5CWV
 
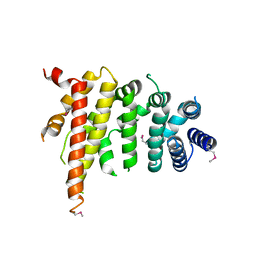 | | Crystal structure of Chaetomium thermophilum Nup192 TAIL domain | | Descriptor: | Nucleoporin NUP192 | | Authors: | Stuwe, T, Bley, C.J, Thierbach, K, Petrovic, S, Schilbach, S, Mayo, D.J, Perriches, T, Rundlet, E.J, Jeon, Y.E, Collins, L.N, Lin, D.H, Paduch, M, Koide, A, Lu, V, Fischer, J, Hurt, E, Koide, S, Kossiakoff, A.A, Hoelz, A. | | Deposit date: | 2015-07-28 | | Release date: | 2015-09-16 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (3.155 Å) | | Cite: | Architecture of the fungal nuclear pore inner ring complex.
Science, 350, 2015
|
|
5D8K
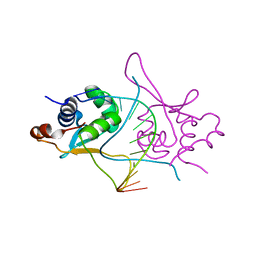 
 | |
5DET
 
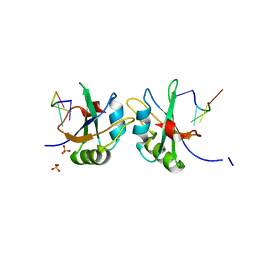 | | X-ray structure of human RBPMS in complex with the RNA | | Descriptor: | RNA (5'-R(*UP*CP*AP*C)-3'), RNA (5'-R(P*UP*CP*AP*CP*U)-3'), RNA-binding protein with multiple splicing, ... | | Authors: | Teplova, M, Farazi, T.A, Tuschl, T, Patel, D.J. | | Deposit date: | 2015-08-25 | | Release date: | 2015-09-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural basis underlying CAC RNA recognition by the RRM domain of dimeric RNA-binding protein RBPMS.
Q. Rev. Biophys., 49, 2016
|
|
5DA5
 
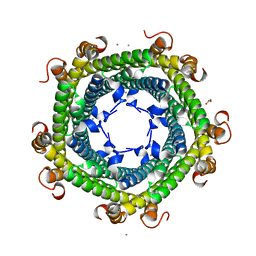 | | Crystal structure of Rhodospirillum rubrum Rru_A0973 | | Descriptor: | CALCIUM ION, FE (III) ION, GLYCOLIC ACID, ... | | Authors: | He, D, Vanden Hehier, S, Georgiev, A, Altenbach, K, Tarrant, E, Mackay, C.L, Waldron, K.J, Clarke, D.J, Marles-Wright, J. | | Deposit date: | 2015-08-19 | | Release date: | 2016-08-10 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.064 Å) | | Cite: | Structural characterization of encapsulated ferritin provides insight into iron storage in bacterial nanocompartments.
Elife, 5, 2016
|
|
5CWU
 
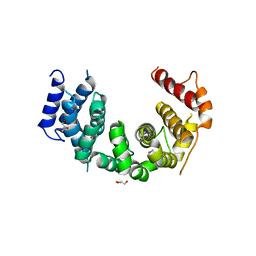 | | Crystal structure of Chaetomium thermophilum Nup188 TAIL domain | | Descriptor: | GLYCEROL, Nucleoporin NUP188 | | Authors: | Stuwe, T, Bley, C.J, Thierbach, K, Petrovic, S, Schilbach, S, Mayo, D.J, Perriches, T, Rundlet, E.J, Jeon, Y.E, Collins, L.N, Lin, D.H, Paduch, M, Koide, A, Lu, V, Fischer, J, Hurt, E, Koide, S, Kossiakoff, A.A, Hoelz, A. | | Deposit date: | 2015-07-28 | | Release date: | 2015-09-16 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Architecture of the fungal nuclear pore inner ring complex.
Science, 350, 2015
|
|
5D3M
 
