3V6Z
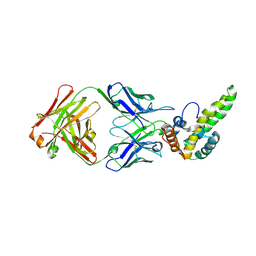 
 | | Crystal Structure of Hepatitis B Virus e-antigen | | Descriptor: | Fab e6 Heavy Chain, Fab e6 Light Chain, e-antigen | | Authors: | Dimattia, M.A, Watts, N.R, Stahl, S.J, Grimes, J.M, Steven, A.C, Stuart, D.I, Wingfield, P.T. | | Deposit date: | 2011-12-20 | | Release date: | 2013-02-06 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (3.34 Å) | | Cite: | Antigenic switching of hepatitis B virus by alternative dimerization of the capsid protein.
Structure, 21, 2013
|
|
3V6F
 
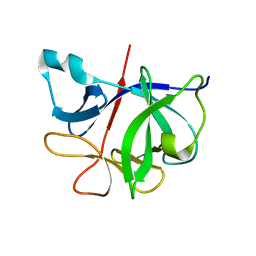
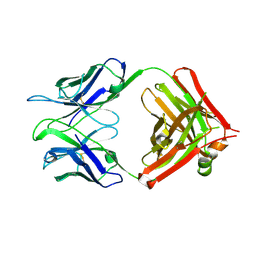 | | Crystal Structure of an anti-HBV e-antigen monoclonal Fab fragment (e6), unbound | | Descriptor: | Fab e6 Heavy Chain, Fab e6 Light Chain | | Authors: | Dimattia, M.A, Watts, N.R, Stahl, S.J, Grimes, J.M, Steven, A.C, Stuart, D.I, Wingfield, P.T. | | Deposit date: | 2011-12-19 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.52 Å) | | Cite: | Antigenic switching of hepatitis B virus by alternative dimerization of the capsid protein.
Structure, 21, 2013
|
|
1QBZ
 
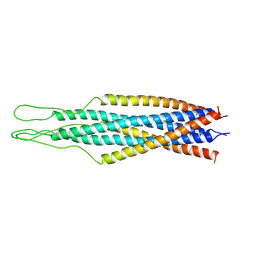
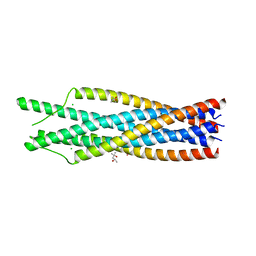 | | THE CRYSTAL STRUCTURE OF THE SIV GP41 ECTODOMAIN AT 1.47 A | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, CHLORIDE ION, MERCURY (II) ION, ... | | Authors: | Yang, Z.-N, Mueser, T.C, Kaufman, J, Stahl, S.J, Wingfield, P.T, Hyde, C.C. | | Deposit date: | 1999-04-28 | | Release date: | 1999-05-17 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | The crystal structure of the SIV gp41 ectodomain at 1.47 A resolution.
J.Struct.Biol., 126, 1999
|
|
1EY1
 
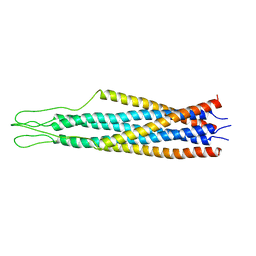
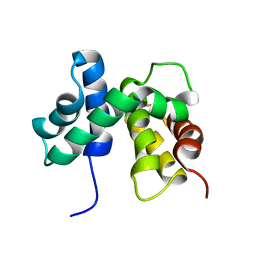 | | SOLUTION STRUCTURE OF ESCHERICHIA COLI NUSB | | Descriptor: | ANTITERMINATION FACTOR NUSB | | Authors: | Altieri, A.S, Mazzulla, M.J, Horita, D.A, Coats, R.H, Wingfield, P.T, Byrd, R.A. | | Deposit date: | 2000-05-05 | | Release date: | 2000-06-14 | | Last modified: | 2022-06-15 | | Method: | SOLUTION NMR | | Cite: | The structure of the transcriptional antiterminator NusB from Escherichia coli.
Nat.Struct.Biol., 7, 2000
|
|
6I1B
 | |
7I1B
 
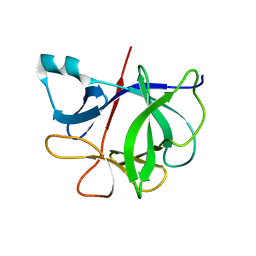 | |
4TRX
 
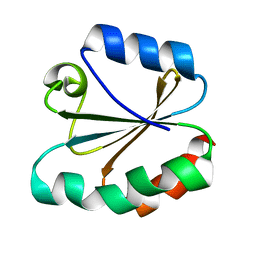 | |
2EZQ
 | |
3TRX
 
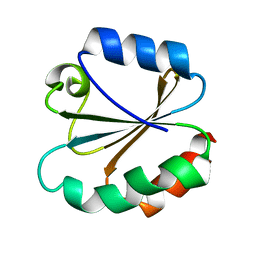 | |
2JR0
 
 | | Solution structure of NusB from Aquifex Aeolicus | | Descriptor: | N utilization substance protein B homolog | | Authors: | Das, R, Loss, S, Li, J, Tarasov, S, Wingfield, P, Waugh, D.S, Byrd, R.A, Altieri, A.S. | | Deposit date: | 2007-06-18 | | Release date: | 2008-02-19 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | Structural biophysics of the NusB:NusE antitermination complex.
J.Mol.Biol., 376, 2008
|
|
2EZP
 | |
2EZS
 | |
2EZO
 | |
2EZR
 
