7SWJ
 
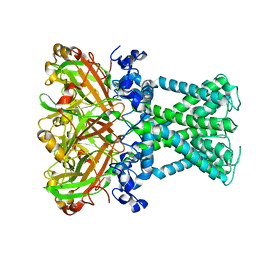 | | KirBac1.1 mutant - I131C | | Descriptor: | Inward rectifier potassium channel | | Authors: | Amani, R, Wylie, B.J. | | Deposit date: | 2021-11-19 | | Release date: | 2022-02-02 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Water Accessibility Refinement of the Extended Structure of KirBac1.1 in the Closed State.
Front Mol Biosci, 8, 2021
|
|
5Z99
 
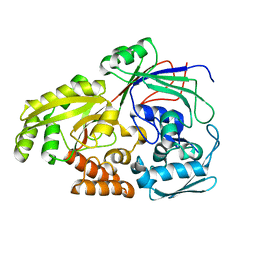 | |
8F0V
 
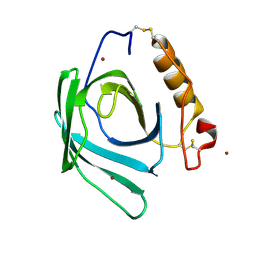 | | Lipocalin-like Milk protein-2 - E38A mutant | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Milk protein, ZINC ION | | Authors: | Subramanian, R, KanagaVijayan, D. | | Deposit date: | 2022-11-04 | | Release date: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.951 Å) | | Cite: | Variability in phenylalanine side chain conformations facilitates broad substrate tolerance of fatty acid binding in cockroach milk proteins.
Plos One, 18, 2023
|
|
8F0Y
 
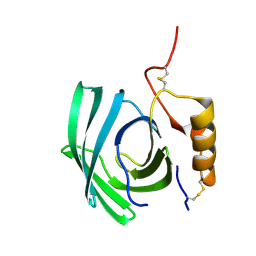 | | Lipocalin-like Milk protein-1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Milk protein | | Authors: | Subramanian, R, KanagaVijayan, D, Shantakumar, R.P.S. | | Deposit date: | 2022-11-04 | | Release date: | 2023-08-23 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Variability in phenylalanine side chain conformations facilitates broad substrate tolerance of fatty acid binding in cockroach milk proteins.
Plos One, 18, 2023
|
|
7KC2
 
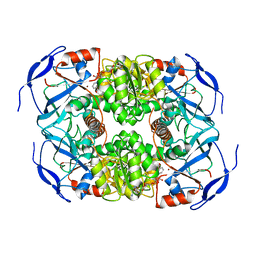 | | Symmetry in Yeast Alcohol Dehydrogenase 1 -Closed Form with NADH | | Descriptor: | Alcohol dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ZINC ION | | Authors: | Subramanian, R, Chang, L, Li, Z, Plapp, B.V. | | Deposit date: | 2020-10-04 | | Release date: | 2021-03-31 | | Last modified: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (2.67 Å) | | Cite: | Cryo-Electron Microscopy Structures of Yeast Alcohol Dehydrogenase.
Biochemistry, 60, 2021
|
|
7KCB
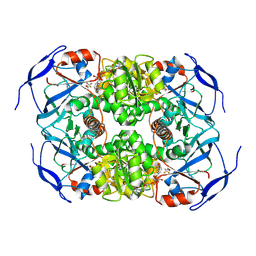 
 | | Symmetry in Yeast Alcohol Dehydrogenase 1 -Closed Form with NAD+ and Trifluoroethanol | | Descriptor: | ADH1 isoform 1, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, TRIFLUOROETHANOL, ... | | Authors: | Subramanian, R, Chang, L, Li, Z, Plapp, B.V, Guntupalli, S.R. | | Deposit date: | 2020-10-05 | | Release date: | 2021-03-31 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2.77 Å) | | Cite: | Cryo-Electron Microscopy Structures of Yeast Alcohol Dehydrogenase.
Biochemistry, 60, 2021
|
|
7KCQ
 