 | | Folate ECF transporter: AMPPNP bound state | | Descriptor: | Energy-coupling factor transporter ATP-binding protein EcfA1, Energy-coupling factor transporter ATP-binding protein EcfA2, Energy-coupling factor transporter transmembrane protein EcfT, ... | | Authors: | Guskov, A, Swier, L.J.Y.M, Slotboom, D.J. | | Deposit date: | 2015-08-06 | | Release date: | 2016-04-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.303 Å) | | Cite: | Structural insight in the toppling mechanism of an energy-coupling factor transporter.
Nat Commun, 7, 2016
|
|
5D8L
 
 | |
5D69
 
 | | Human calpain PEF(S) with (2Z,2Z')-2,2'-disulfanediylbis(3-(6-iodoindol-3-yl)acrylic acid) bound | | Descriptor: | (2E,2'Z)-2,2'-disulfanediylbis[3-(4-iodophenyl)prop-2-enoic acid], CALCIUM ION, Calpain small subunit 1, ... | | Authors: | Adams, S.E, Robinson, E.J, Rizkallah, P.J, Miller, D.J, Hallett, M.B, Allemann, R.K. | | Deposit date: | 2015-08-11 | | Release date: | 2015-09-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Conformationally restricted calpain inhibitors.
Chem Sci, 6, 2015
|
|
5DAH
 
 | |
5DAG
 
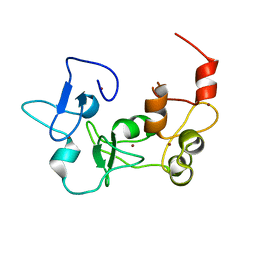 | |
2I1D
 | | DPC micelle-bound NMR structures of Tritrp1 | | Descriptor: | 13-mer from Prophenin-1 containing WWW | | Authors: | Schibli, D.J, Nguyen, L.T. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure-function analysis of tritrpticin analogs: potential relationships between antimicrobial activities, model membrane interactions, and their micelle-bound NMR structures
Biophys.J., 91, 2006
|
|
2I1F
 | | DPC micelle-bound NMR structures of Tritrp3 | | Descriptor: | 13-mer analogue of Prophenin-1 containing WWW | | Authors: | Schibli, D.J, Nguyen, L.T. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Structure-function analysis of tritrpticin analogs: potential relationships between antimicrobial activities, model membrane interactions, and their micelle-bound NMR structures
Biophys.J., 91, 2006
|
|
1SWX
 
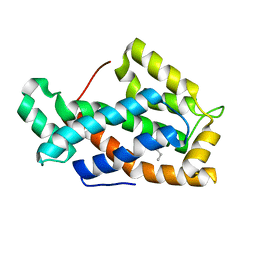 | | Crystal structure of a human glycolipid transfer protein in apo-form | | Descriptor: | Glycolipid transfer protein, HEXANE | | Authors: | Malinina, L, Malakhova, M.L, Teplov, A, Brown, R.E, Patel, D.J. | | Deposit date: | 2004-03-30 | | Release date: | 2004-08-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis for glycosphingolipid transfer specificity.
Nature, 430, 2004
|
|
1SX6
 
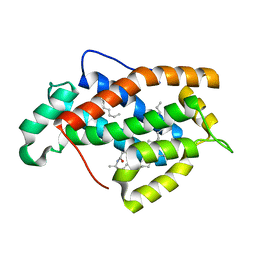 | | Crystal structure of human Glycolipid Transfer protein in lactosylceramide-bound form | | Descriptor: | Glycolipid transfer protein, N-OCTANE, OLEIC ACID, ... | | Authors: | Malinina, L, Malakhova, M.L, Teplov, A, Brown, R.E, Patel, D.J. | | Deposit date: | 2004-03-30 | | Release date: | 2004-08-31 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural basis for glycosphingolipid transfer specificity.
Nature, 430, 2004
|
|
2I13
 
