1CTN
 
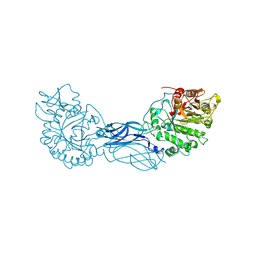 | | CRYSTAL STRUCTURE OF A BACTERIAL CHITINASE AT 2.3 ANGSTROMS RESOLUTION | | Descriptor: | CHITINASE A | | Authors: | Perrakis, A, Tews, I, Dauter, Z, Wilson, K.S, Vorgias, C.E. | | Deposit date: | 1994-10-10 | | Release date: | 1995-02-07 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a bacterial chitinase at 2.3 A resolution.
Structure, 2, 1994
|
|
6QCG
 
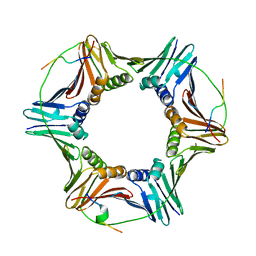 | | PCNA complex with Cdt1 N-terminal PIP-box peptide | | Descriptor: | DNA replication factor Cdt1, Proliferating cell nuclear antigen | | Authors: | Perrakis, A, von Castelmur, E. | | Deposit date: | 2018-12-28 | | Release date: | 2019-01-23 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Direct binding of Cdt2 to PCNA is important for targeting the CRL4Cdt2E3 ligase activity to Cdt1.
Life Sci Alliance, 1, 2018
|
|
8ARF
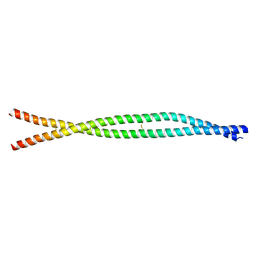 
 | |
7P0K
 
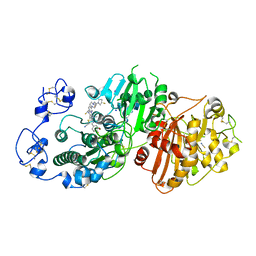 | | Crystal structure of Autotaxin (ENPP2) with 18F-labeled positron emission tomography ligand | | Descriptor: | 2-[[2-ethyl-6-[4-[2-[(3~{R})-3-fluoranylpyrrolidin-1-yl]-2-oxidanylidene-ethyl]piperazin-1-yl]imidazo[1,2-a]pyridin-3-yl]-methyl-amino]-4-(4-fluorophenyl)-2,3-dihydro-1,3-thiazole-5-carbonitrile, 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Salgado-Polo, F, Shao, T, Xiao, Z, Van, R, Chen, J, Rong, J, Haider, A, Shao, Y, Josephson, L, Perrakis, A, Liang, S.H. | | Deposit date: | 2021-06-29 | | Release date: | 2022-07-13 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Imaging Autotaxin In Vivo with 18 F-Labeled Positron Emission Tomography Ligands
J Med Chem, 64, 2021
|
|
9HJT
 
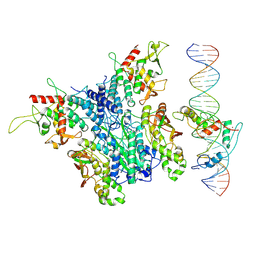 | | Structure of Zincore (SEPHS1:QRICH1) binding to ZFP91 on DNA | | Descriptor: | DNA (31-MER), E3 ubiquitin-protein ligase ZFP91, Selenide, ... | | Authors: | Borza, R, Perrakis, A. | | Deposit date: | 2024-12-02 | | Release date: | 2025-07-09 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Zincore, an atypical coregulator, binds zinc finger transcription factors to control gene expression.
Science, 389, 2025
|
|
9HJU
 
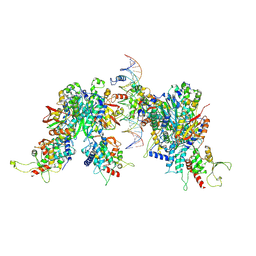 | | Structure of 2x Zincore (SEPHS1:QRICH1) binding to ZFP91 on DNA | | Descriptor: | DNA (31-MER), E3 ubiquitin-protein ligase ZFP91, Selenide, ... | | Authors: | Borza, R, Perrakis, A. | | Deposit date: | 2024-12-02 | | Release date: | 2025-07-09 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | Zincore, an atypical coregulator, binds zinc finger transcription factors to control gene expression.
Science, 389, 2025
|
|
6NVQ
 
