1P0O
 | | HP (2-20) substitution of Trp for Gln and Asp at position 17 and 19 MODIFICATION IN SDS-D25 MICELLES | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P0L
 | | HP (2-20) Substitution GLN To TRP Modification In SDS-D25 MICELLES | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P5L
 
 | | HP (2-20) Substitution PHE5 to SER modification in sds-d25 micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-27 | | Release date: | 2003-06-03 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1OT0
 | | Structure of Antimicrobial Peptide, HP (2-20) and its Analogues Derived from Helicobacter pylori, as Determined by 1H NMR Spectroscopy | | Descriptor: | 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y, Hahm, K.-S, Kim, Y. | | Deposit date: | 2003-03-21 | | Release date: | 2004-09-07 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure of Antimicrobial Peptide, HP (2-20) and its Analogues Derived from Helicobacter pylori, as Determined by 1H NMR Spectroscopy
To be published
|
|
1P0J
 
 | | HP (2-20) Substitution ASP To TRP Modification In SDS-D25 Micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P5K
 
 | | HP (2-20) Substitution SER to LEU11 modification in sds-d25 micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-27 | | Release date: | 2003-06-03 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P0G
 
 | | Structure of Antimicrobial Peptide, HP (2-20) and its Analogues Derived from Helicobacter pylori, as Determined by 1H NMR Spectroscopy | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1VFT
 | | Crystal structure of L-cycloserine-bound form of alanine racemase from D-cycloserine-producing Streptomyces lavendulae | | Descriptor: | CHLORIDE ION, D-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-N,O-CYCLOSERYLAMIDE, alanine racemase | | Authors: | Noda, M, Matoba, Y, Kumagai, T, Sugiyama, M. | | Deposit date: | 2004-04-19 | | Release date: | 2004-09-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural evidence that alanine racemase from a D-cycloserine-producing microorganism exhibits resistance to its own product.
J.Biol.Chem., 279, 2004
|
|
5XU6
 
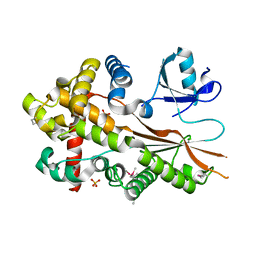 | | Crystal structure of inositol 1,3,4,5,6-pentakisphosphate 2-kinase (IPK1) from Cryptococcus neoformans | | Descriptor: | Inositol-pentakisphosphate 2-kinase, SULFATE ION | | Authors: | Oh, J, Rhee, S. | | Deposit date: | 2017-06-22 | | Release date: | 2017-10-04 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of inositol 1,3,4,5,6-pentakisphosphate 2-kinase from Cryptococcus neoformans.
J. Struct. Biol., 200, 2017
|
|
2LA2
 
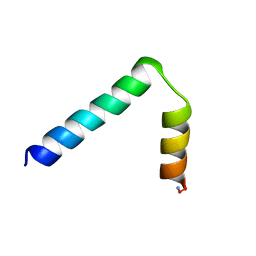 | |
2LN4
 
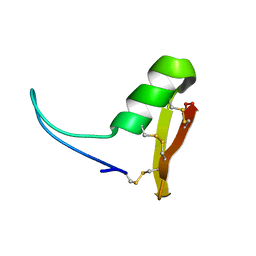 | |
1VFH
 
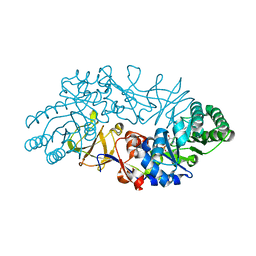 | | Crystal structure of alanine racemase from D-cycloserine producing Streptomyces lavendulae | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, alanine racemase | | Authors: | Noda, M, Matoba, Y, Kumagai, T, Sugiyama, M. | | Deposit date: | 2004-04-13 | | Release date: | 2004-09-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural evidence that alanine racemase from a D-cycloserine-producing microorganism exhibits resistance to its own product.
J.Biol.Chem., 279, 2004
|
|
1VFS
 | | Crystal structure of D-cycloserine-bound form of alanine racemase from D-cycloserine-producing Streptomyces lavendulae | | Descriptor: | CHLORIDE ION, D-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-N,O-CYCLOSERYLAMIDE, alanine racemase | | Authors: | Noda, M, Matoba, Y, Kumagai, T, Sugiyama, M. | | Deposit date: | 2004-04-19 | | Release date: | 2004-09-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural evidence that alanine racemase from a D-cycloserine-producing microorganism exhibits resistance to its own product.
J.Biol.Chem., 279, 2004
|
|
