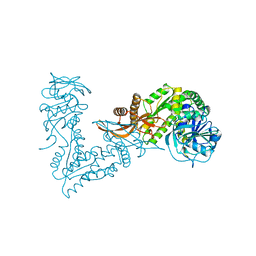
7MO6
 
 | |
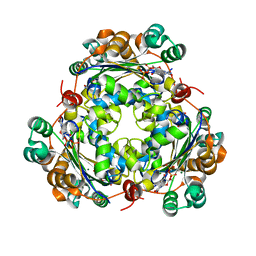
6XPT
 
 | |
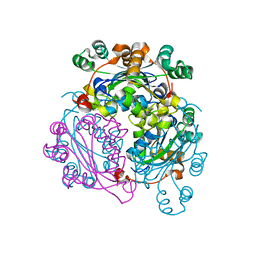
6XP4
 
 | |
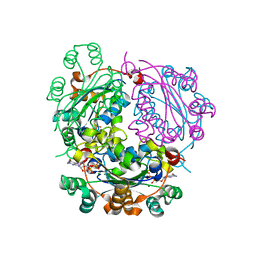
6XPS
 
 | |
6XPV
 
 | |
6XPW
 | |
6XP7
 
 | |
6XPU
 
 | |
7RHV
 
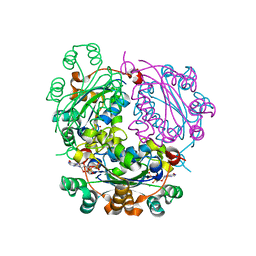
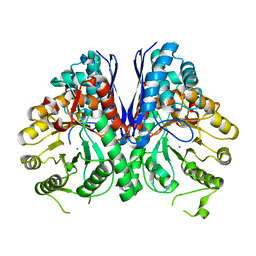 | | Aspergillus fumigatus Enolase | | Descriptor: | Enolase, MAGNESIUM ION | | Authors: | Nguyen, S, Bruning, J.B. | | Deposit date: | 2021-07-19 | | Release date: | 2022-03-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A structural model of the human plasminogen and Aspergillus fumigatus enolase complex.
Proteins, 90, 2022
|
|
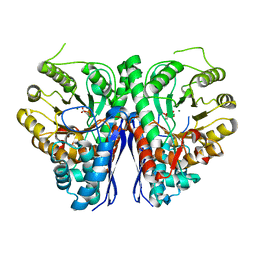
7RI0
 
 | |
7RHW
 
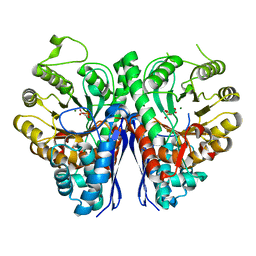 | |
7RK5
 
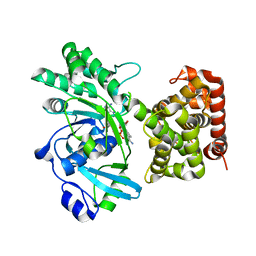 | |
7RK4
 
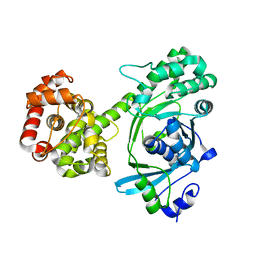 | |
3VD0
 
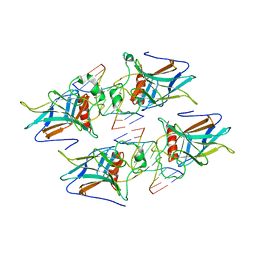 | | structure of p73 DNA binding domain tetramer modulates p73 transactivation | | Descriptor: | DNA (5'-D(*CP*AP*GP*GP*CP*AP*TP*GP*CP*CP*TP*G)-3'), Tumor protein p73, ZINC ION | | Authors: | Ethayathulla, A.S, Tse, P.W, Nguyen, S, Viadiu, H. | | Deposit date: | 2012-01-04 | | Release date: | 2012-04-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structure of p73 DNA-binding domain tetramer modulates p73 transactivation.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3VD1
 