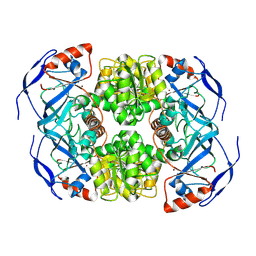 | | Symmetry in Yeast Alcohol Dehydrogenase 1 -Open Form of Apoenzyme | | Descriptor: | Alcohol dehydrogenase, ZINC ION | | Authors: | Subramanian, R, Chang, L, Li, Z, Plapp, B.V, Guntupalli, S.R. | | Deposit date: | 2020-10-07 | | Release date: | 2021-03-31 | | Last modified: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Cryo-Electron Microscopy Structures of Yeast Alcohol Dehydrogenase.
Biochemistry, 60, 2021
|
|
4L6Y
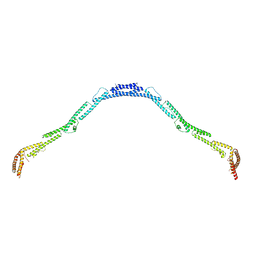 
 | | Structure of the microtubule associated protein PRC1 (Protein Regulator of Cytokinesis 1) | | Descriptor: | Protein regulator of cytokinesis 1 | | Authors: | Subramanian, R, Ti, S, Tan, L, Darst, S.A, Kapoor, T.M. | | Deposit date: | 2013-06-13 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.3015 Å) | | Cite: | Marking and Measuring Single Microtubules by PRC1 and Kinesin-4.
Cell(Cambridge,Mass.), 154, 2013
|
|
4L3I
 
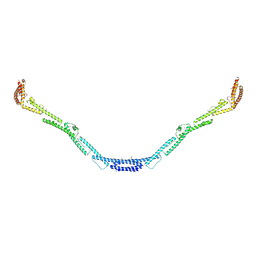 | | Structure of the microtubule associated protein PRC1 (Protein Regulator of Cytokinesis 1) | | Descriptor: | Protein regulator of cytokinesis 1 | | Authors: | Subramanian, R, Ti, S, Tan, L, Darst, S.A, Kapoor, T.M. | | Deposit date: | 2013-06-06 | | Release date: | 2013-07-17 | | Last modified: | 2013-08-07 | | Method: | X-RAY DIFFRACTION (3.6005 Å) | | Cite: | Marking and Measuring Single Microtubules by PRC1 and Kinesin-4.
Cell(Cambridge,Mass.), 154, 2013
|
|
7LQM
 
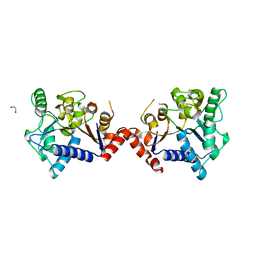 | |
7LQN
 
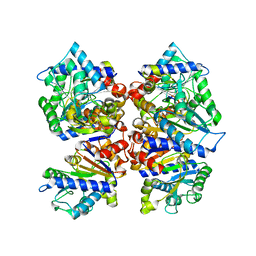 | |
5YYB
 
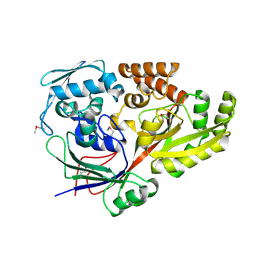 | |
7YX1
 
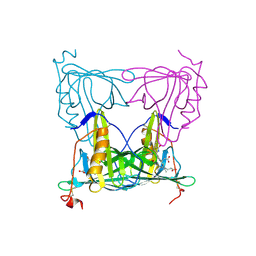 | |
7KJY
 
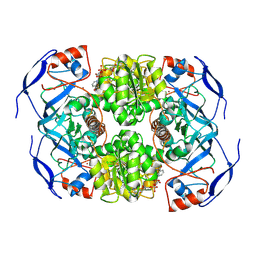 | | Symmetry in Yeast Alcohol Dehydrogenase 1 - Open Form with NADH | | Descriptor: | Alcohol dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ZINC ION | | Authors: | Subramanian, R, Chang, L, Li, Z, Plapp, B.V, Guntupalli, S.R. | | Deposit date: | 2020-10-26 | | Release date: | 2021-03-31 | | Last modified: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Cryo-Electron Microscopy Structures of Yeast Alcohol Dehydrogenase.
Biochemistry, 60, 2021
|
|
6JKU
 
