3QZU
 
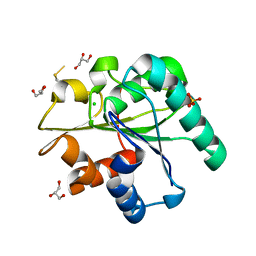 | | Crystal structure of Bacillus subtilis Lipase A 7-fold mutant; the outcome of directed evolution towards thermostability | | Descriptor: | CHLORIDE ION, GLYCEROL, Lipase estA, ... | | Authors: | Pijning, T, Augustyniak, W, Reetz, M.T, Dijkstra, B.W. | | Deposit date: | 2011-03-07 | | Release date: | 2012-02-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Biophysical characterization of mutants of Bacillus subtilis lipase evolved for thermostability: Factors contributing to increased activity retention.
Protein Sci., 21, 2012
|
|
2KL4
 
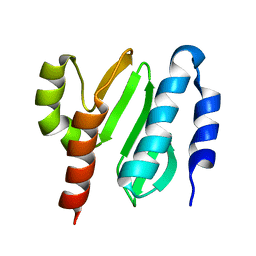 | | NMR structure of the protein NB7804A | | Descriptor: | BH2032 protein | | Authors: | Mohanty, B, Geralt, M, Augustyniak, W, Serrano, P, Horst, R, Wilson, I.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2009-06-30 | | Release date: | 2009-07-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the protein NB7804A from Bacillus Halodurans
To be Published
|
|
2LYY
 
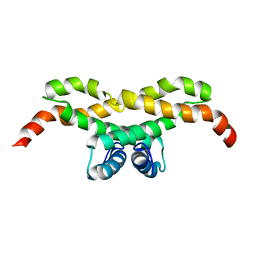 | | NMR structure of the protein NB7890A from Shewanella sp | | Descriptor: | Uncharacterized protein | | Authors: | Serrano, P, Geralt, M, Pedrini, B, Wuthrich, K, Horst, R, Augustyniak, W, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2012-09-21 | | Release date: | 2012-10-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the protein NB7890A from Shewanella sp
To be Published
|
|