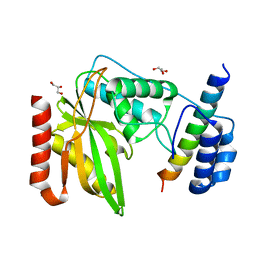 | |
9GSQ
 
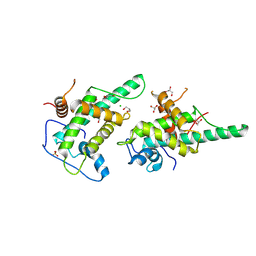 | | DNA binding domain of J-DNA Binding Protein 3 (JBP3) | | Descriptor: | CHLORIDE ION, DNA binding domain of J-DNA binding protein 3, GLYCEROL, ... | | Authors: | de Vries, I, Adamopoulos, A, Joosten, R.P, Perrakis, A. | | Deposit date: | 2024-09-16 | | Release date: | 2024-09-25 | | Last modified: | 2025-01-22 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | JBP1 and JBP3 have conserved structures but different affinity to base-J.
J.Struct.Biol., 217, 2024
|
|
9GSO
 
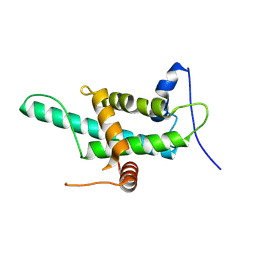 | |
2CKL
 
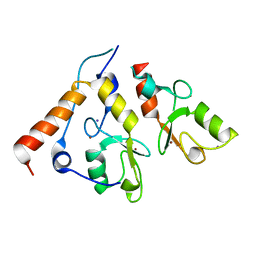 | | Ring1b-Bmi1 E3 catalytic domain structure | | Descriptor: | IODIDE ION, POLYCOMB GROUP RING FINGER PROTEIN 4, UBIQUITIN LIGASE PROTEIN RING2, ... | | Authors: | Buchwald, G, van der Stoop, P, Weichenrieder, O, Perrakis, A, van Lohuizen, M, Sixma, T.K. | | Deposit date: | 2006-04-20 | | Release date: | 2006-05-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and E3-Ligase Activity of the Ring-Ring Complex of Polycomb Proteins Bmi1 and Ring1B.
Embo J., 25, 2006
|
|
3KJY
 
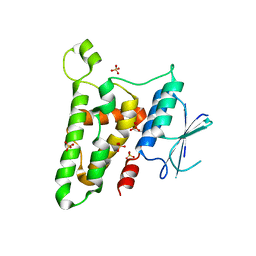 | | Crystal structure of reduced HOMO SAPIENS CLIC3 | | Descriptor: | Chloride intracellular channel protein 3, SULFATE ION | | Authors: | Littler, D.R, Curmi, P.M.G, Breit, S.N, Perrakis, A. | | Deposit date: | 2009-11-04 | | Release date: | 2009-11-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of human CLIC3 at 2 A resolution
Proteins, 78, 2010
|
|
2YLM
 
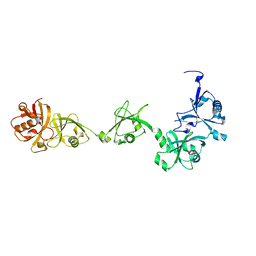 | |
6FPV
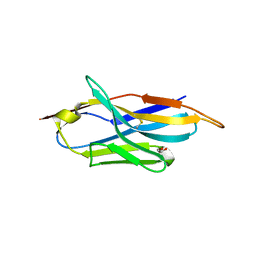 
 | | A llama-derived JBP1-targeting nanobody | | Descriptor: | GLYCEROL, Nanobody | | Authors: | van Beusekom, B, Adamopoulos, A, Heidebrecht, T, Joosten, R.P, Perrakis, A. | | Deposit date: | 2018-02-12 | | Release date: | 2018-10-31 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Characterization and structure determination of a llama-derived nanobody targeting the J-base binding protein 1.
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
5IJS
 
 | | Crystal structure of autotaxin with orthovanadate bound as a trigonal bipyramidal intermediate analog | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 7alpha-hydroxycholesterol, CALCIUM ION, ... | | Authors: | Hausmann, J, Joosten, R.P, Perrakis, A. | | Deposit date: | 2016-03-02 | | Release date: | 2016-06-15 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural snapshots of the catalytic cycle of the phosphodiesterase Autotaxin.
J.Struct.Biol., 195, 2016
|
|
5IJQ
 