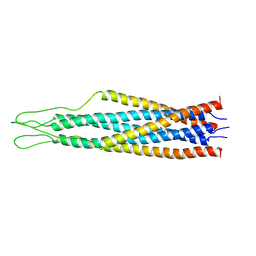 | |
1QA4
 | |
1QCE
 | |
7M18
 
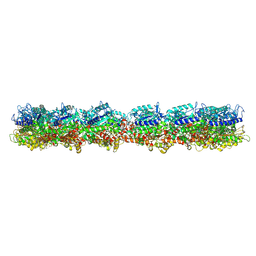 | | HeLa-tubulin in complex with cryptophycin 1 | | Descriptor: | Cryptophycin 1, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Eren, E. | | Deposit date: | 2021-03-12 | | Release date: | 2021-09-08 | | Last modified: | 2021-10-06 | | Method: | ELECTRON MICROSCOPY (3.38 Å) | | Cite: | Conformational changes in tubulin upon binding cryptophycin-52 reveal its mechanism of action.
J.Biol.Chem., 297, 2021
|
|
7M20
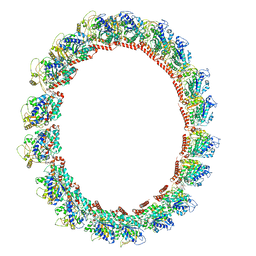 
 | | 18-mer HeLa-tubulin rings in complex with Cryptophycin 1 | | Descriptor: | Cryptophycin 1, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Eren, E. | | Deposit date: | 2021-03-15 | | Release date: | 2021-09-08 | | Last modified: | 2021-10-06 | | Method: | ELECTRON MICROSCOPY (3.84 Å) | | Cite: | Conformational changes in tubulin upon binding cryptophycin-52 reveal its mechanism of action.
J.Biol.Chem., 297, 2021
|
|
7LXB
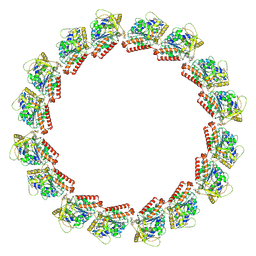 
 | | HeLa-tubulin in complex with cryptophycin 52 | | Descriptor: | Cryptophycin 52, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Eren, E. | | Deposit date: | 2021-03-03 | | Release date: | 2021-09-08 | | Last modified: | 2021-10-06 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Conformational changes in tubulin upon binding cryptophycin-52 reveal its mechanism of action.
J.Biol.Chem., 297, 2021
|
|
2NVH
 
 | |
2GB1
 
 | |
1QA5
 
 | |
5KAO
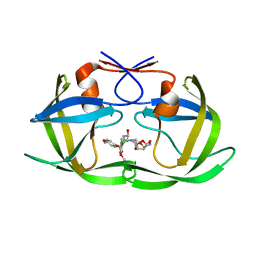 
 | | Crystal structure of wild type HIV-1 protease in complex with GRL-10413 | | Descriptor: | [(3~{a}~{S},4~{R},6~{a}~{R})-2,3,3~{a},4,5,6~{a}-hexahydrofuro[2,3-b]furan-4-yl] ~{N}-[(2~{S},3~{R})-1-(3-chloranyl-4-methoxy-phenyl)-4-[(4-methoxyphenyl)sulfonyl-(2-methylpropyl)amino]-3-oxidanyl-butan-2-yl]carbamate, protease | | Authors: | Yedidi, R.S, Delino, N.S, Das, D, Kaufman, J.D, Wingfield, P.T, Ghosh, A.K, Mitsuya, H. | | Deposit date: | 2016-06-01 | | Release date: | 2016-08-31 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A Modified P1 Moiety Enhances In Vitro Antiviral Activity against Various Multidrug-Resistant HIV-1 Variants and In Vitro Central Nervous System Penetration Properties of a Novel Nonpeptidic Protease Inhibitor, GRL-10413.
Antimicrob.Agents Chemother., 60, 2016
|
|
3GB1
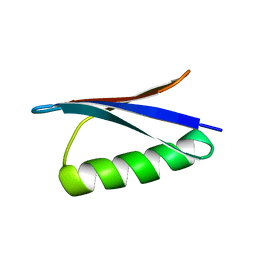 
 | | STRUCTURES OF B1 DOMAIN OF STREPTOCOCCAL PROTEIN G | | Descriptor: | PROTEIN (B1 DOMAIN OF STREPTOCOCCAL PROTEIN G) | | Authors: | Clore, G.M. | | Deposit date: | 1999-05-02 | | Release date: | 1999-06-23 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Improving the Packing and Accuracy of NMR Structures with a Pseudopotential for the Radius of Gyration
J.Am.Chem.Soc., 121, 1999
|
|
7S5C
 
 | | M. xanthus ferritin-like protein EncB | | Descriptor: | CALCIUM ION, EncB, FE (III) ION | | Authors: | Eren, E. | | Deposit date: | 2021-09-10 | | Release date: | 2022-02-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structural characterization of the Myxococcus xanthus encapsulin and ferritin-like cargo system gives insight into its iron storage mechanism.
Structure, 30, 2022
|
|
