8YGD
 
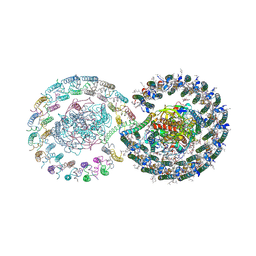 | | Rhodobacter blasticus RC-LH1 dimer | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1-deoxy-D-xylulose-5-phosphate synthase, Antenna pigment protein alpha chain, ... | | Authors: | Liu, L.N, Zhang, Y.Z, Wang, P, Christianson, B.M, Ugurlar, D. | | Deposit date: | 2024-02-26 | | Release date: | 2025-03-05 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Architectures of photosynthetic RC-LH1 supercomplexes from Rhodobacter blasticus.
Sci Adv, 10, 2024
|
|
8YGL
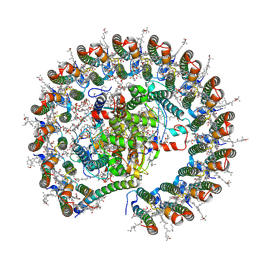 
 | | Rhodobacter blasticus RC-LH1 monomer | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1-deoxy-D-xylulose-5-phosphate synthase, Antenna pigment protein alpha chain, ... | | Authors: | Liu, L.N, Zhang, Y.Z, Wang, P, Christianson, B.M, Ugurlar, D. | | Deposit date: | 2024-02-26 | | Release date: | 2025-03-05 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Architectures of photosynthetic RC-LH1 supercomplexes from Rhodobacter blasticus.
Sci Adv, 10, 2024
|
|
8B12
 
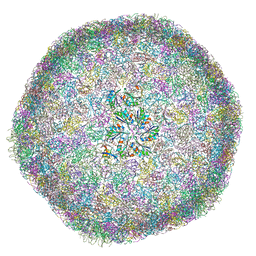 | | cryo-EM structure of carboxysomal mini-shell: icosahedral assembly from CsoS4A/1A and CsoS2 co-expression (T = 9) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Ni, T, Jiang, Q, Liu, L.N, Zhang, P. | | Deposit date: | 2022-09-09 | | Release date: | 2023-08-23 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (1.86 Å) | | Cite: | Intrinsically disordered CsoS2 acts as a general molecular thread for alpha-carboxysome shell assembly.
Nat Commun, 14, 2023
|
|
8B11
 
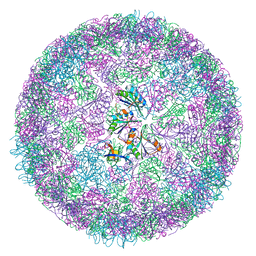 | | cryo-EM structure of carboxysomal mini-shell: icosahedral assembly from CsoS4A/1A and CsoS2 co-expression (T = 4) | | Descriptor: | Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Ni, T, Jiang, Q, Liu, L.N, Zhang, P. | | Deposit date: | 2022-09-08 | | Release date: | 2023-08-23 | | Last modified: | 2023-09-20 | | Method: | ELECTRON MICROSCOPY (2.52 Å) | | Cite: | Intrinsically disordered CsoS2 acts as a general molecular thread for alpha-carboxysome shell assembly.
Nat Commun, 14, 2023
|
|
8B0Y
 
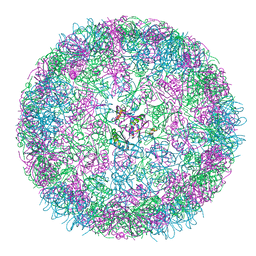 | | cryo-EM structure of carboxysomal mini-shell: icosahedral assembly from CsoS4A/1A co-expression (T = 3) | | Descriptor: | Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Ni, T, Jiang, Q, Liu, L.N, Zhang, P. | | Deposit date: | 2022-09-08 | | Release date: | 2023-08-23 | | Last modified: | 2023-09-20 | | Method: | ELECTRON MICROSCOPY (2.79 Å) | | Cite: | Intrinsically disordered CsoS2 acts as a general molecular thread for alpha-carboxysome shell assembly.
Nat Commun, 14, 2023
|
|
9K3Q
 
 | |
9F0H
 
 | | cryo-EM structure of carboxysomal mini-shell icosahedral assembly from co-expression of CsoS1C, CsoS4A, and CsoS2-C (T = 9) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell protein CsoS1C, Carboxysome shell vertex protein CsoS4A, ... | | Authors: | Wang, P, Marles-Wright, J, Liu, L.N. | | Deposit date: | 2024-04-16 | | Release date: | 2024-12-11 | | Method: | ELECTRON MICROSCOPY (1.8 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
8B64
 
