1KXS
 
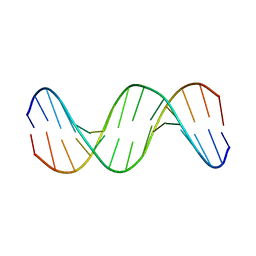 | |
4MGW
 
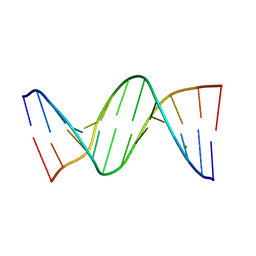 | | Comparison of the structural and dynamic effects of 5-methylcytosine and 5-chlorocytosine in a CpG dinucleotide sequence | | Descriptor: | ClC containing Dickerson-Drew dodecamer, MAGNESIUM ION | | Authors: | Theruvathu, J.A, Whitney, Y, Montgomery Pettitt, B, Sowers, L.C. | | Deposit date: | 2013-08-29 | | Release date: | 2013-11-06 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Comparison of the structural and dynamic effects of 5-methylcytosine and 5-chlorocytosine in a CpG dinucleotide sequence.
Biochemistry, 52, 2013
|
|
4MKW
 | |
4GSG
 
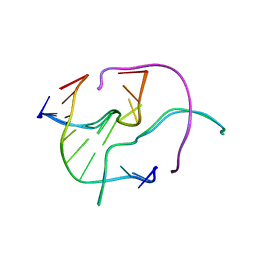 | |
4GQD
 
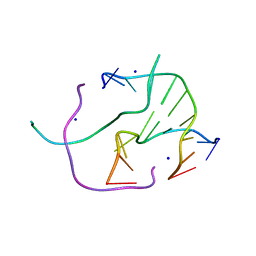 | | DNA Holliday junction stabilized by chlorine halogen bond. | | Descriptor: | DNA (5'-D(*CP*CP*GP*AP*TP*AP*CP*CP*GP*G)-3'), DNA (5'-D(*CP*CP*GP*GP*TP*AP*(UCL)P*CP*GP*G)-3'), SODIUM ION | | Authors: | Carter, M, Ho, P.S. | | Deposit date: | 2012-08-22 | | Release date: | 2013-07-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Enthalpy-entropy compensation in biomolecular halogen bonds measured in DNA junctions.
Biochemistry, 52, 2013
|
|
4GRE
 | |
4GSI
 | |
4GS2
 | |
1FKZ
 
 | | NMR STUDY OF B-DNA CONTAINING A MISMATCHED BASE PAIR IN THE 29-39 K-RAS GENE SEQUENCE: CC CT C+C C+T, 2 STRUCTURES | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*TP*CP*GP*TP*GP*GP*C)-3'), DNA (5'-D(*GP*CP*CP*AP*CP*CP*AP*GP*CP*TP*C)-3') | | Authors: | Boulard, Y, Cognet, J.A.H, Fazakerley, G.V. | | Deposit date: | 1996-10-09 | | Release date: | 1997-03-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure as a function of pH of two central mismatches, C . T and C . C, in the 29 to 39 K-ras gene sequence, by nuclear magnetic resonance and molecular dynamics.
J.Mol.Biol., 268, 1997
|
|
1FKY
 
 | | NMR STUDY OF B-DNA CONTAINING A MISMATCHED BASE PAIR IN THE 29-39 K-RAS GENE SEQUENCE: CC CT C+C C+T, 2 STRUCTURES | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*TP*TP*GP*TP*GP*GP*C)-3'), DNA (5'-D(*GP*CP*CP*AP*CP*CP*AP*GP*CP*TP*C)-3') | | Authors: | Boulard, Y, Cognet, J.A.H, Fazakerley, G.V. | | Deposit date: | 1996-10-09 | | Release date: | 1997-04-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure as a function of pH of two central mismatches, C . T and C . C, in the 29 to 39 K-ras gene sequence, by nuclear magnetic resonance and molecular dynamics.
J.Mol.Biol., 268, 1997
|
|