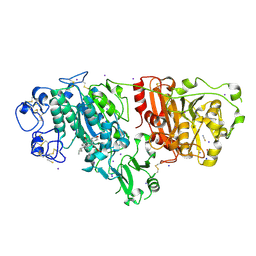 | | Crystal structure of autotaxin (ENPP2) re-refined | | Descriptor: | 7alpha-hydroxycholesterol, CALCIUM ION, Ectonucleotide pyrophosphatase/phosphodiesterase family member 2, ... | | Authors: | Hausmann, J, Joosten, R.P, Perrakis, A. | | Deposit date: | 2016-03-02 | | Release date: | 2016-06-15 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural snapshots of the catalytic cycle of the phosphodiesterase Autotaxin.
J.Struct.Biol., 195, 2016
|
|
9FTN
 
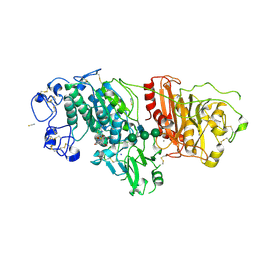 | | Crystal Structure of Autotaxin (ENPP2) with Type VI Inhibitor, a Novel Class of Inhibitors with Three-Point Lock Binding Mode | | Descriptor: | 3-(3-((4-(4-fluorophenyl)thiazol-2-yl)(methyl)amino)-6-(1-(methylsulfonyl)piperidin-4-yl)imidazo[1,2-b]pyridazin-2-yl)-N-(2-oxo-2,3-dihydrobenzo[d]oxazol-6-yl)propenamide, CALCIUM ION, IODIDE ION, ... | | Authors: | Borza, R, Joosten, R.P, Perrakis, A. | | Deposit date: | 2024-06-25 | | Release date: | 2025-04-23 | | Last modified: | 2025-04-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Design, Synthesis, and Biological Implications of Autotaxin inhibitors with a Three-Point lock binding mode.
Bioorg.Med.Chem., 124, 2025
|
|
9FXU
 
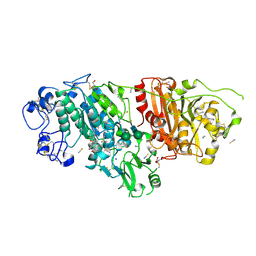 | | Crystal Structure of Autotaxin (ENPP2) with Type VI Inhibitor, a Novel Class of Inhibitors with Three-Point Lock Binding Mode | | Descriptor: | CALCIUM ION, GLYCEROL, IODIDE ION, ... | | Authors: | Borza, R, Joosten, R.P, Perrakis, A. | | Deposit date: | 2024-07-02 | | Release date: | 2025-04-23 | | Last modified: | 2025-04-30 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Design, Synthesis, and Biological Implications of Autotaxin inhibitors with a Three-Point lock binding mode.
Bioorg.Med.Chem., 124, 2025
|
|
9FXW
 
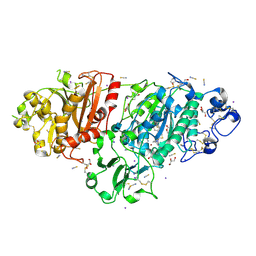 | | Crystal Structure of Autotaxin (ENPP2) with Type VI Inhibitor, a Novel Class of Inhibitors with Three-Point Lock Binding Mode | | Descriptor: | CALCIUM ION, GLYCEROL, IODIDE ION, ... | | Authors: | Borza, R, Joosten, R.P, Perrakis, A. | | Deposit date: | 2024-07-02 | | Release date: | 2025-04-23 | | Last modified: | 2025-04-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design, Synthesis, and Biological Implications of Autotaxin inhibitors with a Three-Point lock binding mode.
Bioorg.Med.Chem., 124, 2025
|
|
9FXY
 
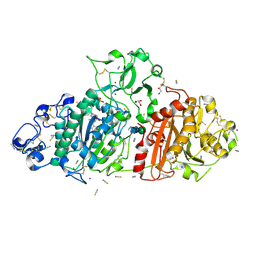 | | Crystal Structure of Autotaxin (ENPP2) with Type IV Inhibitor | | Descriptor: | 3-(3-((4-(4-fluorophenyl)thiazol-2-yl)(methyl)amino)-6-(1-(methylsulfonyl)piperidin-4-yl)imidazo[1,2-b]pyridazin-2-yl)propanenitrile, CALCIUM ION, GLYCEROL, ... | | Authors: | Borza, R, Joosten, R.P, Perrakis, A. | | Deposit date: | 2024-07-02 | | Release date: | 2025-04-23 | | Last modified: | 2025-04-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design, Synthesis, and Biological Implications of Autotaxin inhibitors with a Three-Point lock binding mode.
Bioorg.Med.Chem., 124, 2025
|
|
6GVJ
 