 | | Cryo-EM structure of RC-LH1-PufX photosynthetic core complex from Rba. capsulatus | | Descriptor: | 1,2-Distearoyl-sn-glycerophosphoethanolamine, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, ... | | Authors: | Bracun, L, Yamagata, A, Shirouzu, M, Liu, L.N. | | Deposit date: | 2022-09-26 | | Release date: | 2023-02-01 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (2.589 Å) | | Cite: | Cryo-EM structure of a monomeric RC-LH1-PufX supercomplex with high-carotenoid content from Rhodobacter capsulatus.
Structure, 31, 2023
|
|
8YVC
 
 | | Cryo-EM structure of carboxysomal midi-shell:icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 19) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Wang, P, Li, J.X, Li, T.P, Li, K, Ng, P.C, Wang, S.M, Chriscoli, V, Basle, A, Marles-Wright, J, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-03-28 | | Release date: | 2024-12-11 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
8YVF
 
 | | cryo-EM structure of carboxysomal midi-shell: assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T=9 Q=12) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Wang, P, Li, J.X, Li, T.P, Li, K, Ng, P.C, Wang, S.M, Chriscoli, V, Basle, A, Marles-Wright, J, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-03-28 | | Release date: | 2024-12-11 | | Method: | ELECTRON MICROSCOPY (2.99 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
8YVI
 
 | | Cryo-EM structure of carboxysomal midi-shell: icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 13) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Wang, P, Li, J.X, Li, T.P, Li, K, Ng, P.C, Wang, S.M, Chriscoli, V, Basle, A, Marles-Wright, J, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-03-28 | | Release date: | 2024-12-11 | | Method: | ELECTRON MICROSCOPY (2.93 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
8YXU
 
 | | Crystal structure of CsoS1A/B (modeled with CsoS1A) | | Descriptor: | Major carboxysome shell protein CsoS1A | | Authors: | Wang, P, Li, J.X, Li, T.P, Li, K, Ng, P.C, Wang, S.M, Chriscoli, V, Basle, A, Marles-Wright, J, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-04-03 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
8YVE
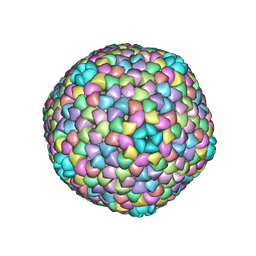 
 | | cryo-EM structure of carboxysomal midi-shell: icosahedral assembly from CsoS4A/4B/1A/1B/1C/1D and CsoS2 C-terminal co-expression (T = 9) | | Descriptor: | Carboxysome assembly protein CsoS2B, Carboxysome shell vertex protein CsoS4A, Major carboxysome shell protein CsoS1A | | Authors: | Wang, P, Li, J.X, Li, T.P, Li, K, Ng, P.C, Wang, S.M, Chriscoli, V, Basle, A, Marles-Wright, J, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-03-28 | | Release date: | 2024-12-11 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Molecular principles of the assembly and construction of a carboxysome shell.
Sci Adv, 10, 2024
|
|
6SMH
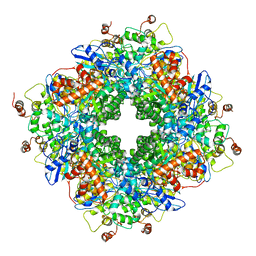 
 | | Cryo-electron microscopy structure of a RbcL-Raf1 supercomplex from Synechococcus elongatus PCC 7942 | | Descriptor: | Ribulose bisphosphate carboxylase large chain, Rubisco accumulation factor 1 (RAF1) peptide | | Authors: | Huang, F, Kong, W.-W, Sun, Y, Chen, T, Dykes, G.F, Jiang, Y.L, Liu, L.N. | | Deposit date: | 2019-08-21 | | Release date: | 2020-07-08 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Rubisco accumulation factor 1 (Raf1) plays essential roles in mediating Rubisco assembly and carboxysome biogenesis.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
8JJR
 
 | | Cryo-EM structure of Symbiodinium photosystem I | | Descriptor: | (3S,3'R,5R,6S,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,beta-carotene-3,3'-diol, 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, ... | | Authors: | Zhao, L.S, Wang, N, Li, K, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2023-05-31 | | Release date: | 2024-05-15 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Architecture of symbiotic dinoflagellate photosystem I-light-harvesting supercomplex in Symbiodinium.
Nat Commun, 15, 2024
|
|
8Z11
 
