1RPV
 
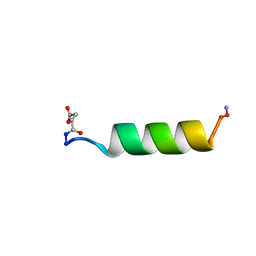 | | HIV-1 REV PROTEIN (RESIDUES 34-50) | | Descriptor: | HIV-1 REV PROTEIN | | Authors: | Scanlon, M.J, Fairlie, D.P, Craik, D.J, Englebretsen, D.R, West, M.L. | | Deposit date: | 1995-05-04 | | Release date: | 1995-10-15 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the RNA-binding peptide from human immunodeficiency virus (type 1) Rev.
Biochemistry, 34, 1995
|
|
1AV3
 
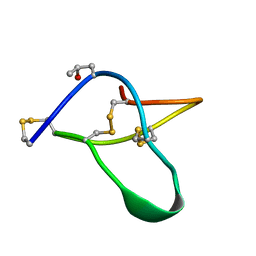 | | POTASSIUM CHANNEL BLOCKER KAPPA CONOTOXIN PVIIA FROM C. PURPURASCENS, NMR, 20 STRUCTURES | | Descriptor: | Kappa-conotoxin PVIIA | | Authors: | Scanlon, M.J, Naranjo, D, Thomas, L, Alewood, P.F, Lewis, R.J, Craik, D.J. | | Deposit date: | 1997-09-24 | | Release date: | 1998-10-14 | | Last modified: | 2025-03-26 | | Method: | SOLUTION NMR | | Cite: | Solution structure and proposed binding mechanism of a novel potassium channel toxin kappa-conotoxin PVIIA.
Structure, 5, 1997
|
|
1CE3
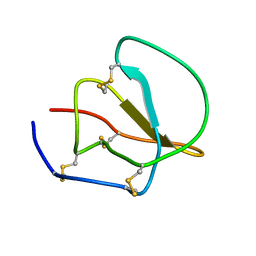 
 | |
6DO6
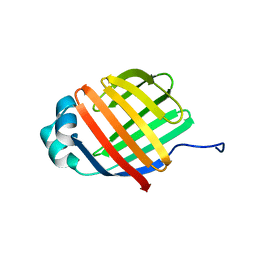 
 | | NMR solution structure of wild type apo hFABP1 at 308 K | | Descriptor: | Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-09 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
6DRG
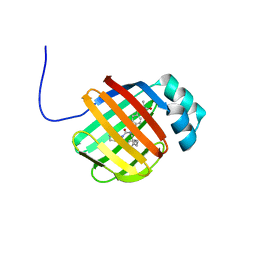 
 | | NMR solution structure of wild type hFABP1 with GW7647 | | Descriptor: | 2-[(4-{2-[(4-cyclohexylbutyl)(cyclohexylcarbamoyl)amino]ethyl}phenyl)sulfanyl]-2-methylpropanoic acid, Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-11 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
6DO7
 
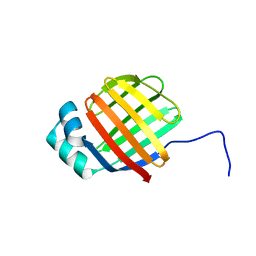 | | NMR solution structure of wild type hFABP1 with GW7647 | | Descriptor: | Fatty acid-binding protein, liver | | Authors: | Scanlon, M.J, Mohanty, B, Doak, B.C, Patil, R. | | Deposit date: | 2018-06-09 | | Release date: | 2019-01-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A ligand-induced structural change in fatty acid-binding protein 1 is associated with potentiation of peroxisome proliferator-activated receptor alpha agonists.
J. Biol. Chem., 294, 2019
|
|
1HX2
 
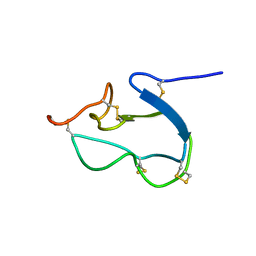 | | SOLUTION STRUCTURE OF BSTI, A TRYPSIN INHIBITOR FROM BOMBINA BOMBINA. | | Descriptor: | BSTI | | Authors: | Rosengren, K.J, Daly, N.L, Scanlon, M.J, Craik, D.J. | | Deposit date: | 2001-01-11 | | Release date: | 2001-01-24 | | Last modified: | 2024-10-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of BSTI: a new trypsin inhibitor from skin secretions of Bombina bombina.
Biochemistry, 40, 2001
|
|
1QH2
 
 | |
5QKC
 
 | |
5QKS
 
 | |
5QL9
 
 | |
5QLS
 
 | |
5QM7
 
 | |
5QKM
 
 | |
5QL1
 
 | |
5QLK
 
 | |
5QLZ
 
 | |
5QKI
 
 | |
5QKT
 
 | |
5QLE
 
 | |
5QLR
 
 | |
5QM3
 
 | |
5QMH
 
 | |
5QMY
 
 | |
9EGR
 
 | | Crystal structure of oxidised E.coli DsbA in complex with allene | | Descriptor: | Thiol:disulfide interchange protein DsbA, methyl 4-{3-[(5-methyl-1,2-oxazole-3-carbonyl)amino]phenyl}butanoate | | Authors: | Balaji, G.R, Tasdan, Y, Ilyichova, O.V, Akhtar, N, Scanlon, M.J. | | Deposit date: | 2024-11-21 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Crystal structure of oxidised E.coli DsbA in complex with allene
To Be Published
|
|
