1D28
 
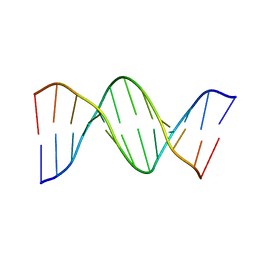 | |
1MI7
 
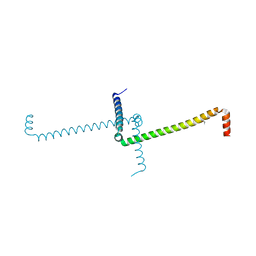 | | Crystal Structure of Domain Swapped trp Aporepressor in 30%(v/v) Isopropanol | | Descriptor: | ISOPROPYL ALCOHOL, Trp operon repressor | | Authors: | Lawson, C.L, Benoff, B, Berger, T, Berman, H.M, Carey, J. | | Deposit date: | 2002-08-22 | | Release date: | 2003-09-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | E. coli trp repressor forms a domain-swapped array in aqueous alcohol.
Structure, 12, 2004
|
|
218D
 
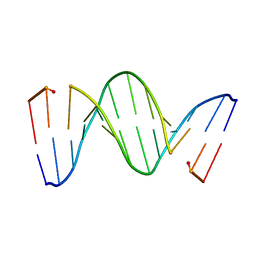 | | THE STRUCTURE OF A NEW CRYSTAL FORM OF A DNA DODECAMER CONTAINING T.(O6ME)G BASE PAIRS | | Descriptor: | DNA (5'-D(*CP*GP*TP*GP*AP*AP*TP*TP*CP*(6OG)P*CP*G)-3') | | Authors: | Vojtechovsky, J, Eaton, M.D, Gaffney, B, Jones, R, Berman, H.M. | | Deposit date: | 1995-07-18 | | Release date: | 1995-10-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of a new crystal form of a DNA dodecamer containing T.(O6Me)G base pairs.
Biochemistry, 34, 1995
|
|
210D
 
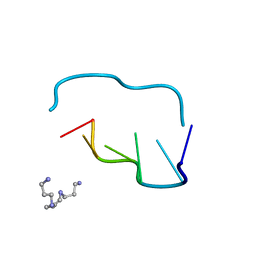 | | CRYSTAL AND MOLECULAR STRUCTURE OF A NEW Z-DNA CRYSTAL FORM: D[CGT(2-NH2-A)CG] AND ITS PLATINATED DERIVATIVE | | Descriptor: | DNA (5'-D(*CP*GP*TP*(1AP)P*CP*G)-3'), SPERMINE | | Authors: | Parkinson, G.N, Arvantis, G.M, Lessinger, L, Ginell, S.L, Jones, R, Gaffney, B, Berman, H.M. | | Deposit date: | 1995-06-13 | | Release date: | 1996-03-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal and molecular structure of a new Z-DNA crystal form: d[CGT(2-NH2-A)CG] and its platinated derivative.
Biochemistry, 34, 1995
|
|
237D
 
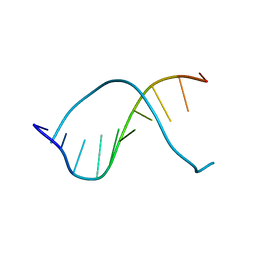 | | CRYSTAL STRUCTURE OF A DNA DECAMER SHOWING A NOVEL PSEUDO FOUR-WAY HELIX-HELIX JUNCTION | | Descriptor: | DNA (5'-D(*CP*GP*CP*AP*AP*TP*TP*GP*CP*G)-3') | | Authors: | Spink, N, Nunn, C.M, Vojtechovsky, J, Berman, H.M, Neidle, S. | | Deposit date: | 1995-09-28 | | Release date: | 1996-03-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a DNA decamer showing a novel pseudo four-way helix-helix junction.
Proc.Natl.Acad.Sci.USA, 92, 1995
|
|
211D
 