 | | Cryo-EM structure of haptophyte photosystem I | | Descriptor: | (3S,3'R,5R,6S,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,beta-carotene-3,3'-diol, (3S,3'S,5R,5'R,6S,6'R,8'R)-3,5'-dihydroxy-8-oxo-6',7'-didehydro-5,5',6,6',7,8-hexahydro-5,6-epoxy-beta,beta-caroten-3'- yl acetate, 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, ... | | Authors: | He, F.Y, Zhao, L.S, Li, K, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-04-10 | | Release date: | 2025-01-22 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Structural insights into the assembly and energy transfer of haptophyte photosystem I-light-harvesting supercomplex.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8XR6
 | | Cryo-EM structure of cryptophyte photosystem II | | Descriptor: | (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E})-3,7,12,16-tetramethyl-18-[(4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohexen-1-yl]octadeca-3,5,7,9,11,13,15-heptaen-1,17-diynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-(2,6,6-trimethylcyclohexen-1-yl)octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, (6'R,11cis,11'cis,13cis,15cis)-4',5'-didehydro-5',6'-dihydro-beta,beta-carotene, ... | | Authors: | Li, K, Zhao, L.S, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2024-01-06 | | Release date: | 2024-06-19 | | Last modified: | 2025-01-01 | | Method: | ELECTRON MICROSCOPY (2.53 Å) | | Cite: | Structure of cryptophyte photosystem II-light-harvesting antennae supercomplex.
Nat Commun, 15, 2024
|
|
7Y7B
 
 | | Cryo-EM structure of cryptophyte photosystem I | | Descriptor: | (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E})-3,7,12,16-tetramethyl-18-[(4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohexen-1-yl]octadeca-3,5,7,9,11,13,15-heptaen-1,17-diynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-(2,6,6-trimethylcyclohexen-1-yl)octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-[(1~{R},4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohex-2-en-1-yl]octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, ... | | Authors: | Zhao, L.S, Li, K, Zhang, Y.Z, Liu, L.N. | | Deposit date: | 2022-06-22 | | Release date: | 2023-04-12 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2.66 Å) | | Cite: | Structural basis and evolution of the photosystem I-light-harvesting supercomplex of cryptophyte algae.
Plant Cell, 35, 2023
|
|
7Y8A
 
 | | Cryo-EM structure of cryptophyte photosystem I | | Descriptor: | (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E})-3,7,12,16-tetramethyl-18-[(4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohexen-1-yl]octadeca-3,5,7,9,11,13,15-heptaen-1,17-diynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-(2,6,6-trimethylcyclohexen-1-yl)octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-[(1~{R},4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohex-2-en-1-yl]octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, ... | | Authors: | Zhao, L.S, Zhang, Y.Z, Liu, L.N, Li, K. | | Deposit date: | 2022-06-23 | | Release date: | 2023-04-12 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (2.71 Å) | | Cite: | Structural basis and evolution of the photosystem I-light-harvesting supercomplex of cryptophyte algae.
Plant Cell, 35, 2023
|
|
7DDQ
 
 | | Structure of RC-LH1-PufX from Rhodobacter veldkampii | | Descriptor: | 1,2-Distearoyl-sn-glycerophosphoethanolamine, Antenna pigment protein alpha chain, Antenna pigment protein beta chain, ... | | Authors: | Bracun, L, Yamagata, A, Shirouzu, M, Liu, L.N. | | Deposit date: | 2020-10-29 | | Release date: | 2021-06-30 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Cryo-EM structure of the photosynthetic RC-LH1-PufX supercomplex at 2.8-angstrom resolution.
Sci Adv, 7, 2021
|
|
7VOR
 | | The structure of dimeric photosynthetic RC-LH1 supercomplex in Class-1 | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, ... | | Authors: | Cao, P, Li, M, Liu, L.N. | | Deposit date: | 2021-10-14 | | Release date: | 2022-04-27 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC-LH1 supercomplex.
Nat Commun, 13, 2022
|
|
7VOT
 | | The structure of dimeric photosynthetic RC-LH1 supercomplex in Class-2 | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN B, ... | | Authors: | Cao, P, Li, M, Liu, L.N. | | Deposit date: | 2021-10-14 | | Release date: | 2022-04-27 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC-LH1 supercomplex.
Nat Commun, 13, 2022
|
|
7VB9
 
 | | Rba sphaeroides PufY-KO RC-LH1 dimer type-2 | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, ... | | Authors: | Bracun, L, Yamagata, A, Liu, L.N, Shirouzu, M. | | Deposit date: | 2021-08-30 | | Release date: | 2022-05-04 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (3.45 Å) | | Cite: | Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC-LH1 supercomplex.
Nat Commun, 13, 2022
|
|
7VNM
 
 | | Rba sphaeroides PufY-KO RC-LH1 monomer | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, ... | | Authors: | Bracun, L, Yamagata, A, Liu, L.N, Shirouzu, M. | | Deposit date: | 2021-10-11 | | Release date: | 2022-05-04 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (2.86 Å) | | Cite: | Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC-LH1 supercomplex.
Nat Commun, 13, 2022
|
|
7VOY
 | | Rba sphaeroides PufX-KO RC-LH1 | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, FE (II) ION, ... | | Authors: | Bracun, L, Yamagata, A, Liu, L.N, Shirouzu, M. | | Deposit date: | 2021-10-15 | | Release date: | 2022-05-04 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC-LH1 supercomplex.
Nat Commun, 13, 2022
|
|
