1R4G
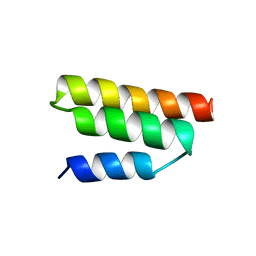 
 | | Solution structure of the Sendai virus protein X C-subdomain | | Descriptor: | RNA polymerase alpha subunit | | Authors: | Blanchard, L, Tarbouriech, N, Blackledge, M, Timmins, P, Burmeister, W.P, Ruigrok, R.W, Marion, D. | | Deposit date: | 2003-10-06 | | Release date: | 2004-03-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and dynamics of the nucleocapsid-binding domain of the Sendai virus phosphoprotein in solution
Virology, 319, 2004
|
|
5JKS
 
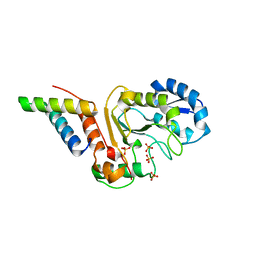 | | vaccinia virus D4 R167A mutant /A20(1-50) | | Descriptor: | DNA polymerase processivity factor component A20, SULFATE ION, Uracil-DNA glycosylase | | Authors: | Contesto-Richefeu, C, Tarbouriech, N, Brazzolotto, X, Burmeister, W.P, Peyrefitte, C.N, Iseni, F. | | Deposit date: | 2016-04-26 | | Release date: | 2016-09-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structural analysis of point mutations at the Vaccinia virus A20/D4 interface.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
5JKR
 
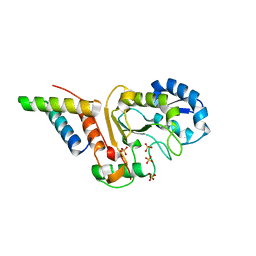 | | vaccinia virus D4/A20(1-50)w43a mutant | | Descriptor: | DNA polymerase processivity factor component A20, SULFATE ION, Uracil-DNA glycosylase | | Authors: | Contesto-Richefeu, C, Tarbouriech, N, Brazzolotto, X, Burmeister, W.P, Peyrefitte, C.N, Iseni, F. | | Deposit date: | 2016-04-26 | | Release date: | 2016-09-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural analysis of point mutations at the Vaccinia virus A20/D4 interface.
Acta Crystallogr.,Sect.F, 72, 2016
|
|
2BSY
 
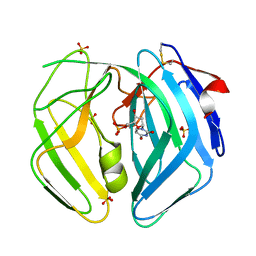 | | Epstein Barr Virus dUTPase | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, DEOXYURIDINE 5'-TRIPHOSPHATE NUCLEOTIDOHYDROLASE, SULFATE ION | | Authors: | Tarbouriech, N, Buisson, M, Seigneurin, J.-M, Cusack, S, Burmeister, W.P. | | Deposit date: | 2005-05-24 | | Release date: | 2005-09-15 | | Last modified: | 2023-03-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Monomeric Dutpase from Epstein-Barr Virus Mimics Trimeric Dutpases
Structure, 13, 2005
|
|
2BT1
 
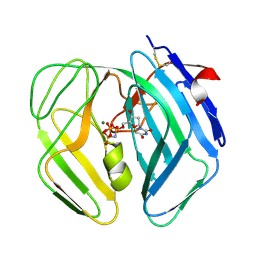 | | Epstein Barr Virus dUTPase in complex with a,b-imino dUTP | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, DEOXYURIDINE 5'-TRIPHOSPHATE NUCLEOTIDOHYDROLASE, MAGNESIUM ION | | Authors: | Tarbouriech, N, Buisson, M, Seigneurin, J.-M, Cusack, S, Burmeister, W.P. | | Deposit date: | 2005-05-24 | | Release date: | 2005-09-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Monomeric Dutpase from Epstein-Barr Virus Mimics Trimeric Dutpases
Structure, 13, 2005
|
|
2WXD
 
 | | A MICROMOLAR O-SULFATED THIOHYDROXIMATE INHIBITOR BOUND TO PLANT MYROSINASE | | Descriptor: | 2-(DIMETHYLAMINO)ETHYL (1Z)-2-PHENYL-N-(SULFOOXY)ETHANIMIDOTHIOATE, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Besle, A, Burmeister, W.P. | | Deposit date: | 2009-11-09 | | Release date: | 2010-02-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A Micromolar O-Sulfated Thiohydroximate Inhibitor Bound to Plant Myrosinase
Acta Crystallogr.,Sect.F, 66, 2010
|
|
4OD8
 