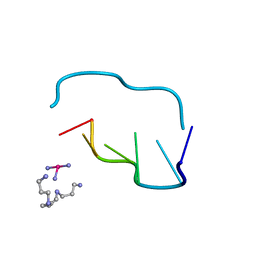 | | THE CRYSTAL AND MOLECULAR STRUCTURE OF A NEW Z-DNA CRYSTAL FORM: D[CGT(2-NH2-A) CG] AND ITS PLATINATED DERIVATIVE | | Descriptor: | DNA (5'-D(*CP*GP*TP*(1AP)P*CP*(PT(NH3)3)G)-3'), PLATINUM TRIAMINE ION, SPERMINE | | Authors: | Parkinson, G.N, Arvantis, G.M, Lessinger, L, Ginell, S.L, Jones, R, Gaffney, B, Berman, H.M. | | Deposit date: | 1995-06-13 | | Release date: | 1996-03-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal and molecular structure of a new Z-DNA crystal form: d[CGT(2-NH2-A)CG] and its platinated derivative.
Biochemistry, 34, 1995
|
|
1HIB
 
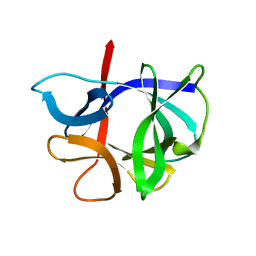 | | THE STRUCTURE OF AN INTERLEUKIN-1 BETA MUTANT WITH REDUCED BIOACTIVITY SHOWS MULTIPLE SUBTLE CHANGES IN CONFORMATION THAT AFFECT PROTEIN-PROTEIN RECOGNITION | | Descriptor: | INTERLEUKIN-1 BETA | | Authors: | Camacho, N.P, Smith, D.R, Goldman, A, Schneider, B, Green, D, Young, P.R, Berman, H.M. | | Deposit date: | 1993-03-29 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of an interleukin-1 beta mutant with reduced bioactivity shows multiple subtle changes in conformation that affect protein-protein recognition.
Biochemistry, 32, 1993
|
|
1HQ7
 
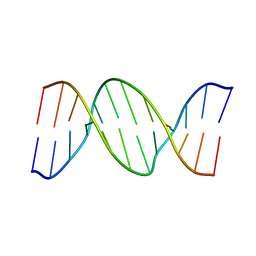 | |
3SSW
 
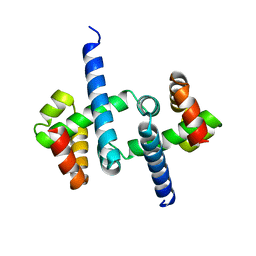 | | E. coli trp aporepressor | | Descriptor: | Trp operon repressor | | Authors: | Benoff, B, Carey, J, Berman, H.M, Lawson, C.L. | | Deposit date: | 2011-07-08 | | Release date: | 2011-07-20 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6693 Å) | | Cite: | Environment-dependent long-range structural distortion in a temperature-sensitive point mutant.
Protein Sci., 21, 2012
|
|
3SSX
 
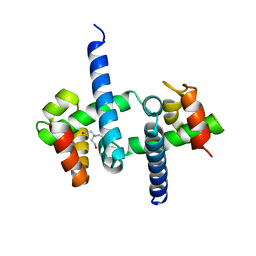 | | E. coli trp aporeporessor L75F mutant | | Descriptor: | TRIS(HYDROXYETHYL)AMINOMETHANE, Trp operon repressor | | Authors: | Benoff, B, Carey, J, Berman, H.M, Lawson, C.L. | | Deposit date: | 2011-07-08 | | Release date: | 2011-07-20 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.5801 Å) | | Cite: | Environment-dependent long-range structural distortion in a temperature-sensitive point mutant.
Protein Sci., 21, 2012
|
|
7PSC
 
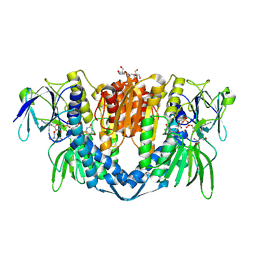 | | Crystal structure of the disease-causing I358T mutant of the human dihydrolipoamide dehydrogenase | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Dihydrolipoyl dehydrogenase, mitochondrial, ... | | Authors: | Nemes-Nikodem, E, Szabo, E, Zambo, Z, Vass, K.R, Taberman, H, Torocsik, B, Weiss, M.S, Adam-Vizi, V, Ambrus, A. | | Deposit date: | 2021-09-22 | | Release date: | 2023-04-05 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.436 Å) | | Cite: | Structural and Biochemical Investigation of Selected Pathogenic Mutants of the Human Dihydrolipoamide Dehydrogenase.
Int J Mol Sci, 24, 2023
|
|
