3QNE
 
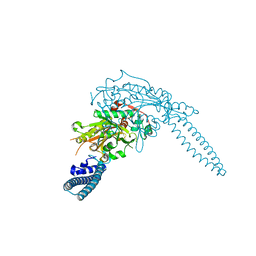 | | Candida albicans seryl-tRNA synthetase | | Descriptor: | Seryl-tRNA synthetase, cytoplasmic | | Authors: | Rocha, R, Santos, M.A, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2011-02-08 | | Release date: | 2011-08-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Unveiling the structural basis for translational ambiguity tolerance in a human fungal pathogen.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3QO5
 
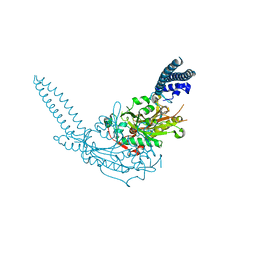 | | Crystal Structure of the seryl-tRNA synthetase from Candida albicans | | Descriptor: | Seryl-tRNA synthetase, cytoplasmic | | Authors: | Rocha, R, Santos, M.A, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2011-02-09 | | Release date: | 2011-08-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Unveiling the structural basis for translational ambiguity tolerance in a human fungal pathogen.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3QO7
 | | Crystal structure of the seryl-tRNA synthetase from Candida albicans | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Seryl-tRNA synthetase, cytoplasmic | | Authors: | Rocha, R, Santos, M.A, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2011-02-09 | | Release date: | 2011-08-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Unveiling the structural basis for translational ambiguity tolerance in a human fungal pathogen.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3QO8
 
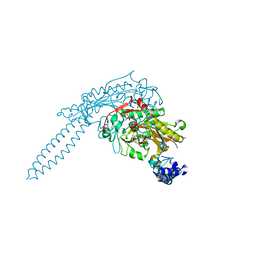 | | Crystal Structure of seryl-tRNA synthetase from Candida albicans | | Descriptor: | 5'-O-(N-(L-SERYL)-SULFAMOYL)ADENOSINE, MAGNESIUM ION, Seryl-tRNA synthetase, ... | | Authors: | Rocha, R, Santos, M.A, Pereira, P.J.B, Macedo-Ribeiro, S. | | Deposit date: | 2011-02-09 | | Release date: | 2011-08-03 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Unveiling the structural basis for translational ambiguity tolerance in a human fungal pathogen.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6I8V
 
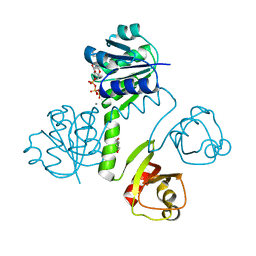 | | KtrC with ATP bound | | Descriptor: | 1,4-BUTANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Rocha, R, Morais-Cabral, J.H. | | Deposit date: | 2018-11-21 | | Release date: | 2019-10-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.991 Å) | | Cite: | Characterization of the molecular properties of KtrC, a second RCK domain that regulates a Ktr channel in Bacillus subtilis.
J.Struct.Biol., 205, 2019
|
|
8POO
 
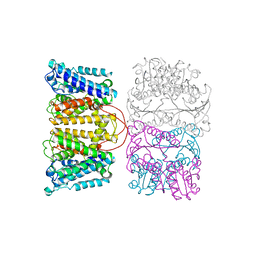 | |
7L7U
 
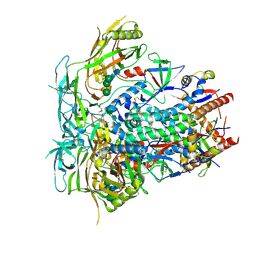 | | BG505 SOSIP reconstructed from a designed nanoparticle, BG505 SOSIP-T33-31 (Component B) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BG505 SOSIP.v5.2(7S) - gp120, ... | | Authors: | Antanasijevic, A, Froes Rocha, R, Sewall, L.M, Ward, A.B. | | Deposit date: | 2020-12-30 | | Release date: | 2021-08-04 | | Last modified: | 2021-08-25 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Polyclonal antibody responses to HIV Env immunogens resolved using cryoEM.
Nat Commun, 12, 2021
|
|
5BUT
 
