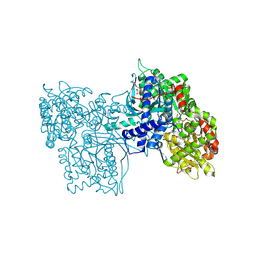
6GPB
 
 | |
2QIL
 
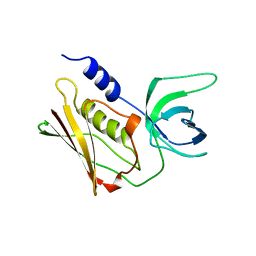 | |
1QIL
 
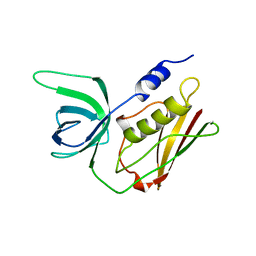 | |
1STE
 
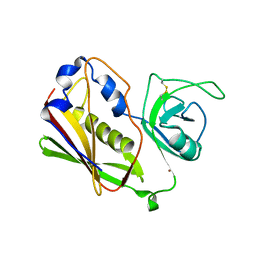 | |
1ANG
 
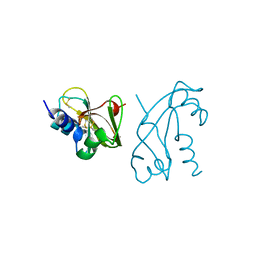 | | CRYSTAL STRUCTURE OF HUMAN ANGIOGENIN REVEALS THE STRUCTURAL BASIS FOR ITS FUNCTIONAL DIVERGENCE FROM RIBONUCLEASE | | Descriptor: | ANGIOGENIN | | Authors: | Acharya, K.R, Allen, S, Shapiro, R, Riordan, J.F, Vallee, B.L. | | Deposit date: | 1994-01-18 | | Release date: | 1995-04-20 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of human angiogenin reveals the structural basis for its functional divergence from ribonuclease.
Proc.Natl.Acad.Sci.USA, 91, 1994
|
|
1BBT
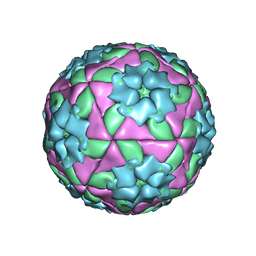 
 | | METHODS USED IN THE STRUCTURE DETERMINATION OF FOOT AND MOUTH DISEASE VIRUS | | Descriptor: | FOOT-AND-MOUTH DISEASE VIRUS (SUBUNIT VP1), FOOT-AND-MOUTH DISEASE VIRUS (SUBUNIT VP2), FOOT-AND-MOUTH DISEASE VIRUS (SUBUNIT VP3), ... | | Authors: | Acharya, K.R, Fry, E.E, Logan, D.T, Stuart, D.I. | | Deposit date: | 1992-05-18 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Methods used in the structure determination of foot-and-mouth disease virus.
Acta Crystallogr.,Sect.A, 49, 1993
|
|
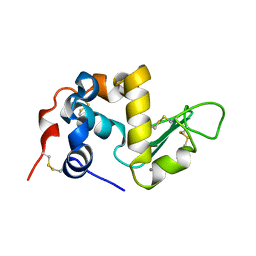
1ALC
 
 | |
1LCL
 
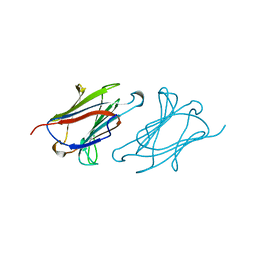 | | CHARCOT-LEYDEN CRYSTAL PROTEIN | | Descriptor: | LYSOPHOSPHOLIPASE | | Authors: | Acharya, K.R, Leonidas, D.D. | | Deposit date: | 1996-01-18 | | Release date: | 1997-01-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of human Charcot-Leyden crystal protein, an eosinophil lysophospholipase, identifies it as a new member of the carbohydrate-binding family of galectins.
Structure, 3, 1995
|
|
1AGI
 
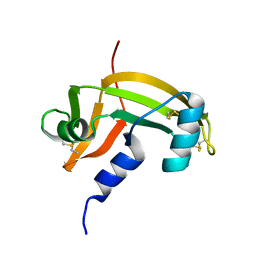 | |
6XVP
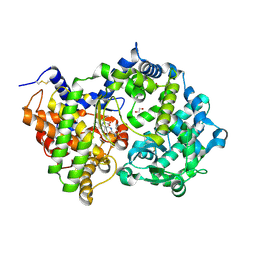 
 | | Crystal structure of Neprilysin in complex with Sampatrilat. | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Neprilysin, ... | | Authors: | Cozier, G.E, Acharya, K.R, Sharma, U. | | Deposit date: | 2020-01-22 | | Release date: | 2020-05-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Molecular Basis for Omapatrilat and Sampatrilat Binding to Neprilysin-Implications for Dual Inhibitor Design with Angiotensin-Converting Enzyme.
J.Med.Chem., 63, 2020
|
|
2GAL
 