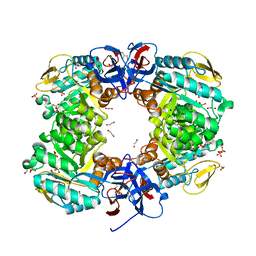 | | Crystal structure of N-acetylglucosamine-6-phosphate deacetylase from Pasteurella Multocida | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, N-acetylglucosamine-6-phosphate deacetylase, ... | | Authors: | Manjunath, L, Bose, S, Subramanian, R. | | Deposit date: | 2019-03-01 | | Release date: | 2020-03-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Quaternary variations in the structural assembly of N-acetylglucosamine-6-phosphate deacetylase from Pasteurella multocida.
Proteins, 2020
|
|
5ZNH
 
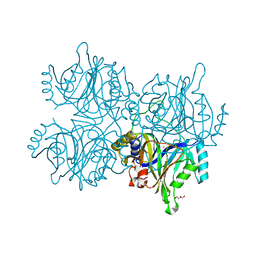 | | Catechol 2,3-dioxygenase with 4-methyl catechol from Diaphorobacter sp DS2 | | Descriptor: | 1,2-ETHANEDIOL, 4-METHYLCATECHOL, CALCIUM ION, ... | | Authors: | Mishra, K, Arya, C.K, Subramaniyan, R, Ramanathan, G. | | Deposit date: | 2018-04-09 | | Release date: | 2019-04-17 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | catechol 2,3-dioxygenase with 4-methyl catechol from Diaphorobacter sp DS2
To Be Published
|
|
5ZSX
 
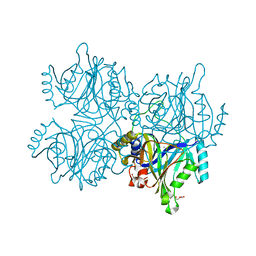 | | Catechol 2,3-dioxygenase with 3-fluorocatechol from Diaphorobacter sp DS2 | | Descriptor: | 1,2-ETHANEDIOL, 3-FLUOROBENZENE-1,2-DIOL, CALCIUM ION, ... | | Authors: | Mishra, K, Arya, C.K, Subramanian, R, Ramanathan, G. | | Deposit date: | 2018-04-30 | | Release date: | 2019-06-12 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Catechol 2,3-dioxygenase with 3-fluorocatechol from Diaphorobacter sp DS2.
To Be Published
|
|
5ZSZ
 
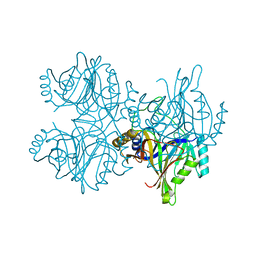 | | Catechol 2,3-dioxygenase (C23O64) from Diaphorobacter sp DS2 | | Descriptor: | CALCIUM ION, Catechol 2,3-dioxygenase, Extradiol ring cleavage protein, ... | | Authors: | Mishra, K, Arya, C.K, Subramanian, R, Ramanathan, G. | | Deposit date: | 2018-04-30 | | Release date: | 2019-05-22 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Catechol 2,3-dioxygenase (C23O64) from Diaphorobacter sp DS2
To Be Published
|
|
7BKX
 
