5QMB
 
 | |
5QMR
 
 | |
5QN6
 
 | |
5QNO
 
 | |
5QO4
 
 | |
5QMI
 
 | |
5QN0
 
 | |
5QNI
 
 | |
5QNY
 
 | |
5QOF
 
 | |
5QND
 
 | |
5QNT
 
 | |
5QO9
 
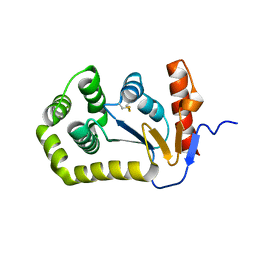 | |
7GPB
 
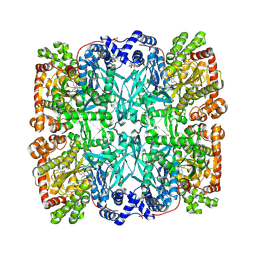 | | STRUCTURAL MECHANISM FOR GLYCOGEN PHOSPHORYLASE CONTROL BY PHOSPHORYLATION AND AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, GLYCOGEN PHOSPHORYLASE B, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Barford, D, Hu, S.-H, Johnson, L.N. | | Deposit date: | 1990-11-13 | | Release date: | 1992-10-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural mechanism for glycogen phosphorylase control by phosphorylation and AMP.
J.Mol.Biol., 218, 1991
|
|
4DWN
 
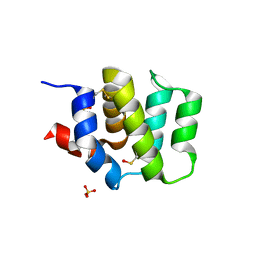 | | Crystal Structure of Human BinCARD CARD | | Descriptor: | Bcl10-interacting CARD protein, SULFATE ION | | Authors: | Chen, K.-E, Kobe, B, Martin, J.L. | | Deposit date: | 2012-02-26 | | Release date: | 2013-02-06 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.581 Å) | | Cite: | The structure of the caspase recruitment domain of BinCARD reveals that all three cysteines can be oxidized.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
7S1D
 
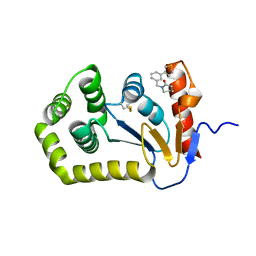 | | Crystal structure of E.coli DsbA in complex with compound MIPS-0001877 (compound 39) | | Descriptor: | 1-[3-(thiophen-3-yl)benzyl]piperidin-2-one, COPPER (II) ION, Thiol:disulfide interchange protein DsbA | | Authors: | Heras, B, Scanlon, M.J, Martin, J.L, Caria, S. | | Deposit date: | 2021-09-02 | | Release date: | 2023-02-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Fluoromethylketone-fragment conjugates designed as covalent modifiers of EcDsbA are atypical substrates
Chemrxiv, 2022
|
|
7S1F
 
 | | Crystal structure of E.coli DsbA in complex with compound MIPS-0001886 (compound 38) | | Descriptor: | 1-[(3-thiophen-3-ylphenyl)methyl]-3~{H}-pyrrol-2-one, COPPER (II) ION, GLYCEROL, ... | | Authors: | Heras, B, Scanlon, M.J, Martin, J.L, Caria, S. | | Deposit date: | 2021-09-02 | | Release date: | 2023-02-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Fluoromethylketone-fragment conjugates designed as covalent modifiers of EcDsbA are atypical substrates
Chemrxiv, 2022
|
|
7S1C
 
 | | Crystal structure of E.coli DsbA in complex with compound MIPS-0001897 (compound 1) | | Descriptor: | COPPER (II) ION, Thiol:disulfide interchange protein DsbA, ~{N}-methyl-1-(3-thiophen-3-ylphenyl)methanamine | | Authors: | Heras, B, Scanlon, M.J, Martin, J.L, Sharma, P. | | Deposit date: | 2021-09-02 | | Release date: | 2023-02-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.949 Å) | | Cite: | Fluoromethylketone-fragment conjugates designed as covalent modifiers of EcDsbA are atypical substrates
Chemrxiv, 2022
|
|
7S1L
 
 | | Crystal structure of E.coli DsbA in complex with compound MIPS-0001896 (compound 72) | | Descriptor: | COPPER (II) ION, Thiol:disulfide interchange protein DsbA, methyl cis-4-({[3-(thiophen-3-yl)benzyl]amino}methyl)cyclohexanecarboxylate | | Authors: | Heras, B, Scanlon, M.J, Martin, J.L, Caria, S. | | Deposit date: | 2021-09-02 | | Release date: | 2023-02-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.623 Å) | | Cite: | Fluoromethylketone-fragment conjugates designed as covalent modifiers of EcDsbA are atypical substrates
Chemrxiv, 2022
|
|
2PRI
 
 | | BINDING OF 2-DEOXY-GLUCOSE-6-PHOSPHATE TO GLYCOGEN PHOSPHORYLASE B | | Descriptor: | 2-deoxy-6-O-phosphono-alpha-D-glucopyranose, GLYCOGEN PHOSPHORYLASE B, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Oikonomakos, N.G, Zographos, S.E, Johnson, L.N, Papageorgiou, A.C, Acharya, K.R. | | Deposit date: | 1998-12-11 | | Release date: | 1998-12-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The binding of 2-deoxy-D-glucose 6-phosphate to glycogen phosphorylase b: kinetic and crystallographic studies.
J.Mol.Biol., 254, 1995
|
|
4DM3
 
 | | Crystal structure of human PNMT in complex adohcy, resorcinol and imidazole | | Descriptor: | IMIDAZOLE, Phenylethanolamine N-methyltransferase, RESORCINOL, ... | | Authors: | Nair, P.C, Malde, A.K, Mark, A.E. | | Deposit date: | 2012-02-06 | | Release date: | 2012-04-25 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2.4001 Å) | | Cite: | Missing fragments: detecting cooperative binding in fragment-based drug design.
ACS Med Chem Lett, 3, 2012
|
|