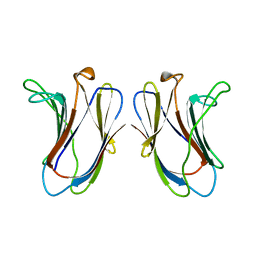 | |
5A2R
 
 | | A New Crystal Structure of the Drosophila melanogaster Angiotensin Converting Enzyme Homologue AnCE. | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ANGIOTENSIN-CONVERTING ENZYME, ... | | Authors: | Harrison, C, Acharya, K.R. | | Deposit date: | 2015-05-22 | | Release date: | 2015-08-26 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A New High-Resolution Crystal Structure of the Drosophila Melanogaster Angiotensin Converting Enzyme Homologue, Ance.
FEBS Open Bio, 5, 2015
|
|
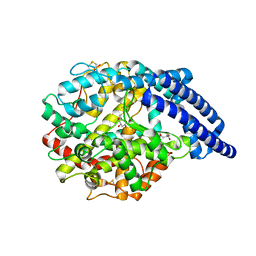
8OW8
 
 | | Crystal Structure of the Catalytic Domain of a Botulinum Neurotoxin Homologue from Enterococcus faecium | | Descriptor: | 1,2-ETHANEDIOL, Botulinum-like toxin eBoNT/J light chain, PHOSPHATE ION, ... | | Authors: | Gregory, K.S, Acharya, K.R, Liu, S.M. | | Deposit date: | 2023-04-27 | | Release date: | 2023-08-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Catalytic Domain of a Botulinum Neurotoxin Homologue from Enterococcus faecium : Potential Insights into Substrate Recognition.
Int J Mol Sci, 24, 2023
|
|
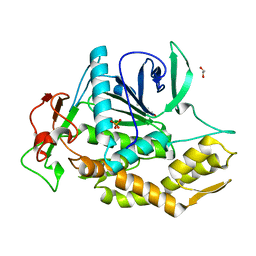
8AGK
 
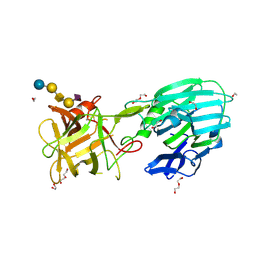 | | Botulinum neurotoxin subtype A6 cell binding domain in complex with GD1a ganglioside | | Descriptor: | 1,2-ETHANEDIOL, Bont/A1, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Gregory, K.S, Acharya, K.R, Liu, S.M, Newell, A.R, Mojanaga, O.O. | | Deposit date: | 2022-07-20 | | Release date: | 2022-09-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structures of the Clostridium botulinum Neurotoxin A6 Cell Binding Domain Alone and in Complex with GD1a Reveal Significant Conformational Flexibility.
Int J Mol Sci, 23, 2022
|
|
8ALP
 
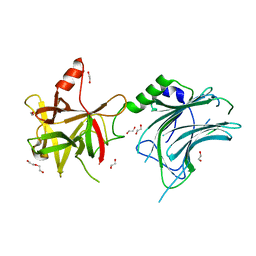 | | Botulinum neurotoxin A6 cell binding domain crystal form II | | Descriptor: | 1,2-ETHANEDIOL, Bont/A1, DI(HYDROXYETHYL)ETHER | | Authors: | Gregory, K.S, Acharya, K.R, Liu, S.M. | | Deposit date: | 2022-08-01 | | Release date: | 2022-09-21 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structures of the Clostridium botulinum Neurotoxin A6 Cell Binding Domain Alone and in Complex with GD1a Reveal Significant Conformational Flexibility.
Int J Mol Sci, 23, 2022
|
|
4GAL
 
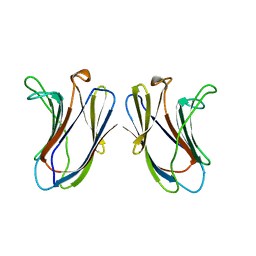 | |
2SKE
 
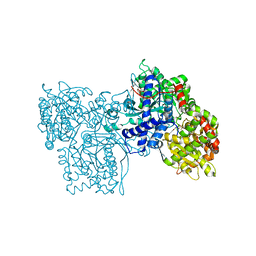 | | PYRIDOXAL PHOSPHORYLASE B IN COMPLEX WITH PHOSPHITE, GLUCOSE AND INOSINE-5'-MONOPHOSPHATE | | Descriptor: | INOSINIC ACID, PHOSPHITE ION, PYRIDOXAL PHOSPHORYLASE B, ... | | Authors: | Oikonomakos, N.G, Zographos, S.E, Tsitsanou, K.E, Johnson, L.N, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Activator anion binding site in pyridoxal phosphorylase b: the binding of phosphite, phosphate, and fluorophosphate in the crystal.
Protein Sci., 5, 1996
|
|
2SKC
 