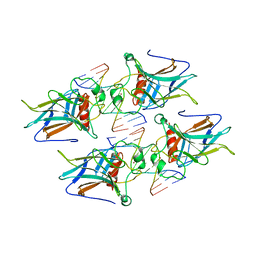 | | structure of p73 DNA binding domain tetramer modulates p73 transactivation | | Descriptor: | DNA (5'-D(*CP*GP*GP*GP*CP*AP*TP*GP*CP*CP*CP*G)-3'), Tumor protein p73, ZINC ION | | Authors: | Ethayathulla, A.S, Tse, P.W, Nguyen, S, Viadiu, H. | | Deposit date: | 2012-01-04 | | Release date: | 2012-04-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structure of p73 DNA-binding domain tetramer modulates p73 transactivation.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3VD2
 
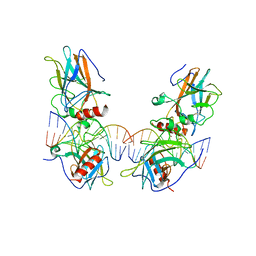 | | structure of p73 DNA binding domain tetramer modulates p73 transactivation | | Descriptor: | DNA (5'-D(*AP*TP*GP*GP*AP*CP*AP*TP*GP*TP*CP*CP*AP*T)-3'), Tumor protein p73, ZINC ION | | Authors: | Ethayathulla, A.S, Tse, P.W, Nguyen, S, Viadiu, H. | | Deposit date: | 2012-01-04 | | Release date: | 2012-04-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Structure of p73 DNA-binding domain tetramer modulates p73 transactivation.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
7KH5
 
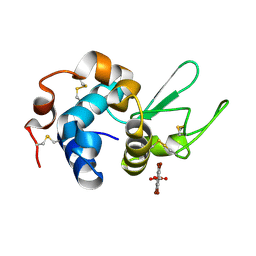 | | Hen Egg White Lysozyme in complex with tetrabromoterephthalic acid | | Descriptor: | 2,3,5,6-tetrabromobenzene-1,4-dicarboxylic acid, Lysozyme C | | Authors: | Truong, J, Nguyen, S. | | Deposit date: | 2020-10-20 | | Release date: | 2021-04-28 | | Last modified: | 2021-05-19 | | Method: | X-RAY DIFFRACTION (1.295 Å) | | Cite: | Simplified heavy-atom derivatization of protein structures via co-crystallization with the MAD tetragon tetrabromoterephthalic acid.
Acta Crystallogr.,Sect.F, 77, 2021
|
|
6CWG
 
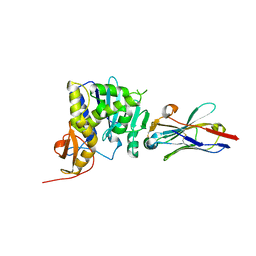 | | Ricin catalytic subunit bound go A9 VHH antibody | | Descriptor: | CHLORIDE ION, Ricin, VHH antibody | | Authors: | Rudolph, M.J, Mantis, N. | | Deposit date: | 2018-03-30 | | Release date: | 2019-01-16 | | Last modified: | 2019-12-18 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Contribution of an unusual CDR2 element of a single domain antibody in ricin toxin binding affinity and neutralizing activity.
Protein Eng. Des. Sel., 31, 2018
|
|
6OJY
 
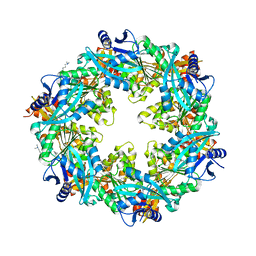 | |
6OJX
 
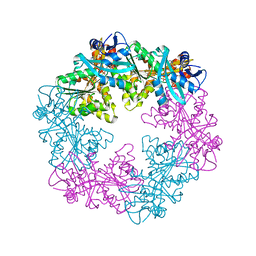 | |
6OKV
 
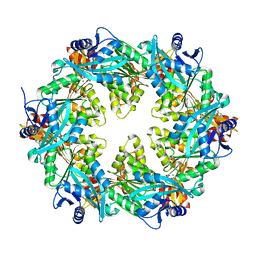 | |
6OLK
 | |
6OLM
 
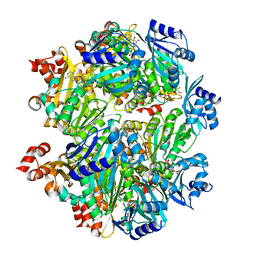 | |
6OJZ
 
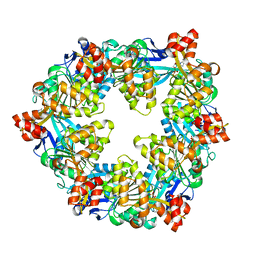 | |
6OLL
 
 | |