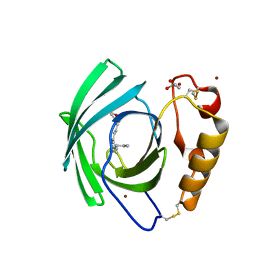 | | Diploptera punctata inspired lipocalin-like Milk protein expressed in Saccharomyces cerevisiae | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Banerjee, S, Kanagavijayan, D, Subramanian, R, Santhakumari, P.R, Chavas, L.M.G, Ramaswamy, S. | | Deposit date: | 2021-01-17 | | Release date: | 2021-12-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure of recombinantly expressed cockroach Lili-Mip protein in glycosylated and deglycosylated forms.
Biochim Biophys Acta Gen Subj, 1866, 2022
|
|
7RX0
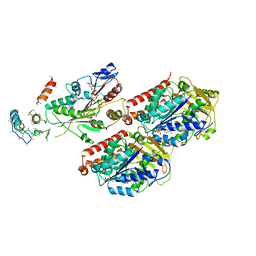 
 | | Complex of AMPPNP-Kif7 and Gli2 Zinc-Finger domain bound to microtubules | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, Kinesin-like protein KIF7, MAGNESIUM ION, ... | | Authors: | Mani, N, Wilson-Kubalek, E.M, Haque, F, Freniere, C, Milligan, R.A, Subramanian, R. | | Deposit date: | 2021-08-20 | | Release date: | 2022-06-15 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Cytoskeletal regulation of a transcription factor by DNA mimicry via coiled-coil interactions.
Nat.Cell Biol., 24, 2022
|
|
8G4V
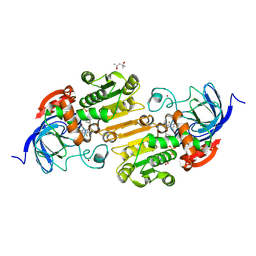 
 | | Horse liver alcohol dehydrogense His-51-Gln form complexed with NAD+ and 2,3,4,5,6-pentafluorobenzyl alcohol | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2,3,4,5,6-PENTAFLUOROBENZYL ALCOHOL, Alcohol dehydrogenase E chain, ... | | Authors: | Plapp, B.V, Subramanian, R. | | Deposit date: | 2023-02-10 | | Release date: | 2023-02-22 | | Last modified: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Histidine-51 facilitates deprotonation of the zinc-bound ligand during catalysis by horse liver alcohol dehydrogenase
To Be Published
|
|
7W8J
 
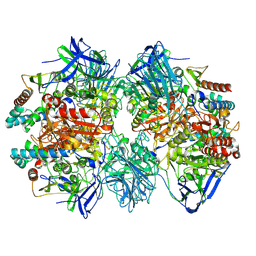 | | Dimethylformamidase, 2x(A2B2) | | Descriptor: | FE (III) ION, N,N-dimethylformamidase large subunit, N,N-dimethylformamidase small subunit | | Authors: | Vinothkumar, K.R, Subramanian, R, Arya, C, Ramanathan, G. | | Deposit date: | 2021-12-07 | | Release date: | 2022-04-06 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Dimethylformamidase with a Unique Iron Center
To Be Published
|
|
1MX4
 
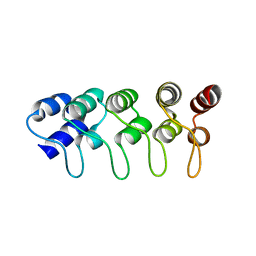 | | Structure of p18INK4c (F82Q) | | Descriptor: | Cyclin-dependent kinase 6 inhibitor | | Authors: | Marmorstein, R, Venkataramani, R.N, MacLachlan, T.K, Chai, X, El-Deiry, W.S. | | Deposit date: | 2002-10-01 | | Release date: | 2002-10-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-based design of p18INK4c proteins with increased thermodynamic stability and cell cycle inhibitory activity
J.Biol.Chem., 277, 2002
|
|
1MX2
 
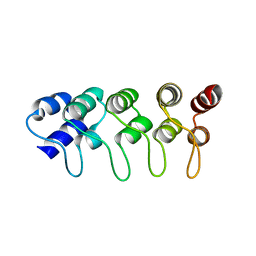 | | Structure of F71N mutant of p18INK4c | | Descriptor: | Cyclin-dependent kinase 6 inhibitor | | Authors: | Marmorstein, R, Venkataramani, R.N, MacLachlan, T.K, Chai, X, El-Deiery, W.S. | | Deposit date: | 2002-10-01 | | Release date: | 2002-10-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure-based design of p18INK4c proteins with increased thermodynamic stability and cell cycle inhibitory activity
J.Biol.Chem., 277, 2002
|
|
1MX6
 
 | | Structure of p18INK4c (F92N) | | Descriptor: | Cyclin-dependent kinase 6 inhibitor | | Authors: | Marmorstein, R, Venkataramani, R.N, MacLachlan, T.K, Chai, X, El-Deiry, W.S. | | Deposit date: | 2002-10-01 | | Release date: | 2002-10-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-based design of p18INK4c proteins with increased thermodynamic stability and cell cycle inhibitory activity
J.Biol.Chem., 277, 2002
|
|
