Probable lipoamide acyltransferase | | Authors: | Marrott, N.L, Crennell, S.J, Hough, D.W, Danson, M.J, van den Elsen, J.M.H. | | Deposit date: | 2011-04-28 | | Release date: | 2012-01-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (4.01 Å) | | Cite: | The catalytic core of an archaeal 2-oxoacid dehydrogenase multienzyme complex is a 42-mer protein assembly.
Febs J., 279, 2012
|
|
3SGC
 
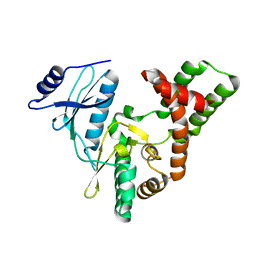 | |
2C7C
 
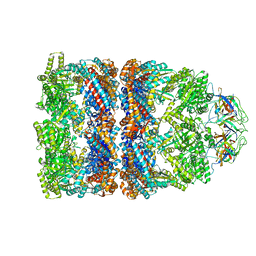 | | FITTED COORDINATES FOR GROEL-ATP7-GROES CRYO-EM COMPLEX (EMD-1180) | | Descriptor: | 10 KDA CHAPERONIN MOLECULE: GROES, PROTEIN CPN10, GROES PROTEIN, ... | | Authors: | Ranson, N.A, Clare, D.K, Farr, G.W, Houldershaw, D, Horwich, A.L, Saibil, H.R. | | Deposit date: | 2005-11-22 | | Release date: | 2006-01-25 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (7.7 Å) | | Cite: | Allosteric Signalling of ATP Hydrolysis in Groel-Groes Complexes.
Nat.Struct.Mol.Biol., 13, 2006
|
|
1W3N
 
 | | Sulfolobus solfataricus 2-keto-3-deoxygluconate (KDG) aldolase complex with D-KDG | | Descriptor: | 2-KETO-3-DEOXY GLUCONATE ALDOLASE, 3-DEOXY-D-ARABINO-HEXONIC ACID, GLYCEROL | | Authors: | Theodossis, A, Walden, H, Westwick, E.J, Connaris, H, Lamble, H.J, Hough, D.W, Danson, M.J, Taylor, G.L. | | Deposit date: | 2004-07-17 | | Release date: | 2004-09-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The structural basis for substrate promiscuity in 2-keto-3-deoxygluconate aldolase from the Entner-Doudoroff pathway in Sulfolobus solfataricus.
J. Biol. Chem., 279, 2004
|
|
2YDA
 
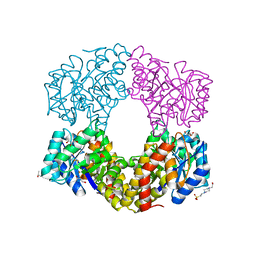 | | Sulfolobus sulfataricus 2-keto-3-deoxygluconate aldolase Y103F,Y130F, A198F variant | | Descriptor: | 2-KETO-3-DEOXY GLUCONATE ALDOLASE, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, GLYCEROL, ... | | Authors: | Crennell, S.J, Royer, S.F, Angelopoulou, M, Hough, D.W, Danson, M.J, Bull, S.D. | | Deposit date: | 2011-03-18 | | Release date: | 2012-03-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Sulfolobus Sulfataricus 2-Keto-3-Deoxygluconate Aldolase
To be Published
|
|
1W9V
 
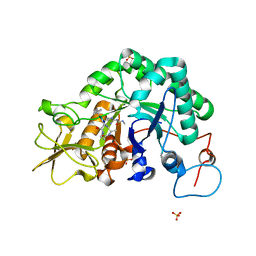 | | Specificity and affinity of natural product cyclopentapeptide argifin against Aspergillus fumigatus | | Descriptor: | ARGIFIN, CHITINASE, SULFATE ION | | Authors: | Rao, F.V, Houston, D.R, Boot, R.G, Aerts, J.M.F.G, Hodkinson, M, Adams, D.J, Shiomi, K, Omura, S, van Aalten, D.M.F. | | Deposit date: | 2004-10-19 | | Release date: | 2005-01-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Specificity and Affinity of Natural Product Cyclopentapeptide Inhibitors Against Aspergillus Fumigatus, Human and Bacterial Chitinases
Chem.Biol., 12, 2005
|
|
1W9U
 
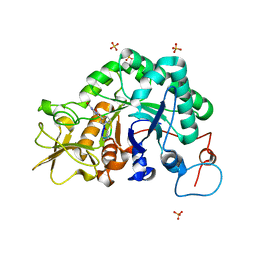 | | Specificity and affnity of natural product cyclopentapeptide inhibitor Argadin against Aspergillus fumigatus chitinase | | Descriptor: | ARGADIN, CHITINASE, SULFATE ION | | Authors: | Rao, F.V, Houston, D.R, Boot, R.G, Aerts, J.M.F.G, Hodkinson, M, Adams, D.J, Shiomi, K, Omura, S, van Aalten, D.M.F. | | Deposit date: | 2004-10-19 | | Release date: | 2004-11-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Specificity and Affinity of Natural Product Cyclopentapeptide Inhibitors Against Aspergillus Fumigatus, Human and Bacterial Chitinases
Chem.Biol., 12, 2005
|
|
1WAW
 
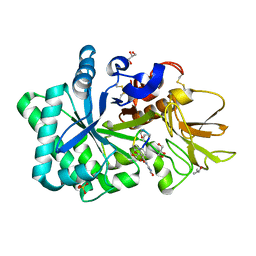 | | Specificity and affinity of natural product cyclopentapeptide inhibitor Argadin against human chitinase | | Descriptor: | ARGADIN, CHITOTRIOSIDASE 1, GLYCEROL, ... | | Authors: | Rao, F.V, Houston, D.R, Boot, R.G, Aerts, J.M.F.G, Hodkinson, M, Adams, D.J, Shiomi, K, Omura, S, van Aalten, D.M.F. | | Deposit date: | 2004-10-28 | | Release date: | 2005-01-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Specificity and Affinity of Natural Product Cyclopentapeptide Inhibitors Against Aspergillus Fumigatus, Human and Bacterial Chitinases
Chem.Biol., 12, 2005
|
|