 | | Crystal structure of the vaccinia virus DNA polymerase holoenzyme subunit D4 in complex with the A20 N-terminus | | Descriptor: | DNA polymerase processivity factor component A20, GLYCEROL, SULFATE ION, ... | | Authors: | Contesto-Richefeu, C, Tarbouriech, N, Brazzolotto, X, Burmeister, W.P, Iseni, F. | | Deposit date: | 2014-01-10 | | Release date: | 2014-03-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of the vaccinia virus DNA polymerase holoenzyme subunit d4 in complex with the a20 N-terminal domain.
Plos Pathog., 10, 2014
|
|
4ODA
 
 | | Crystal structure of the vaccinia virus DNA polymerase holoenzyme subunit D4 in complex with the A20 N-terminus | | Descriptor: | DNA polymerase processivity factor component A20, GLYCEROL, SULFATE ION, ... | | Authors: | Contesto-Richefeu, C, Tarbouriech, N, Brazzolotto, X, Burmeister, W.P, Iseni, F. | | Deposit date: | 2014-01-10 | | Release date: | 2014-03-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the vaccinia virus DNA polymerase holoenzyme subunit d4 in complex with the a20 N-terminal domain.
Plos Pathog., 10, 2014
|
|
1H7Z
 
 | | Adenovirus Ad3 fibre head | | Descriptor: | ADENOVIRUS FIBRE PROTEIN, SULFATE ION | | Authors: | Durmort, C, Stehlin, C, Schoehn, G, Mitraki, A, Drouet, E, Cusack, S, Burmeister, W.P. | | Deposit date: | 2001-01-21 | | Release date: | 2001-07-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of the Fiber Head of Ad3, a Non-Car-Binding Serotype of Adenovirus
Virology, 285, 2001
|
|
1O6E
 
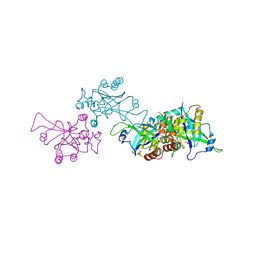 | | Epstein-Barr virus protease | | Descriptor: | CAPSID PROTEIN P40, PHOSPHORYLISOPROPANE | | Authors: | Buisson, M, Hernandez, J, Lascoux, D, Schoehn, G, Forest, E, Arlaud, G, Seigneurin, J, Ruigrok, R.W.H, Burmeister, W.P. | | Deposit date: | 2002-09-13 | | Release date: | 2002-11-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of the Epstein-Barr Virus Protease Shows Rearrangement of the Processed C Terminus
J.Mol.Biol., 324, 2002
|
|
1WCG
 
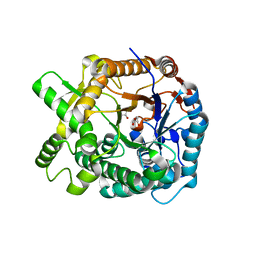 | | Aphid myrosinase | | Descriptor: | GLYCEROL, THIOGLUCOSIDASE | | Authors: | Husebye, H, Arzt, S, Burmeister, W.P, Haertel, F.V, Brandt, A, Rossiter, J.T, Bones, A.M. | | Deposit date: | 2004-11-15 | | Release date: | 2005-11-23 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal Structure at 1.1A Resolution of an Insect Myrosinase from Brevicoryne Brassicae Shows its Close Relationship to Beta-Glucosidases.
Insect Biochem.Mol.Biol., 35, 2005
|
|
1CZ7
 
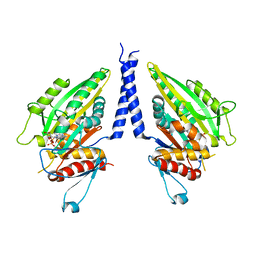 | | THE CRYSTAL STRUCTURE OF A MINUS-END DIRECTED MICROTUBULE MOTOR PROTEIN NCD REVEALS VARIABLE DIMER CONFORMATIONS | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, MICROTUBULE MOTOR PROTEIN NCD | | Authors: | Kozielski, F.K, De Bonis, S, Burmeister, W, Cohen-Addad, C, Wade, R. | | Deposit date: | 1999-09-01 | | Release date: | 1999-11-05 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The crystal structure of the minus-end-directed microtubule motor protein ncd reveals variable dimer conformations.
Structure Fold.Des., 7, 1999
|
|
1KU6
 
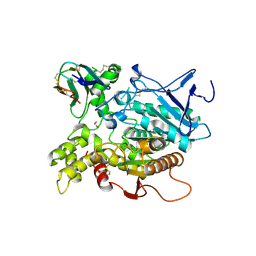 | | Fasciculin 2-Mouse Acetylcholinesterase Complex | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETYLCHOLINESTERASE, ... | | Authors: | Bourne, Y, Burmeister, W, Taylor, P, Marchot, P. | | Deposit date: | 2002-01-21 | | Release date: | 2003-12-23 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into ligand interactions at the acetylcholinesterase peripheral anionic site.
EMBO J., 22, 2003
|
|
