8OQW
 
 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK | | Descriptor: | ATPase, GLYCEROL, SULFATE ION | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OQK
 
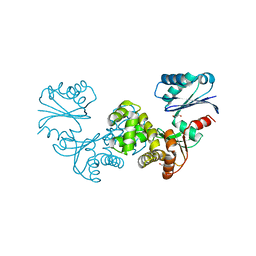 | | Crystal structure of Tannerella forsythia sugar kinase K1058 | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, N-acetylglucosamine kinase | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OQX
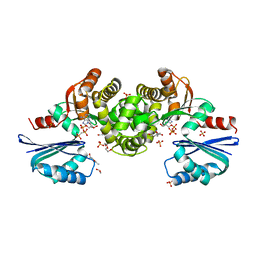 
 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK with a phosphate analogue | | Descriptor: | 1,2-ETHANEDIOL, ATPase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Gogler, K, Fink, P, Stasiak, A.C, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-12 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OW9
 
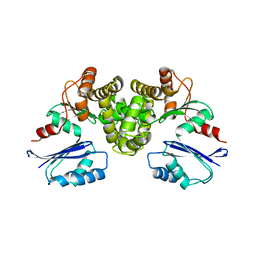 | | Crystal structure of Tannerella forsythia MurNAc kinase MurK in complex with N-acetylmuramic acid (MurNAc) | | Descriptor: | N-acetyl-beta-muramic acid, Putative novel MurNAc kinase | | Authors: | Stasiak, A.C, Gogler, K, Fink, P, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-08-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
8OW7
 
 | | Crystal structure of Tannerella forsythia sugar kinase K1058 in complex with N-acetylmuramic acid (MurNAc) | | Descriptor: | N-acetyl-beta-muramic acid, N-acetylglucosamine kinase, SULFATE ION | | Authors: | Stasiak, A.C, Gogler, K, Fink, P, Stehle, T, Zocher, G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-08-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | N-acetylmuramic acid recognition by MurK kinase from the MurNAc auxotrophic oral pathogen Tannerella forsythia.
J.Biol.Chem., 299, 2023
|
|
4Y7T
 
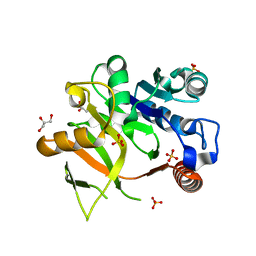 | | Structural analysis of MurU | | Descriptor: | GLYCEROL, Nucleotidyl transferase, SULFATE ION | | Authors: | Renner-Schneck, M.G, Stehle, T. | | Deposit date: | 2015-02-16 | | Release date: | 2015-03-18 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of the N-Acetylmuramic Acid alpha-1-Phosphate (MurNAc-alpha 1-P) Uridylyltransferase MurU, a Minimal Sugar Nucleotidyltransferase and Potential Drug Target Enzyme in Gram-negative Pathogens.
J.Biol.Chem., 290, 2015
|
|
4Y7U
 
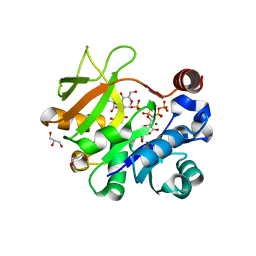 | | Structural analysis of MurU | | Descriptor: | 2-acetamido-3-O-[(1R)-1-carboxyethyl]-2-deoxy-1-O-phosphono-alpha-D-glucopyranose, 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phosphoryl]uridine, GLYCEROL, ... | | Authors: | Renner-Schneck, M.G, Stehle, T. | | Deposit date: | 2015-02-16 | | Release date: | 2015-03-18 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of the N-Acetylmuramic Acid alpha-1-Phosphate (MurNAc-alpha 1-P) Uridylyltransferase MurU, a Minimal Sugar Nucleotidyltransferase and Potential Drug Target Enzyme in Gram-negative Pathogens.
J.Biol.Chem., 290, 2015
|
|
4Y7V
 
