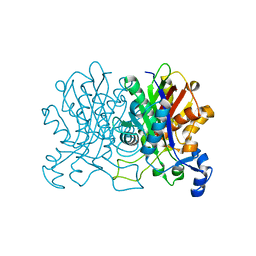
4YUF
 
 | |
4YUC
 
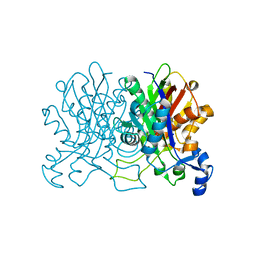 | | Crystal Structure of CorB derivatized with S-(2-acetamidoethyl) 4-methyl-3-oxohexanethioate | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CorB, SODIUM ION | | Authors: | Zocher, G, Vilstrup, J, Stehle, T. | | Deposit date: | 2015-03-18 | | Release date: | 2016-03-30 | | Last modified: | 2017-08-23 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | Structural basis of head to head polyketide fusion by CorB.
Chem Sci, 6, 2015
|
|
3TW5
 
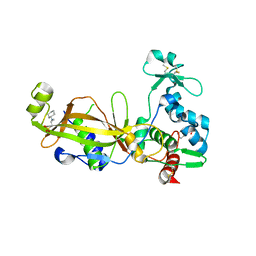 | | Crystal structure of the GP42 transglutaminase from Phytophthora sojae | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, Transglutaminase elicitor | | Authors: | Reiss, K, Kirchner, E, Zocher, G, Stehle, T. | | Deposit date: | 2011-09-21 | | Release date: | 2011-10-12 | | Last modified: | 2020-10-21 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural and Phylogenetic Analyses of the GP42 Transglutaminase from Phytophthora sojae Reveal an Evolutionary Relationship between Oomycetes and Marine Vibrio Bacteria.
J.Biol.Chem., 286, 2011
|
|
4ZH1
 
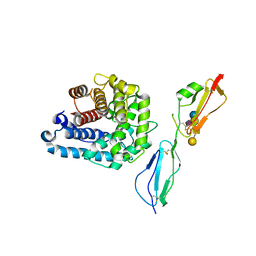 | | Complement factor H in complex with the GM1 glycan | | Descriptor: | Complement C3, Complement factor H, GLYCEROL, ... | | Authors: | Blaum, B.S, Stehle, T. | | Deposit date: | 2015-04-24 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Complement Factor H and Simian Virus 40 bind the GM1 ganglioside in distinct conformations.
Glycobiology, 26, 2016
|
|
4GR4
 
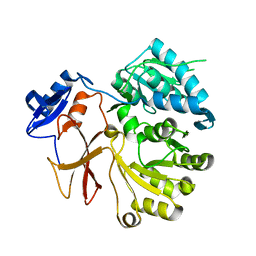 | | Crystal structure of SlgN1deltaAsub | | Descriptor: | CHLORIDE ION, Non-ribosomal peptide synthetase | | Authors: | Herbst, D.A, Zocher, G, Stehle, T. | | Deposit date: | 2012-08-24 | | Release date: | 2012-11-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structural Basis of the Interaction of MbtH-like Proteins, Putative Regulators of Nonribosomal Peptide Biosynthesis, with Adenylating Enzymes.
J.Biol.Chem., 288, 2013
|
|
4GR5
 
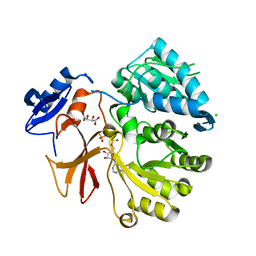 | | Crystal structure of SlgN1deltaAsub in complex with AMPcPP | | Descriptor: | CHLORIDE ION, DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, L(+)-TARTARIC ACID, ... | | Authors: | Herbst, D.A, Zocher, G, Stehle, T. | | Deposit date: | 2012-08-24 | | Release date: | 2012-11-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Structural Basis of the Interaction of MbtH-like Proteins, Putative Regulators of Nonribosomal Peptide Biosynthesis, with Adenylating Enzymes.
J.Biol.Chem., 288, 2013
|
|
4KNL
 | |
4KNK
 
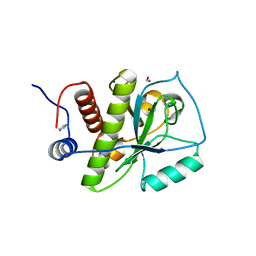
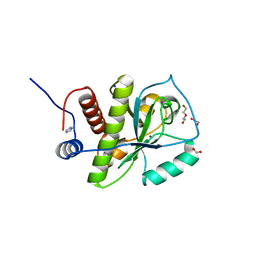 | | Crystal structure of Staphylococcus aureus hydrolase AmiA | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional autolysin, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Buettner, F.M, Zoll, S, Stehle, T. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.124 Å) | | Cite: | Structure-function analysis of Staphylococcus aureus amidase reveals the determinants of peptidoglycan recognition and cleavage.
J.Biol.Chem., 289, 2014
|
|
4LD7
 