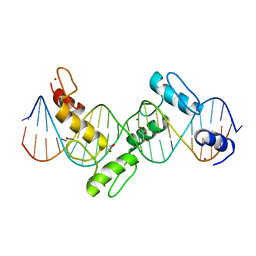 | | Aart, a six finger zinc finger designed to recognize ANN triplets | | Descriptor: | 5'-D(*CP*AP*GP*AP*TP*GP*TP*AP*GP*GP*GP*AP*AP*AP*AP*GP*CP*CP*CP*GP*GP*G)-3', 5'-D(*GP*CP*CP*CP*GP*GP*GP*CP*TP*TP*TP*TP*CP*CP*CP*TP*AP*CP*AP*TP*CP*T)-3', Aart, ... | | Authors: | Horton, N.C, Segal, D.J, Bhakta, M, Crotty, J.W, Barbas III, C.F. | | Deposit date: | 2006-08-12 | | Release date: | 2006-10-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structure of Aart, a Designed Six-finger Zinc Finger Peptide, Bound to DNA.
J.Mol.Biol., 363, 2006
|
|
2I1G
 
 | | DPC micelle-bound NMR structures of Tritrp5 | | Descriptor: | 13-mer analogue of Prophenin-1 containing WWW | | Authors: | Schibli, D.J, Nguyen, L.T. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structure-function analysis of tritrpticin analogs: potential relationships between antimicrobial activities, model membrane interactions, and their micelle-bound NMR structures
Biophys.J., 91, 2006
|
|
2I1I
 
 | | DPC micelle-bound NMR structures of Tritrp8 | | Descriptor: | 13-mer analogue of Prophenin-1 containing WWW | | Authors: | Schibli, D.J, Nguyen, L.T. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structure-function analysis of tritrpticin analogs: potential relationships between antimicrobial activities, model membrane interactions, and their micelle-bound NMR structures
Biophys.J., 91, 2006
|
|
1TJX
 | | Crystallographic Identification of Ca2+ Coordination Sites in Synaptotagmin I C2B Domain | | Descriptor: | ACETATE ION, CALCIUM ION, GLYCEROL, ... | | Authors: | Cheng, Y, Sequeira, S.M, Malinina, L, Tereshko, V, Sollner, T.H, Patel, D.J. | | Deposit date: | 2004-06-07 | | Release date: | 2004-11-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Crystallographic identification of Ca2+ and Sr2+ coordination sites in synaptotagmin I C2B domain.
Protein Sci., 13, 2004
|
|
2I03
 
 | | Crystal structure of human dipeptidyl peptidase 4 (DPP IV) with potent alkynyl cyanopyrrolidine (ABT-279) | | Descriptor: | 2-[4-({2-[(2S,5R)-2-(AMINOMETHYL)-5-ETHYNYLPYRROLIDIN-1-YL]-2-OXOETHYL}AMINO)-4-METHYLPIPERIDIN-1-YL]ISONICOTINIC ACID, Dipeptidyl peptidase 4 | | Authors: | Longenecker, K.L, Madar, D.J. | | Deposit date: | 2006-08-09 | | Release date: | 2006-12-12 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Discovery of 2-[4-{{2-(2S,5R)-2-cyano-5-ethynyl-1-pyrrolidinyl]-2-oxoethyl]amino]- 4-methyl-1-piperidinyl]-4-pyridinecarboxylic acid (ABT-279): a very potent, selective, effective, and well-tolerated inhibitor of dipeptidyl peptidase-IV, useful for the treatment of diabetes.
J.Med.Chem., 49, 2006
|
|
1TI8
 
 | | H7 Haemagglutinin | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, alpha-D-mannopyranose, ... | | Authors: | Russell, R.J, Gamblin, S.J, Haire, L.F, Stevens, D.J, Xaio, B, Ha, Y, Skehel, J.J. | | Deposit date: | 2004-06-02 | | Release date: | 2005-06-21 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | H1 and H7 influenza haemagglutinin structures extend a structural classification of haemagglutinin subtypes.
Virology, 325, 2004
|
|
2I1E
 | | DPC micelle-bound NMR structures of Tritrp2 | | Descriptor: | 13-mer analogue of Prophenin-1 containing WWW | | Authors: | Schibli, D.J, Nguyen, L.T. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structure-function analysis of tritrpticin analogs: potential relationships between antimicrobial activities, model membrane interactions, and their micelle-bound NMR structures
Biophys.J., 91, 2006
|
|
1T6L
 