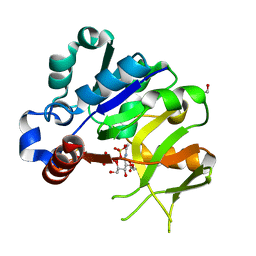 | | Structural analysis of MurU | | Descriptor: | 2-acetamido-3-O-[(1R)-1-carboxyethyl]-2-deoxy-1-O-phosphono-alpha-D-glucopyranose, GLYCEROL, IMIDODIPHOSPHORIC ACID, ... | | Authors: | Renner-Schneck, M.G, Stehle, T. | | Deposit date: | 2015-02-16 | | Release date: | 2015-03-18 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of the N-Acetylmuramic Acid alpha-1-Phosphate (MurNAc-alpha 1-P) Uridylyltransferase MurU, a Minimal Sugar Nucleotidyltransferase and Potential Drug Target Enzyme in Gram-negative Pathogens.
J.Biol.Chem., 290, 2015
|
|
4WZJ
 
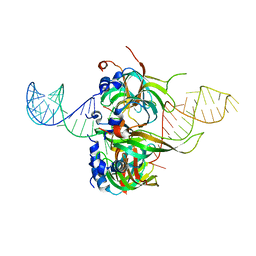 | | Spliceosomal U4 snRNP core domain | | Descriptor: | Small nuclear ribonucleoprotein E, Small nuclear ribonucleoprotein F, Small nuclear ribonucleoprotein G, ... | | Authors: | Leung, A.K.W, Nagai, K, Li, J. | | Deposit date: | 2014-11-19 | | Release date: | 2015-01-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Structure of the spliceosomal U4 snRNP core domain and its implication for snRNP biogenesis.
Nature, 473, 2011
|
|
3LK6
 
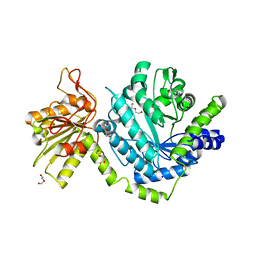 | |
3NVD
 
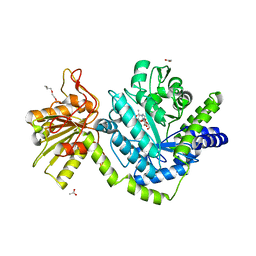 | | Structure of YBBD in complex with pugnac | | Descriptor: | ACETATE ION, O-(2-ACETAMIDO-2-DEOXY D-GLUCOPYRANOSYLIDENE) AMINO-N-PHENYLCARBAMATE, SODIUM ION, ... | | Authors: | Diederichs, K. | | Deposit date: | 2010-07-08 | | Release date: | 2010-08-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.836 Å) | | Cite: | Structural and kinetic analysis of Bacillus subtilis N-acetylglucosaminidase reveals a unique Asp-His dyad mechanism
J.Biol.Chem., 285, 2010
|
|
7QD7
 
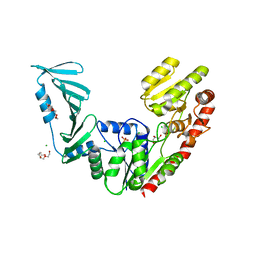 | | TarM(Se)_G117R | | Descriptor: | CHLORIDE ION, GLYCEROL, PENTAETHYLENE GLYCOL, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-11-26 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
7QNT
 
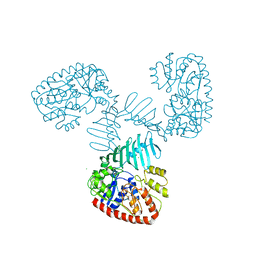 | | TarM(Se) native | | Descriptor: | 1,2-ETHANEDIOL, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-12-22 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.21 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
7QH9
 
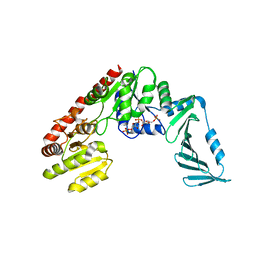 | | TarM(Se)_G117R-4RboP | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, TarM(Se)_G117R-4RboP, ... | | Authors: | Guo, Y, Stehle, T. | | Deposit date: | 2021-12-10 | | Release date: | 2023-05-10 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.689 Å) | | Cite: | Invasive Staphylococcus epidermidis uses a unique processive wall teichoic acid glycosyltransferase to evade immune recognition.
Sci Adv, 9, 2023
|
|
4II9
 
