3GNR
 
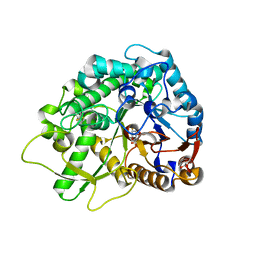 | | Crystal Structure of a Rice Os3BGlu6 Beta-Glucosidase with covalently bound 2-deoxy-2-fluoroglucoside to the catalytic nucleophile E396 | | Descriptor: | 2-deoxy-2-fluoro-alpha-D-glucopyranose, GLYCEROL, Os03g0212800 protein | | Authors: | Seshadri, S, Akiyama, T, Opassiri, R, Kuaprasert, B, Cairns, J.R.K. | | Deposit date: | 2009-03-17 | | Release date: | 2009-07-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structural and enzymatic characterization of Os3BGlu6, a rice beta-glucosidase hydrolyzing hydrophobic glycosides and (1->3)- and (1->2)-linked disaccharides.
Plant Physiol., 151, 2009
|
|
3GNO
 
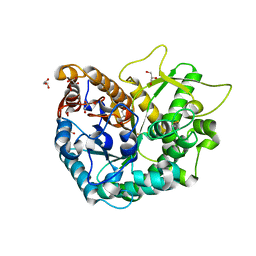 | | Crystal Structure of a Rice Os3BGlu6 Beta-Glucosidase | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLYCEROL, Os03g0212800 protein | | Authors: | Seshadri, S, Akiyama, T, Opassiri, R, Kuaprasert, B, Cairns, J.R.K. | | Deposit date: | 2009-03-17 | | Release date: | 2009-07-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural and enzymatic characterization of Os3BGlu6, a rice {beta}-glucosidase hydrolyzing hydrophobic glycosides and (1->3)- and (1->2)-linked disaccharides.
Plant Physiol., 2009
|
|
3GNP
 
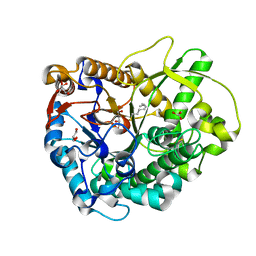 | | Crystal Structure of a Rice Os3BGlu6 Beta-Glucosidase with Octyl-Beta-D-Thio-Glucoside | | Descriptor: | GLYCEROL, Os03g0212800 protein, octyl 1-thio-beta-D-glucopyranoside | | Authors: | Seshadri, S, Akiyama, T, Opassiri, R, Kuaprasert, B, Cairns, J.R.K. | | Deposit date: | 2009-03-17 | | Release date: | 2009-07-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and enzymatic characterization of Os3BGlu6, a rice {beta}-glucosidase hydrolyzing hydrophobic glycosides and (1->3)- and (1->2)-linked disaccharides.
Plant Physiol., 2009
|
|
7SXP
 
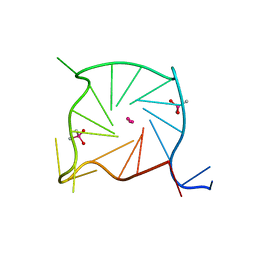 | |
6U7N
 
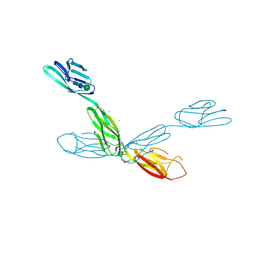 | | Crystal structure of neurotrimin (NTM) | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Neurotrimin, ... | | Authors: | Machius, M, Venkannagari, H, Misra, A, Rudenko, G, Rush, S. | | Deposit date: | 2019-09-03 | | Release date: | 2020-08-12 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.321 Å) | | Cite: | Highly Conserved Molecular Features in IgLONs Contrast Their Distinct Structural and Biological Outcomes.
J.Mol.Biol., 432, 2020
|
|
6U6T
 
 | | Neuronal growth regulator 1 (NEGR1) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Neuronal growth regulator 1, ... | | Authors: | Machius, M, Venkannagari, H, Misra, A, Rudenko, G. | | Deposit date: | 2019-08-30 | | Release date: | 2020-08-12 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Highly Conserved Molecular Features in IgLONs Contrast Their Distinct Structural and Biological Outcomes.
J.Mol.Biol., 432, 2020
|
|
5V5W
 
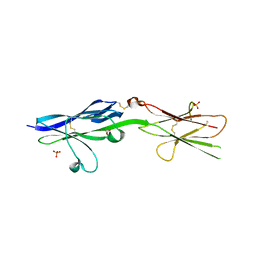 | |
5V5V
 
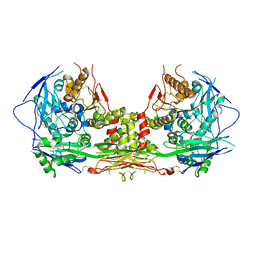 | | Complex of NLGN2 with MDGA1 Ig1-Ig2 | | Descriptor: | MAM domain-containing glycosylphosphatidylinositol anchor protein 1, Neuroligin-2 | | Authors: | Gangwar, S.P, Machius, M, Rudenko, G. | | Deposit date: | 2017-03-15 | | Release date: | 2017-07-05 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (4.11 Å) | | Cite: | Molecular Mechanism of MDGA1: Regulation of Neuroligin 2:Neurexin Trans-synaptic Bridges.
Neuron, 94, 2017
|
|