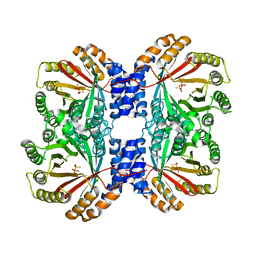 | | Crystal structure of AnaPT from Neosartorya fischeri | | Descriptor: | Dimethylallyl tryptophan synthase, SODIUM ION, TRIHYDROGEN THIODIPHOSPHATE | | Authors: | Zocher, G, Stehle, T. | | Deposit date: | 2013-06-24 | | Release date: | 2013-12-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Catalytic Mechanism of Stereospecific Formation of cis-Configured Prenylated Pyrroloindoline Diketopiperazines by Indole Prenyltransferases.
Chem.Biol., 20, 2013
|
|
4LZU
 
 | |
4LZV
 
 | |
4ODB
 
 | |
4PCG
 
 | | Structure of Human Polyomavirus 6 (HPyV6) VP1 pentamer | | Descriptor: | SODIUM ION, THIOCYANATE ION, VP1 | | Authors: | Stroh, L.J, Stehle, T. | | Deposit date: | 2014-04-15 | | Release date: | 2014-08-06 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure analysis of the major capsid proteins of human polyomaviruses 6 and 7 reveals an obstructed sialic Acid binding site.
J.Virol., 88, 2014
|
|
4PCH
 | |
4Q4W
 
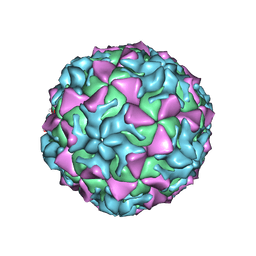 | |
6GAO
 | |
6GAP
 
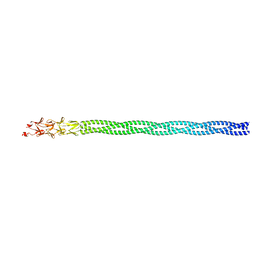 | |
6GAK
 | |
6GAJ
 | |
6H1J
 
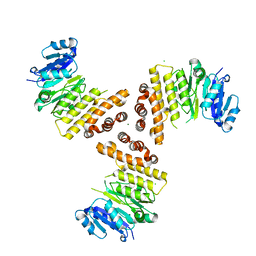 | | TarP native | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Probable ss-1,3-N-acetylglucosaminyltransferase | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-07-11 | | Release date: | 2018-09-26 | | Last modified: | 2018-12-12 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
6H2N
 
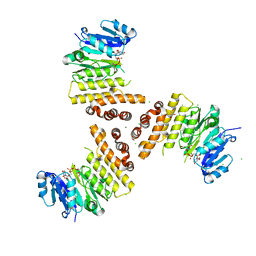 | | TarP-UDP-GlcNAc-Mg | | Descriptor: | CHLORIDE ION, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-07-13 | | Release date: | 2018-09-26 | | Last modified: | 2018-12-12 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
6H21
 
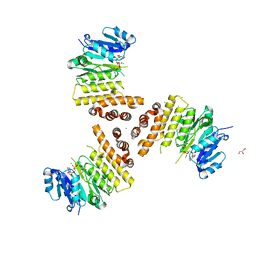 | | TarP-UDP-GlcNAc-Mn | | Descriptor: | CHLORIDE ION, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-07-12 | | Release date: | 2018-09-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
6H4F
 
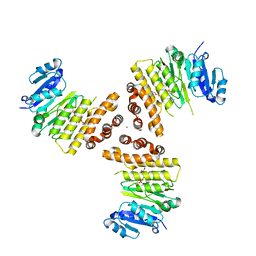 | | TarP-3RboP | | Descriptor: | CHLORIDE ION, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-07-21 | | Release date: | 2018-09-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
6H4M
 
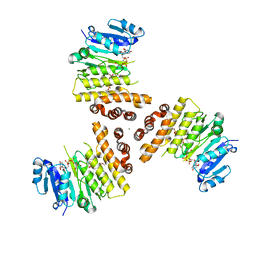 | | TarP-UDP-GlcNAc-3RboP | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Probable ss-1,3-N-acetylglucosaminyltransferase, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-07-22 | | Release date: | 2018-09-26 | | Last modified: | 2018-12-12 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
6HNQ
 
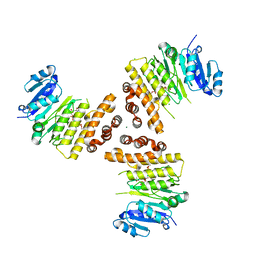 | | TarP-6RboP-(CH2)6NH2 | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, Probable ss-1,3-N-acetylglucosaminyltransferase, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2018-09-17 | | Release date: | 2018-10-24 | | Last modified: | 2018-12-12 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity.
Nature, 563, 2018
|
|