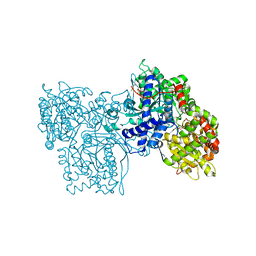 | | PYRIDOXAL PHOSPHORYLASE B IN COMPLEX WITH FLUOROPHOSPHATE, GLUCOSE AND INOSINE-5'-MONOPHOSPHATE | | Descriptor: | FLUORO-PHOSPHITE ION, INOSINIC ACID, PYRIDOXAL PHOSPHORYLASE B, ... | | Authors: | Oikonomakos, N.G, Zographos, S.E, Tsitsanou, K.E, Johnson, L.N, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Activator anion binding site in pyridoxal phosphorylase b: the binding of phosphite, phosphate, and fluorophosphate in the crystal.
Protein Sci., 5, 1996
|
|
2SKD
 
 | | PYRIDOXAL PHOSPHORYLASE B IN COMPLEX WITH PHOSPHATE, GLUCOSE AND INOSINE-5'-MONOPHOSPHATE | | Descriptor: | INOSINIC ACID, PHOSPHATE ION, PYRIDOXAL PHOSPHORYLASE B, ... | | Authors: | Oikonomakos, N.G, Zographos, S.E, Tsitsanou, K.E, Johnson, L.N, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Activator anion binding site in pyridoxal phosphorylase b: the binding of phosphite, phosphate, and fluorophosphate in the crystal.
Protein Sci., 5, 1996
|
|
1W4Q
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | 2'-FLUORO-2'-DEOXYURIDINE 3'-MONOPHOSPHATE, PANCREATIC RIBONUCLEASE A | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
1W4O
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | PANCREATIC RIBONUCLEASE A, URACIL ARABINOSE-3'-PHOSPHATE | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
1W4P
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | 2'-DEOXYURIDINE 3'-MONOPHOSPHATE, PANCREATIC RIBONUCLEASE A | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
5OQ3
 
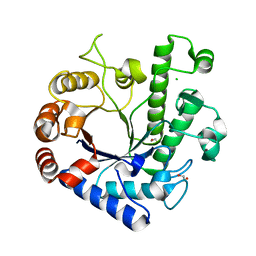 | | High resolution structure of the functional region of Cwp19 from Clostridium difficile | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Cwp19, ... | | Authors: | Bradshaw, W.J, Kirby, J.M, Roberts, A.K, Shone, C.C, Acharya, K.R. | | Deposit date: | 2017-08-10 | | Release date: | 2017-11-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | The molecular structure of the glycoside hydrolase domain of Cwp19 from Clostridium difficile.
FEBS J., 284, 2017
|
|
5OQ2
 
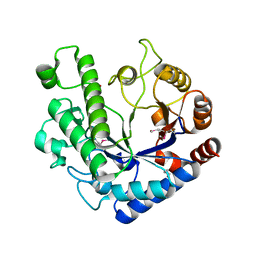 | | Se-SAD structure of the functional region of Cwp19 from Clostridium difficile | | Descriptor: | 1,2-ETHANEDIOL, Cwp19, PHOSPHATE ION | | Authors: | Bradshaw, W.J, Kirby, J.M, Roberts, A.K, Shone, C.C, Acharya, K.R. | | Deposit date: | 2017-08-10 | | Release date: | 2017-11-01 | | Last modified: | 2017-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The molecular structure of the glycoside hydrolase domain of Cwp19 from Clostridium difficile.
FEBS J., 284, 2017
|
|
3SEB
 
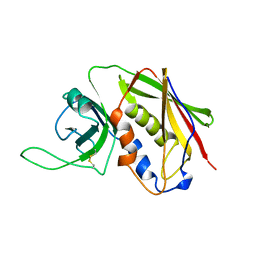 | | STAPHYLOCOCCAL ENTEROTOXIN B | | Descriptor: | STAPHYLOCOCCAL ENTEROTOXIN B | | Authors: | Papageorgiou, A.C, Acharya, K.R. | | Deposit date: | 1997-11-26 | | Release date: | 1998-05-27 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Crystal structure of microbial superantigen staphylococcal enterotoxin B at 1.5 A resolution: implications for superantigen recognition by MHC class II molecules and T-cell receptors.
J.Mol.Biol., 277, 1998
|
|