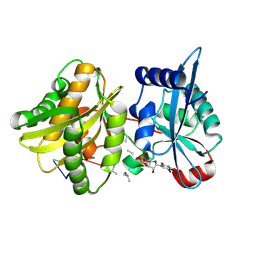 | | Crystal structure of Weissella viridescens FemXVv non-ribosomal amino acid transferase in complex with a peptidyl-RNA conjugate | | Descriptor: | 5-mer peptide, FemX, GLYCEROL, ... | | Authors: | Li de la Sierra-Gallay, I, Fonvielle, M, van Tilbeurgh, H, Arthur, M, Etheve-Quelquejeu, M. | | Deposit date: | 2012-12-20 | | Release date: | 2013-07-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | The Structure of FemXWv in Complex with a Peptidyl-RNA Conjugate: Mechanism of Aminoacyl Transfer from Ala-tRNA(Ala) to Peptidoglycan Precursors
Angew.Chem.Int.Ed.Engl., 52, 2013
|
|
3BMX
 
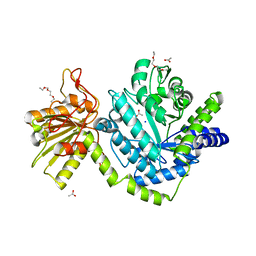 | | Beta-N-hexosaminidase (YbbD) from Bacillus subtilis | | Descriptor: | 1-ETHOXY-2-(2-ETHOXYETHOXY)ETHANE, ACETATE ION, SODIUM ION, ... | | Authors: | Fischer, S. | | Deposit date: | 2007-12-13 | | Release date: | 2008-12-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and kinetic analysis of Bacillus subtilis N-acetylglucosaminidase reveals a unique Asp-His dyad mechanism
J.Biol.Chem., 285, 2010
|
|
3HGB
 
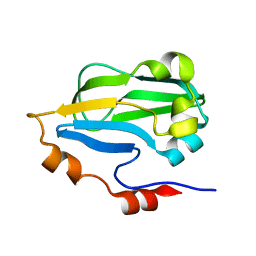 | |
3PL1
 
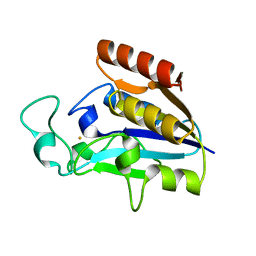 | | Determination of the crystal structure of the pyrazinamidase from M.tuberculosis : a structure-function analysis for prediction resistance to pyrazinamide. | | Descriptor: | FE (II) ION, PYRAZINAMIDASE/NICOTINAMIDASE PNCA (PZase) | | Authors: | Petrella, S, Gelus-Ziental, N, Mayer, C, Sougakoff, W. | | Deposit date: | 2010-11-12 | | Release date: | 2011-01-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the Pyrazinamidase of Mycobacterium tuberculosis: Insights into Natural and Acquired Resistance to Pyrazinamide.
Plos One, 6, 2011
|
|
1LJO
 
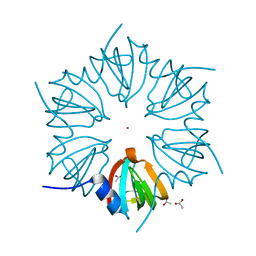 | | CRYSTAL STRUCTURE OF AN SM-LIKE PROTEIN (AF-SM2) FROM ARCHAEOGLOBUS FULGIDUS AT 1.95A RESOLUTION | | Descriptor: | ACETIC ACID, Archaeal Sm-like protein AF-Sm2, CADMIUM ION | | Authors: | Toro, I, Basquin, J, Teo-Dreher, H, Suck, D. | | Deposit date: | 2002-04-22 | | Release date: | 2002-07-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Archaeal Sm proteins form heptameric and hexameric complexes: crystal structures of the Sm1 and Sm2 proteins from the hyperthermophile Archaeoglobus fulgidus.
J.Mol.Biol., 320, 2002
|
|
