8GMW
 
 | | Crystal structure of lysozyme | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION | | Authors: | Nam, K.H. | | Deposit date: | 2022-08-22 | | Release date: | 2022-09-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal structure of lysozyme
To Be Published
|
|
8GMV
 
 | | Crystal structure of lysozyme | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION | | Authors: | Nam, K.H. | | Deposit date: | 2022-08-22 | | Release date: | 2022-09-21 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of lysozyme
To Be Published
|
|
9K73
 
 | | Crystal structure of TsaBgl | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SODIUM ION, beta-glucosidase | | Authors: | Nam, K.H. | | Deposit date: | 2024-10-23 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of TsaBgl
To Be Published
|
|
9K9Z
 
 | |
9KA0
 
 | |
9K72
 
 | | Crystal structure of TsaBgl using merged datasets | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SODIUM ION, beta-glucosidase | | Authors: | Nam, K.H. | | Deposit date: | 2024-10-23 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal structure of TsaBgl using merged datasets
To Be Published
|
|
9L9R
 
 | |
9LMK
 
 | |
9LML
 
 | |
9UGI
 
 | |
3L1J
 
 | | Crystal structure of EstE5, was soaked by ZnSO4 | | Descriptor: | Esterase/lipase | | Authors: | Nam, K.H, Hwang, K.Y. | | Deposit date: | 2009-12-11 | | Release date: | 2010-01-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into the noninvasive inhibition of HSL-homolog EstE5 by organic solvents and metal ions
To be Published
|
|
3L1H
 
 | | Crystal structure of EstE5, was soaked by FeCl3 | | Descriptor: | Esterase/lipase | | Authors: | Nam, K.H, Hwang, K.Y. | | Deposit date: | 2009-12-11 | | Release date: | 2010-01-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural insights into the noninvasive inhibition of HSL-homolog EstE5 by organic solvents and metal ions
To be Published
|
|
3L1I
 
 | | Crystal structure of EstE5, was soaked by CuSO4 | | Descriptor: | Esterase/lipase | | Authors: | Nam, K.H, Hwang, K.Y. | | Deposit date: | 2009-12-11 | | Release date: | 2010-01-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural insights into the noninvasive inhibition of HSL-homolog EstE5 by organic solvents and metal ions
To be Published
|
|
8WXO
 
 | |
8WXN
 
 | |
8WXM
 
 | |
8X1D
 
 | |
8WXP
 
 | |
8HVF
 
 | | Crystal structure of Thaumatin (100 ms) | | Descriptor: | 1,2-ETHANEDIOL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Nam, K.H. | | Deposit date: | 2022-12-26 | | Release date: | 2023-11-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of Thaumatin (100 ms)
To Be Published
|
|
8HVE
 
 | | Crystal structure of Thaumatin (1 s) | | Descriptor: | 1,2-ETHANEDIOL, L(+)-TARTARIC ACID, Thaumatin I | | Authors: | Nam, K.H. | | Deposit date: | 2022-12-26 | | Release date: | 2023-11-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of Thaumatin (1 s)
To Be Published
|
|
7BVL
 
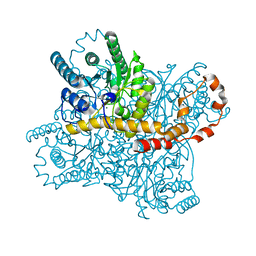 | |
7BVO
 
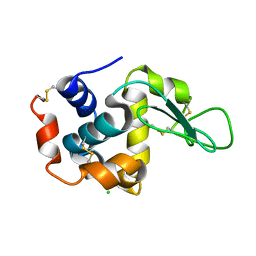 | |
7BVM
 
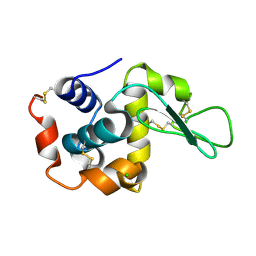 | |
7BVN
 
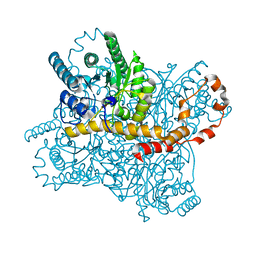 | |
7CJZ
 
 | |