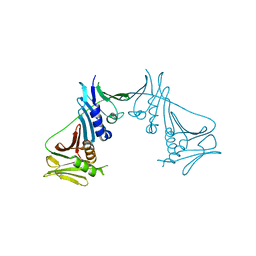 | | Crystal Structure of the Human Cytomegalovirus DNA Polymerase Subunit, UL44 | | Descriptor: | DNA polymerase processivity factor | | Authors: | Appleton, B.A, Loregian, A, Filman, D.J, Coen, D.M, Hogle, J.M. | | Deposit date: | 2004-05-06 | | Release date: | 2004-08-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Cytomegalovirus DNA Polymerase Subunit UL44 Forms a C Clamp-Shaped Dimer.
Mol.Cell, 15, 2004
|
|
1T7K
 
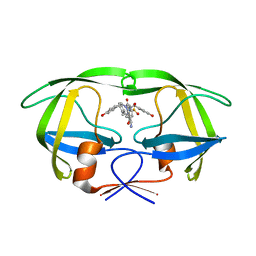 | | Crystal Structure of HIV Protease complexed with Arylsulfonamide azacyclic urea | | Descriptor: | 3-({5-BENZYL-6-HYDROXY-2,4-BIS-(4-HYDROXY-BENZYL)-3-OXO-[1,2,4]-TRIAZEPANE-1-SULFONYL)-BENZONITRILE, Pol polyprotein [Contains: Protease (Retropepsin)] | | Authors: | Huang, P.P, Randolph, J.T, Klein, L.L, Vasavanonda, S, Dekhtyar, T, Stoll, V.S, Kempf, D.J. | | Deposit date: | 2004-05-10 | | Release date: | 2004-10-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Synthesis and Antiviral Activity of P1' Arylsulfonamide Azacyclic Urea HIV Protease Inhibitors
Bioorg.Med.Chem.Lett., 14, 2004
|
|
1TFZ
 
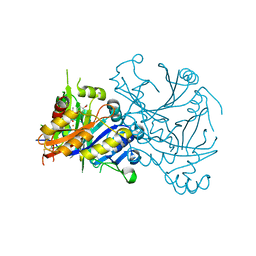 | | Structural basis for herbicidal inhibitor selectivity revealed by comparison of crystal structures of plant and mammalian 4-hydroxyphenylpyruvate dioxygenases | | Descriptor: | (1-TERT-BUTYL-5-HYDROXY-1H-PYRAZOL-4-YL)[6-(METHYLSULFONYL)-4'-METHOXY-2-METHYL-1,1'-BIPHENYL-3-YL]METHANONE, 4-hydroxyphenylpyruvate dioxygenase, FE (III) ION | | Authors: | Yang, C, Pflugrath, J.W, Camper, D.L, Foster, M.L, Pernich, D.J, Walsh, T.A. | | Deposit date: | 2004-05-27 | | Release date: | 2004-08-17 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for herbicidal inhibitor selectivity revealed by comparison of crystal structures of plant and Mammalian 4-hydroxyphenylpyruvate dioxygenases
Biochemistry, 43, 2004
|
|
2IHE
 
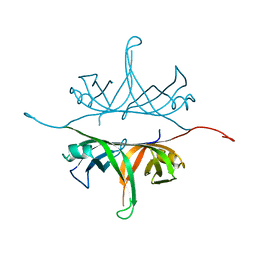 | | Crystal structure of wild-type single-stranded DNA binding protein from Thermus aquaticus | | Descriptor: | Single-stranded DNA-binding protein | | Authors: | Fedorov, R, Witte, G, Urbanke, C, Manstein, D.J, Curth, U. | | Deposit date: | 2006-09-26 | | Release date: | 2007-01-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | 3D structure of Thermus aquaticus single-stranded DNA-binding protein gives insight into the functioning of SSB proteins.
Nucleic Acids Res., 34, 2006
|
|
