1X7O
 
 | |
1X7P
 
 | |
1H7H
 
 | |
1GT7
 
 | | L-rhamnulose-1-phosphate aldolase from Escherichia coli | | Descriptor: | PHOSPHOGLYCOLOHYDROXAMIC ACID, RHAMNULOSE-1-PHOSPHATE ALDOLASE, ZINC ION | | Authors: | Kroemer, M, Schulz, G.E. | | Deposit date: | 2002-01-14 | | Release date: | 2002-05-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Structure of L-Rhamnulose-1-Phosphate Aldolase (Class II) Solved by Low-Resolution Sir Phasing and 20-Fold Ncs Averaging
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1H7F
 
 | |
1H7T
 
 | |
1H7G
 
 | |
1H7E
 
 | |
2SQC
 
 | |
1ZAK
 
 | |
2AG1
 
 | |
1OG1
 
 | | CRYSTAL STRUCTURE OF THE EUCARYOTIC MONO-ADP-RIBOSYLTRANSFERASE ART2.2 IN COMPLEX WITH TAD | | Descriptor: | BETA-METHYLENE-THIAZOLE-4-CARBOXYAMIDE-ADENINE DINUCLEOTIDE, T-CELL ECTO-ADP-RIBOSYLTRANSFERASE 2 | | Authors: | Ritter, H, Koch-Nolte, F, Marquez, V.E, Schulz, G.E. | | Deposit date: | 2003-04-23 | | Release date: | 2003-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Substrate Binding and Catalysis of Ecto-Adp-Ribosyltransferase 2.2 From Rat
Biochemistry, 42, 2003
|
|
1OG3
 
 | | Crystal structure of the eukaryotic mono-ADP-ribosyltransferase ART2.2 mutant E189I in complex with NAD | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, T-CELL ECTO-ADP-RIBOSYLTRANSFERASE 2 | | Authors: | Ritter, H, Koch-Nolte, F, Marquez, V.E, Schulz, G.E. | | Deposit date: | 2003-04-24 | | Release date: | 2003-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Substrate Binding and Catalysis of Ecto-Adp-Ribosyltransferase 2.2 From Rat
Biochemistry, 42, 2003
|
|
1PBG
 
 | |
1RPX
 
 | |
1TZL
 | | Crystal Structure of Pyranose 2-Oxidase from the White-Rot Fungus Peniophora sp. | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, pyranose oxidase | | Authors: | Bannwarth, M, Bastian, S, Heckmann-Pohl, D, Giffhorn, F, Schulz, G.E. | | Deposit date: | 2004-07-10 | | Release date: | 2004-10-19 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of pyranose 2-oxidase from the white-rot fungus peniophora sp.
Biochemistry, 43, 2004
|
|
1UKZ
 
 | |
1UUN
 
 | |
1UWK
 
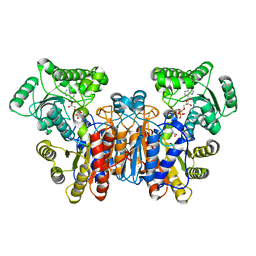 | |
1UWL
 
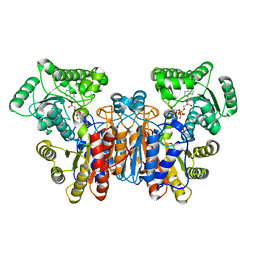 | |
3E1T
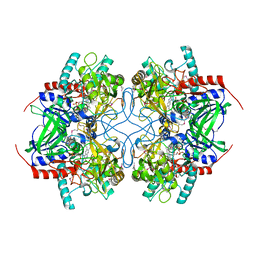
 | | Structure and action of the myxobacterial chondrochloren halogenase CndH, a new variant of FAD-dependent halogenases | | Descriptor: | CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, Halogenase | | Authors: | Buedenbender, S, Rachid, S, Mueller, R, Schulz, G.E. | | Deposit date: | 2008-08-04 | | Release date: | 2009-02-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure and action of the myxobacterial chondrochloren halogenase CndH: a new variant of FAD-dependent halogenases.
J.Mol.Biol., 385, 2009
|
|
3EDF
 
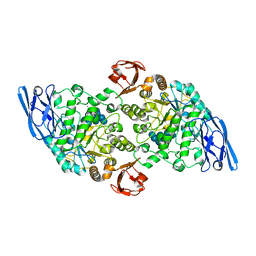 | | Structural base for cyclodextrin hydrolysis | | Descriptor: | CALCIUM ION, Cyclohexakis-(1-4)-(alpha-D-glucopyranose), Cyclomaltodextrinase, ... | | Authors: | Buedenbender, S, Schulz, G.E. | | Deposit date: | 2008-09-03 | | Release date: | 2009-03-03 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural base for enzymatic cyclodextrin hydrolysis
J.Mol.Biol., 385, 2009
|
|
3EDJ
 
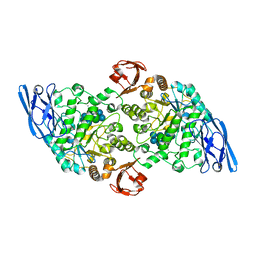 | | Structural base for cyclodextrin hydrolysis | | Descriptor: | CALCIUM ION, Cycloheptakis-(1-4)-(alpha-D-glucopyranose), Cyclomaltodextrinase, ... | | Authors: | Buedenbender, S, Schulz, G.E. | | Deposit date: | 2008-09-03 | | Release date: | 2009-03-03 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural base for enzymatic cyclodextrin hydrolysis
J.Mol.Biol., 385, 2009
|
|
3EDE
 
 | |
3EDK
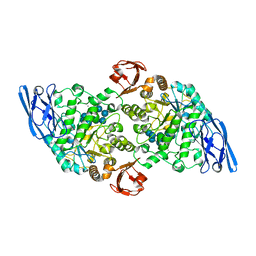 
 | | Structural base for cyclodextrin hydrolysis | | Descriptor: | CALCIUM ION, Cyclomaltodextrinase, Cyclooctakis-(1-4)-(alpha-D-glucopyranose), ... | | Authors: | Buedenbender, S, Schulz, G.E. | | Deposit date: | 2008-09-03 | | Release date: | 2009-03-03 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structural base for enzymatic cyclodextrin hydrolysis
J.Mol.Biol., 385, 2009
|
|