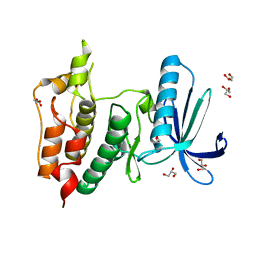 | | Human Mps1 kinase domain with ordered activation loop | | Descriptor: | CHLORIDE ION, Dual specificity protein kinase TTK, GLYCEROL | | Authors: | Roorda, J.C, Hiruma, Y, Joosten, R.P, Perrakis, A. | | Deposit date: | 2018-06-21 | | Release date: | 2019-01-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | A crystal structure of the human protein kinase Mps1 reveals an ordered conformation of the activation loop.
Proteins, 87, 2019
|
|
5M0M
 
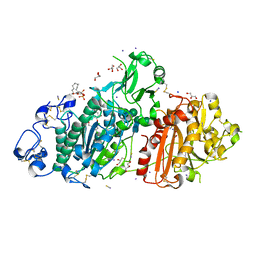 | | Structure-based evolution of a hybrid steroid series of Autotaxin inhibitors | | Descriptor: | (2R)-2-hydroxy-3-(phosphonooxy)propyl (9E)-octadec-9-enoate, CALCIUM ION, Ectonucleotide pyrophosphatase/phosphodiesterase family member 2, ... | | Authors: | Keune, W.-J, Heidebrecht, T, Perrakis, A. | | Deposit date: | 2016-10-05 | | Release date: | 2017-08-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Rational Design of Autotaxin Inhibitors by Structural Evolution of Endogenous Modulators.
J. Med. Chem., 60, 2017
|
|
5M0S
 
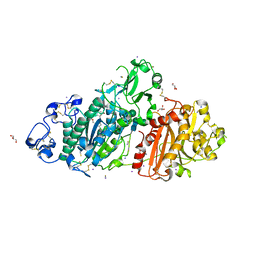 | | Structure-based evolution of a hybrid steroid series of Autotaxin inhibitors | | Descriptor: | CALCIUM ION, Ectonucleotide pyrophosphatase/phosphodiesterase family member 2, GLYCEROL, ... | | Authors: | Keune, W.-J, Heidebrecht, T, Perrakis, A. | | Deposit date: | 2016-10-05 | | Release date: | 2017-08-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Rational Design of Autotaxin Inhibitors by Structural Evolution of Endogenous Modulators.
J. Med. Chem., 60, 2017
|
|
5LJJ
 
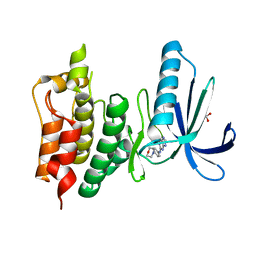 | | Crystal structure of human Mps1 (TTK) in complex with Reversine | | Descriptor: | 1,2-ETHANEDIOL, Dual specificity protein kinase TTK, N~6~-cyclohexyl-N~2~-(4-morpholin-4-ylphenyl)-9H-purine-2,6-diamine | | Authors: | Hiruma, Y, Joosten, R.P, Perrakis, A. | | Deposit date: | 2016-07-18 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis of reversine selectivity in inhibiting Mps1 more potently than aurora B kinase.
Proteins, 84, 2016
|
|
5M0D
 
 | | Structure-based evolution of a hybrid steroid series of Autotaxin inhibitors | | Descriptor: | CALCIUM ION, Ectonucleotide pyrophosphatase/phosphodiesterase family member 2,Ectonucleotide pyrophosphatase/phosphodiesterase family member 2, GLYCEROL, ... | | Authors: | Keune, W.-J, Heidebrecht, T, Perrakis, A. | | Deposit date: | 2016-10-04 | | Release date: | 2017-08-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Rational Design of Autotaxin Inhibitors by Structural Evolution of Endogenous Modulators.
J. Med. Chem., 60, 2017
|
|
5M0E
 
 | | Structure-based evolution of a hybrid steroid series of Autotaxin inhibitors | | Descriptor: | 7alpha-hydroxycholesterol, CALCIUM ION, Ectonucleotide pyrophosphatase/phosphodiesterase family member 2, ... | | Authors: | Keune, W.-J, Heidebrecht, T, Perrakis, A. | | Deposit date: | 2016-10-04 | | Release date: | 2017-08-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Rational Design of Autotaxin Inhibitors by Structural Evolution of Endogenous Modulators.
J. Med. Chem., 60, 2017
|
|
