2AL2
 
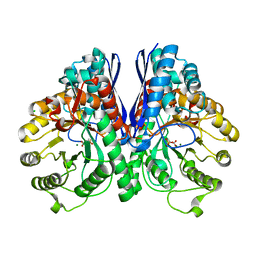 | | Crystal Structure Analysis of Enolase Mg Subunit Complex at pH 8.0 | | Descriptor: | 2-PHOSPHOGLYCERIC ACID, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Sims, P.A, Menefee, A.L, Larsen, T.M, Mansoorabadi, S.O, Reed, G.H. | | Deposit date: | 2005-08-04 | | Release date: | 2006-01-24 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure and catalytic properties of an engineered heterodimer of enolase composed of one active and one inactive subunit
J.Mol.Biol., 355, 2006
|
|
2AL1
 
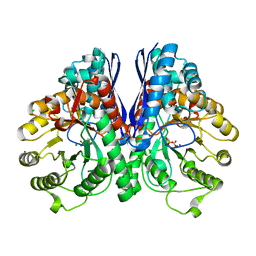 | | Crystal Structure Analysis of Enolase Mg Subunit Complex at pH 8.0 | | Descriptor: | 2-PHOSPHOGLYCERIC ACID, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Sims, P.A, Menefee, A.L, Larsen, T.M, Mansoorabadi, S.O, Reed, G.H. | | Deposit date: | 2005-08-04 | | Release date: | 2006-01-24 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure and catalytic properties of an engineered heterodimer of enolase composed of one active and one inactive subunit
J.Mol.Biol., 355, 2006
|
|
1P43
 
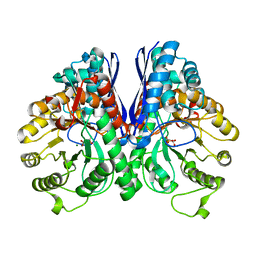 | | REVERSE PROTONATION IS THE KEY TO GENERAL ACID-BASE CATALYSIS IN ENOLASE | | Descriptor: | 2-PHOSPHOGLYCERIC ACID, Enolase 1, MAGNESIUM ION | | Authors: | Sims, P.A, Larsen, T.M, Poyner, R.R, Cleland, W.W, Reed, G.H. | | Deposit date: | 2003-04-21 | | Release date: | 2003-11-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Reverse protonation is the key to general acid-base catalysis in enolase
Biochemistry, 42, 2003
|
|
1P48
 
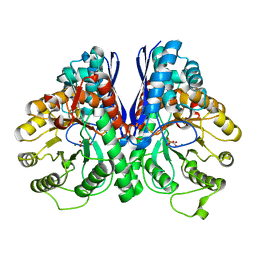 | | REVERSE PROTONATION IS THE KEY TO GENERAL ACID-BASE CATALYSIS IN ENOLASE | | Descriptor: | Enolase 1, MAGNESIUM ION, PHOSPHOENOLPYRUVATE | | Authors: | Sims, P.A, Larsen, T.M, Poyner, R.R, Cleland, W.W, Reed, G.H. | | Deposit date: | 2003-04-21 | | Release date: | 2003-11-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Reverse protonation is the key to general acid-base catalysis in enolase
Biochemistry, 42, 2003
|
|
4GKV
 
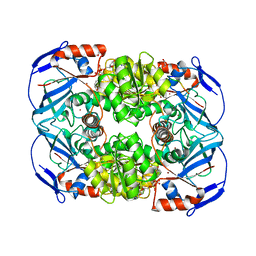 | | Structure of Escherichia coli AdhP (ethanol-inducible dehydrogenase) with bound NAD | | Descriptor: | Alcohol dehydrogenase, propanol-preferring, GLYCEROL, ... | | Authors: | Sims, P.A, Thomas, L.M, Harper, A.R, Miner, W.A, Ajufo, H.O, Branscrum, K.M, Kao, L. | | Deposit date: | 2012-08-13 | | Release date: | 2013-07-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.008 Å) | | Cite: | Structure of Escherichia coli AdhP (ethanol-inducible dehydrogenase) with bound NAD.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