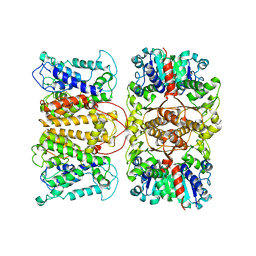 | |
8TGO
 
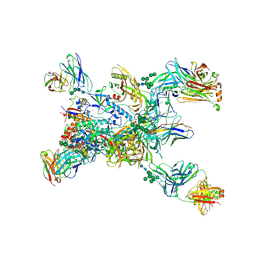 | | Crystal structure of the BG505 triple tandem trimer gp140 HIV-1 Env in complex with PGT124 and 35O22 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 35O22 scFv, ... | | Authors: | Xian, Y, Yuan, M, Wilson, I.A. | | Deposit date: | 2023-07-12 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (5.75 Å) | | Cite: | Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines.
Npj Vaccines, 9, 2024
|
|
2VKR
 
 | | 3Fe-4S, 4Fe-4S plus Zn Acidianus ambivalens ferredoxin | | Descriptor: | FE3-S4 CLUSTER, IRON/SULFUR CLUSTER, ZINC ION, ... | | Authors: | Frazao, C, Aragao, D, Coelho, R, Leal, S.S, Gomes, C.M, Teixeira, M, Carrondo, M.A. | | Deposit date: | 2007-12-23 | | Release date: | 2008-03-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Crystallographic analysis of the intact metal centres [3Fe-4S](1+/0) and [4Fe-4S](2+/1+) in a Zn(2+) -containing ferredoxin.
FEBS Lett., 582, 2008
|
|
6TUR
 
 | | human XPG, Apo1 form | | Descriptor: | DNA repair protein complementing XP-G cells,DNA repair protein complementing XP-G cells | | Authors: | Ruiz, F.M, Fernandez-Tornero, C. | | Deposit date: | 2020-01-08 | | Release date: | 2020-09-16 | | Last modified: | 2020-10-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The crystal structure of human XPG, the xeroderma pigmentosum group G endonuclease, provides insight into nucleotide excision DNA repair.
Nucleic Acids Res., 48, 2020
|
|
6TUX
 
 | | human XPG-DNA, Complex 2 | | Descriptor: | DNA (5'-D(P*AP*AP*CP*TP*CP*TP*GP*C)-3'), DNA (5'-D(P*GP*CP*AP*GP*AP*GP*TP*T)-3'), DNA repair protein complementing XP-G cells,DNA repair protein complementing XP-G cells | | Authors: | Ruiz, F.M, Fernandez-Tornero, C. | | Deposit date: | 2020-01-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The crystal structure of human XPG, the xeroderma pigmentosum group G endonuclease, provides insight into nucleotide excision DNA repair.
Nucleic Acids Res., 48, 2020
|
|
6TUW
 
 | | human XPG-DNA, Complex 1 | | Descriptor: | DNA (5'-D(P*GP*AP*AP*CP*TP*CP*TP*G)-3'), DNA (5'-D(P*TP*GP*CP*AP*GP*AP*GP*TP*TP*C)-3'), DNA repair protein complementing XP-G cells,DNA repair protein complementing XP-G cells | | Authors: | Ruiz, F.M, Fernandez-Tornero, C. | | Deposit date: | 2020-01-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | The crystal structure of human XPG, the xeroderma pigmentosum group G endonuclease, provides insight into nucleotide excision DNA repair.
Nucleic Acids Res., 48, 2020
|
|
6TUS
 
 | | human XPG, Apo2 form | | Descriptor: | DNA repair protein complementing XP-G cells,DNA repair protein complementing XP-G cells, GLYCEROL, SULFATE ION | | Authors: | Ruiz, F.M, Fernandez-Tornero, C. | | Deposit date: | 2020-01-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of human XPG, the xeroderma pigmentosum group G endonuclease, provides insight into nucleotide excision DNA repair.
Nucleic Acids Res., 48, 2020
|
|
3JUT
 
 | | Acidic Fibroblast Growth Factor (FGF-1) complexed with gentisic acid | | Descriptor: | 2,5-dihydroxybenzoic acid, Heparin-binding growth factor 1 | | Authors: | Fernandez, I.S, Gimenez-Gallego, G, Romero, A. | | Deposit date: | 2009-09-15 | | Release date: | 2010-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Gentisic acid, a compound associated with plant defense and a metabolite of aspirin, heads a new class of in vivo fibroblast growth factor inhibitors.
J.Biol.Chem., 285, 2010
|
|
3K1X
 
 | | Acidic Fibroblast Growth Factor (FGF-1) complexed with dobesilate | | Descriptor: | 2,5-dihydroxybenzenesulfonic acid, Heparin-binding growth factor 1 | | Authors: | Romero, A, Fernandez, I.S, Gimenez-Gallego, G. | | Deposit date: | 2009-09-29 | | Release date: | 2010-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Gentisic acid, a compound associated with plant defense and a metabolite of aspirin, heads a new class of in vivo fibroblast growth factor inhibitors.
J.Biol.Chem., 285, 2010
|
|
