4N3D
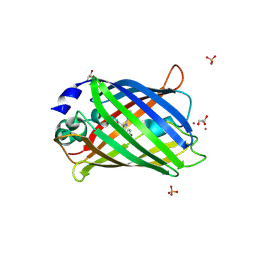 
 | | Crystal structure of the dimeric variant EGFP-K162Q in P61 space group | | Descriptor: | GLYCEROL, Green fluorescent protein, PHOSPHATE ION, ... | | Authors: | Pletneva, N.V, Pletnev, V.Z, Pletnev, S.V. | | Deposit date: | 2013-10-07 | | Release date: | 2014-08-27 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Three dimensional structure of the dimeric gene-engineered variant of green fluorescent protein egfp-K162Q in P61 crystal space group
Rus.J.Bioorg.Chem., 40, 2014
|
|
3LVD
 
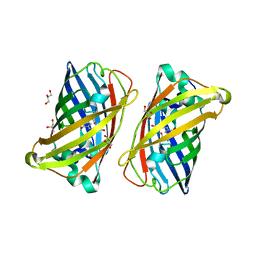 | |
3LVC
 
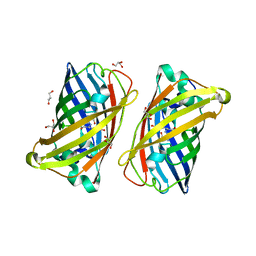 | |
3LVA
 
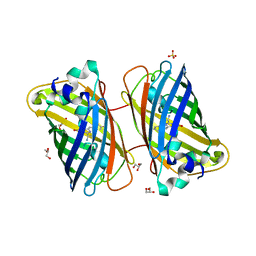 | |
4ZFS
 
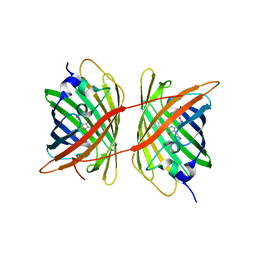 | |
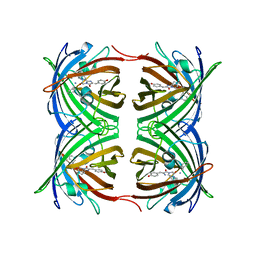
2OGR
 | |
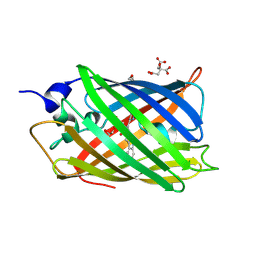
4ZBL
 | |
6ZQO
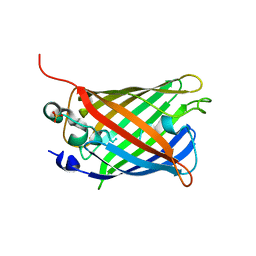 
 | | EYFP mutant - F165G | | Descriptor: | G protein/GFP fusion protein, SULFATE ION | | Authors: | Pletnev, V.Z, Pletnev, S.V, Pletneva, N.V. | | Deposit date: | 2020-07-10 | | Release date: | 2021-06-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Amino acid residue at the 165th position tunes EYFP chromophore maturation. A structure-based design.
Comput Struct Biotechnol J, 19, 2021
|
|
5EXB
 | |
5EXC
 
 | |
3PJB
 
 | |
3PIB
 | |
3PJ7
 | |
3PJ5
 
 | |
4EDS
 | |
4EDO
 | |
7BHG
 | |
3GB3
 
 | |
3GL4
 
 | |
4HVF
 
 | |
4JEO
 
 | |
4JGE
 
 | |
4JF9
 
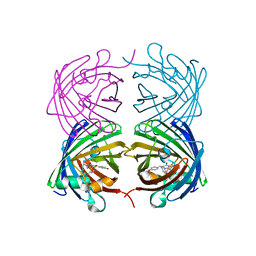 | |
4RTC
 | | Crystal structure of the green fluorescent variant, nowGFP, of the cyan Cerulean at pH 9.0 | | Descriptor: | GLYCEROL, nowGFP | | Authors: | Pletnev, V.Z, Pletneva, N.V, Pletnev, S.V. | | Deposit date: | 2014-11-14 | | Release date: | 2015-09-02 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structure of the green fluorescent protein NowGFP with an anionic tryptophan-based chromophore.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4RYS
 | |
